Submitted:
26 June 2025
Posted:
30 June 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
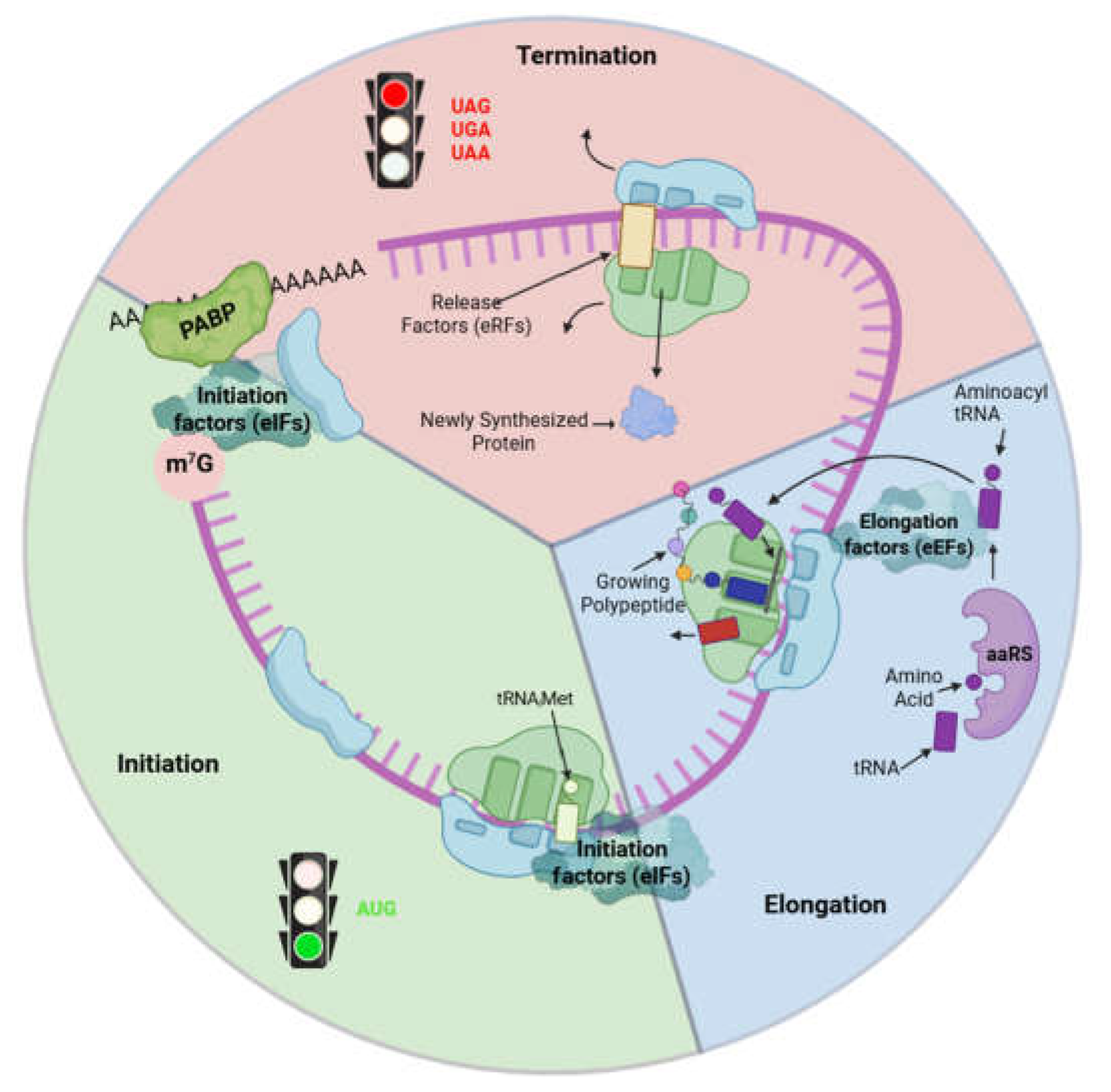
2. Techniques and Methods to Investigate mRNA Translation in the Heart
2.1. Biochemical methods for studying translational control
2.1.1. RNA-binding protein immunoprecipitation (RIP) to identify bound target RNAs
2.1.2. Crosslinking and immunoprecipitation (CLIP) to map RBP-binding sites on RNAs
2.1.3. In vitro pulldown of interacting proteins of biotinylated RNA
2.1.4. Proximity ligation assay associated with immunoblot or mass spectrometry
2.1.5. Puromycin incorporation assay to assess global translation efficiency
2.2. Deep sequencing-based translatome profiling in cells and animals
2.2.1. Polysome profiling-sequencing (polysome-seq)
2.2.2. Translating ribosome affinity purification sequencing (TRAP-seq)
2.2.3. Translational landscape in human and mouse heart failure determined by ribosome profiling (Ribo-seq)
2.3. Imaging-based techniques for evaluating translation efficiency and localized translation in cardiomyocytes
3. Translational Control in Cardiac Development and Congenital Heart Disease
3.1. Human genetic mutations in translation machinery and congenital heart disease
3.1.1. Diamond Blackfan Anemia and other heart disease-causing mutations in cytoplasmic translation factors
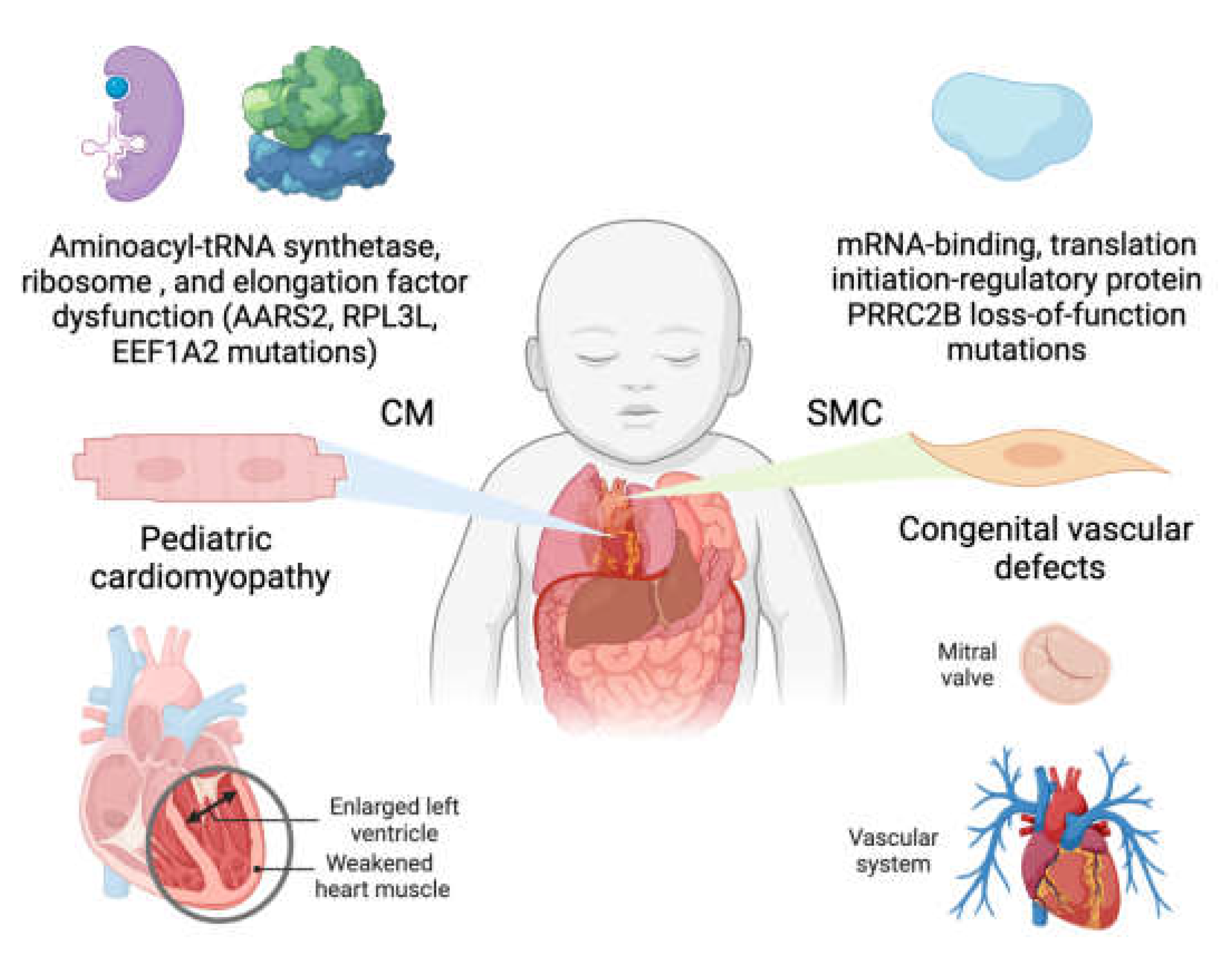
3.1.2. Human mutations in mitochondrial translation machinery lead to genetic cardiomyopathy
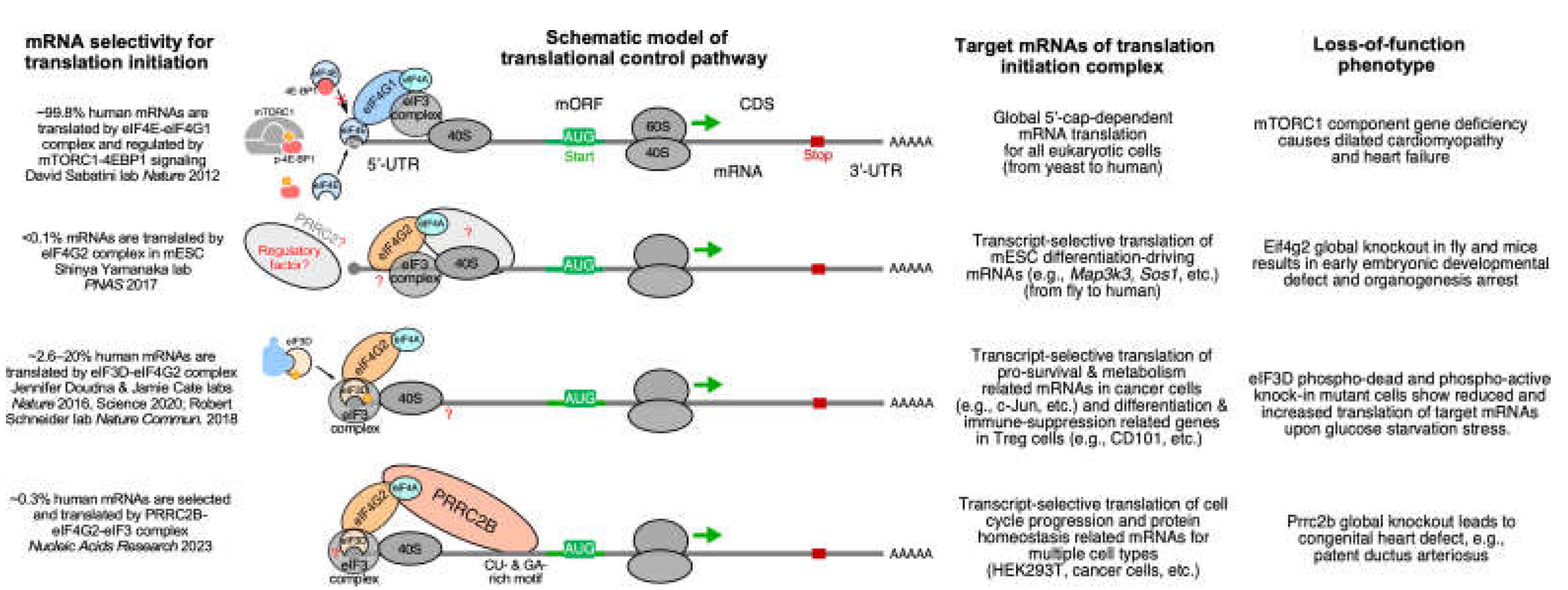
3.1.3. Loss-of-function of PRRC2B-mediated translation initiation regulation causes congenital cardiovascular defect in humans and mice
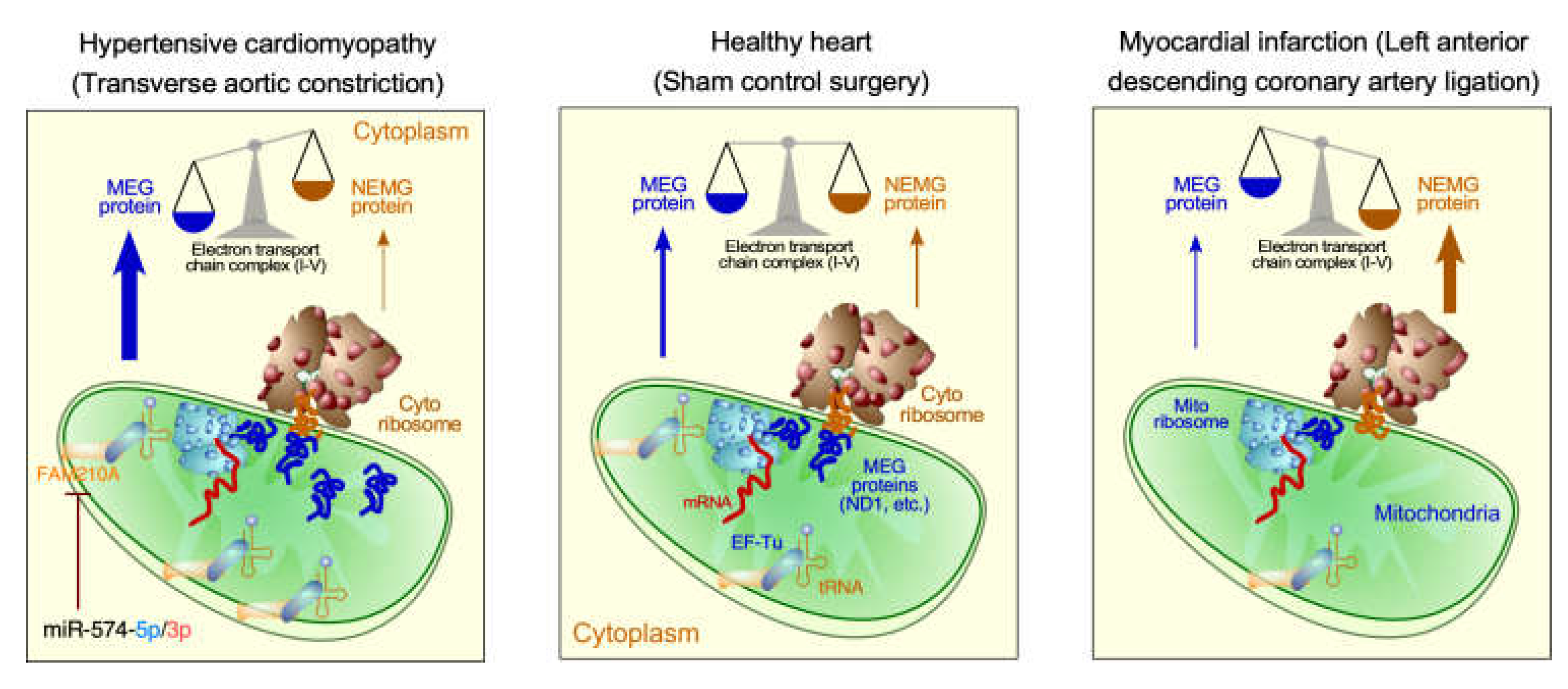
3.2. Translational control in mitochondrial cardiomyopathy
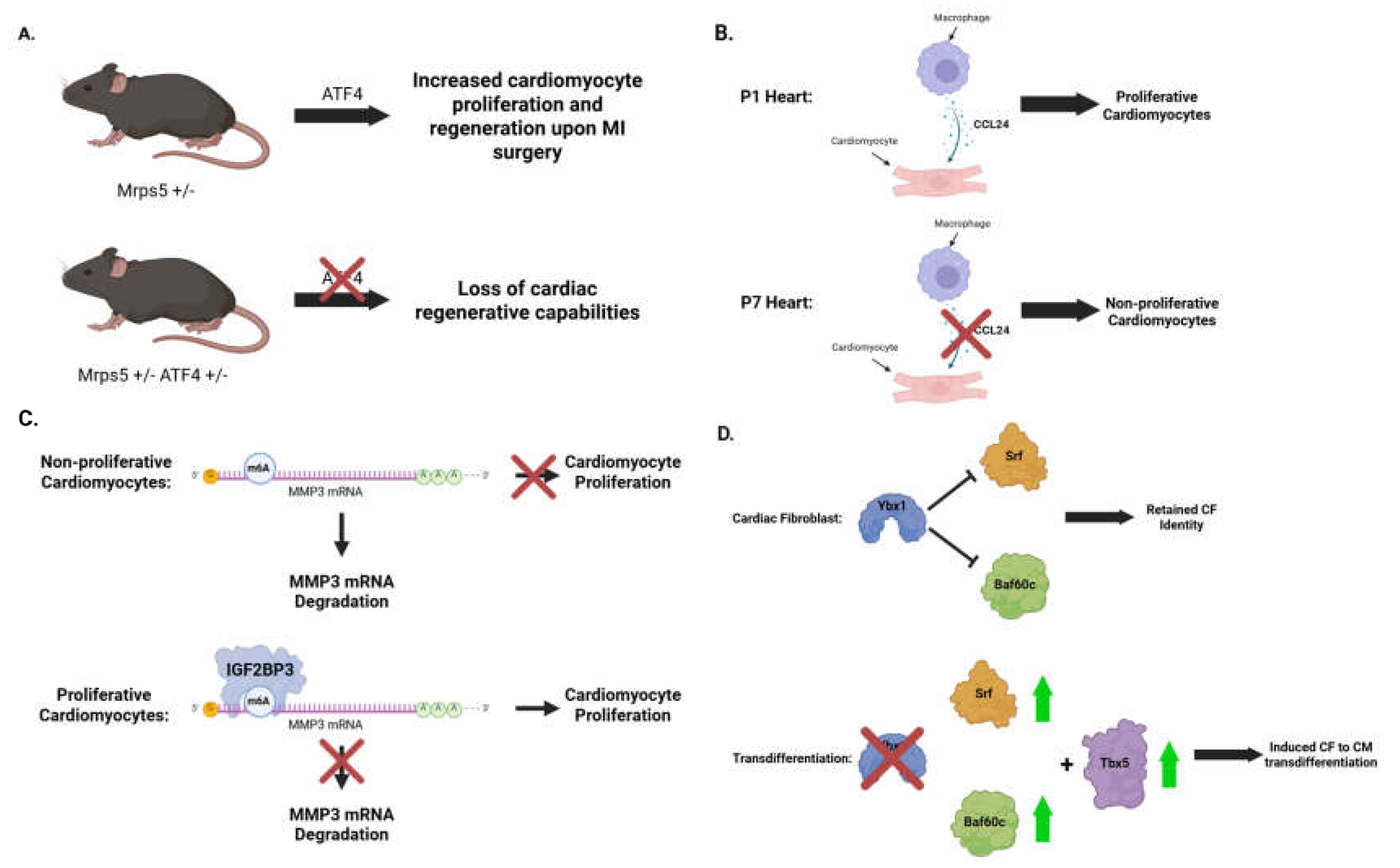
3.3. Translational regulation of cardiac cell proliferation and differentiation
4. Translational Control in Adult Cardiac Disease
4.1. Translational control in cardiomyocyte hypertrophy
4.1.1. Role of translation initiation factors in cardiac hypertrophy
4.1.2. Role of translation elongation factors in cardiac hypertrophy
4.1.3. Genetic loss-of-function of mTORC1 causes heart failure in mice
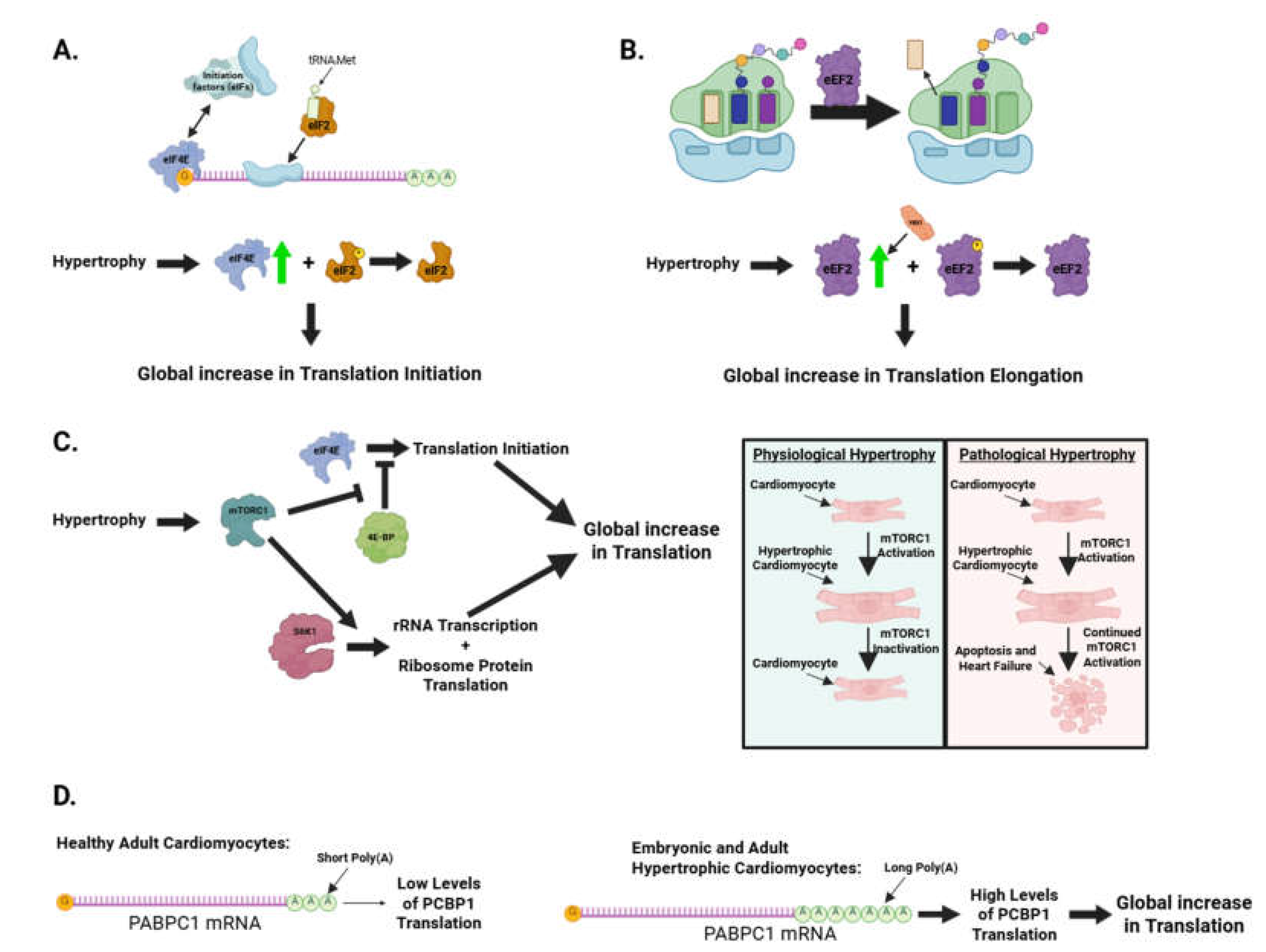
4.1.4. PABPC1-mediated translational control of physiological and pathological cardiac hypertrophy
4.1.5. Translational control of Ybx1 expression regulates cardiac function in response to pressure overload in vivo
4.2. Translational control in cardiac fibroblast activation during fibrosis
4.2.1. Translational regulation in human TGFβ-activated cardiac fibroblasts
4.2.2. EPRS1 promotes cardiac fibrosis by enhancing proline-rich extracellular matrix protein translation
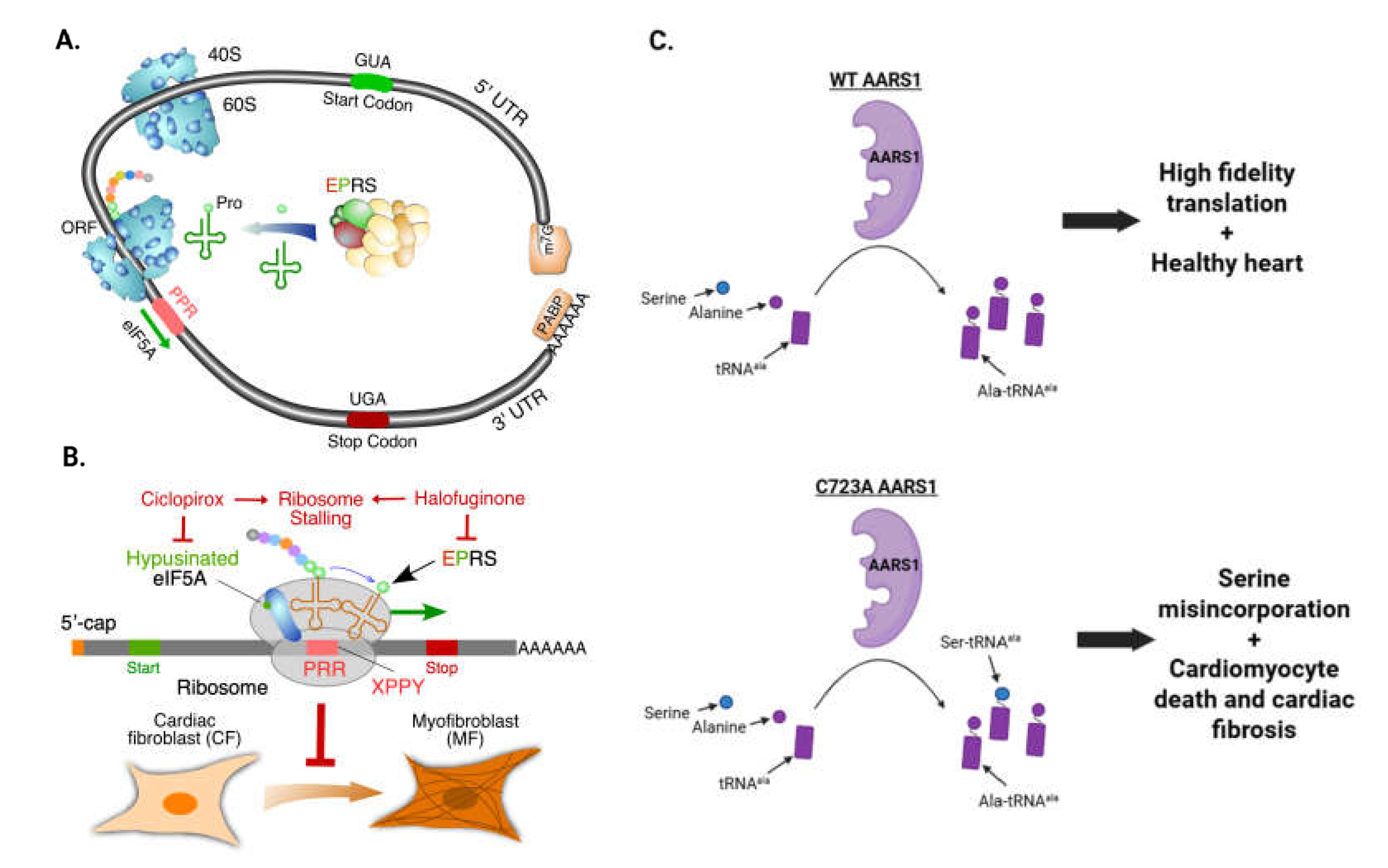
4.2.4. Editing-defective Aars1 mouse shows spontaneous cardiac proteinopathy and fibrosis
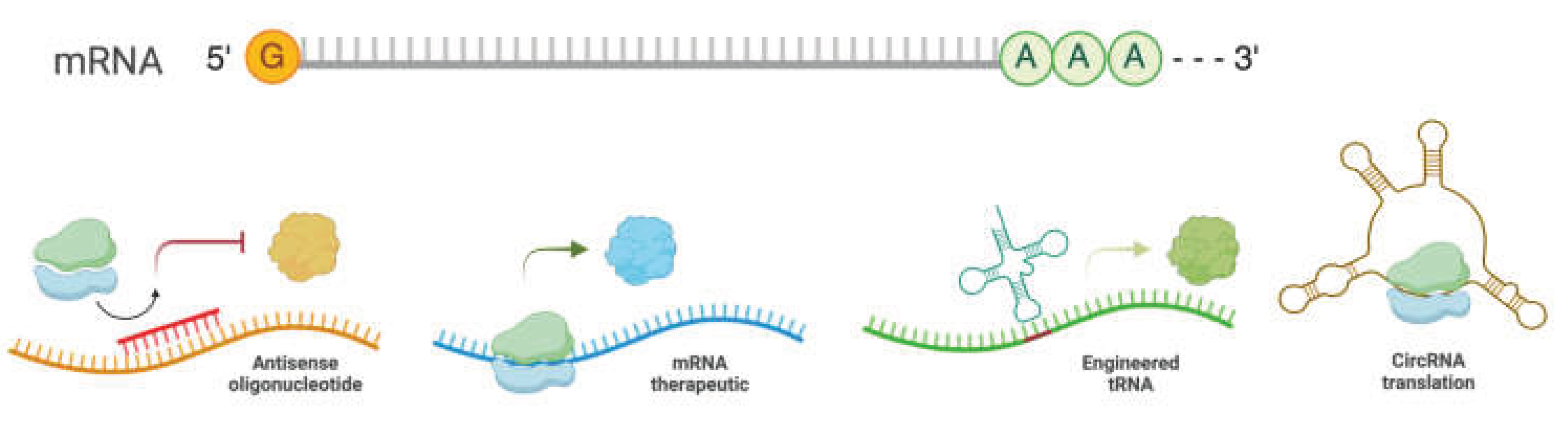
5. Translation-Manipulating Therapeutics for Heart Disease Treatment
5.1. Translation-targeted medicines for cardiac disorders
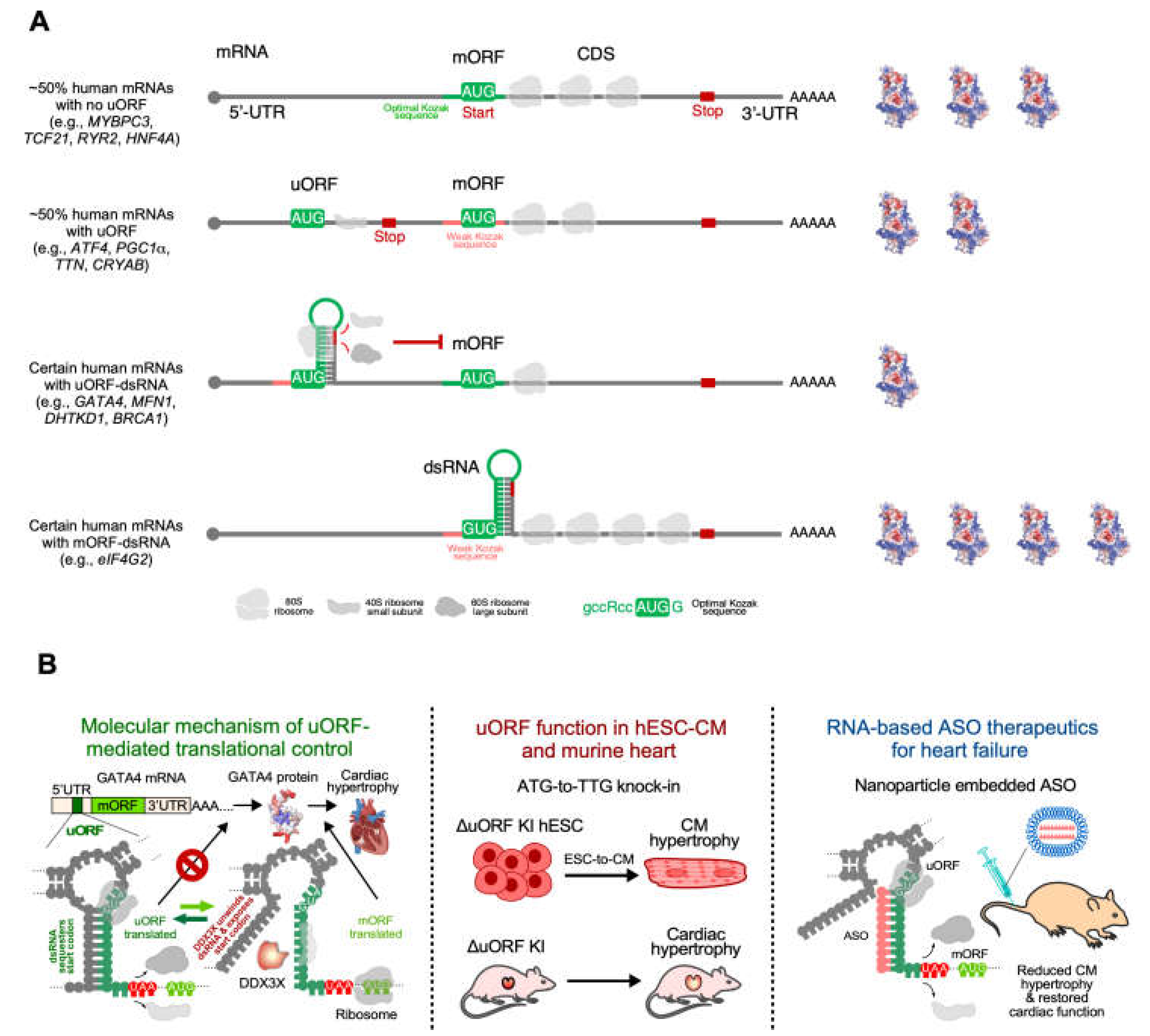
5.2. RNA secondary structure as a therapeutic target for ASO treatment of cardiac hypertrophy
5.3. Chemically modified mRNA-based CAR-T-mediated therapeutics for cardiac fibrosis
5.4. Chemically modified mRNA-based therapeutics for ischemic heart disease
6. Concluding Remarks and Future Perspective
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Martin SS, Aday AW, Almarzooq ZI, Anderson CAM, Arora P, Avery CL, et al. 2024 Heart Disease and Stroke Statistics: A Report of US and Global Data From the American Heart Association. Circulation. 2024;149(8):e347-e913.
- Simmonds SJ, Cuijpers I, Heymans S, Jones EAV. Cellular and Molecular Differences between HFpEF and HFrEF: A Step Ahead in an Improved Pathological Understanding. Cells. 2020;9(1).
- Schimmel, P. Aminoacyl tRNA synthetases: general scheme of structure-function relationships in the polypeptides and recognition of transfer RNAs. Annu Rev Biochem. 1987;56:125-58.
- Ibba M, Söll D. Aminoacyl-tRNA synthesis. Annu Rev Biochem. 2000;69:617-50.
- Yadavalli SS, Ibba M. Quality control in aminoacyl-tRNA synthesis its role in translational fidelity. Adv Protein Chem Struct Biol. 2012;86:1-43.
- Ling J, Reynolds N, Ibba M. Aminoacyl-tRNA synthesis and translational quality control. Annu Rev Microbiol. 2009;63:61-78.
- Liu Y, Beyer A, Aebersold R. On the Dependency of Cellular Protein Levels on mRNA Abundance. Cell. 2016;165(3):535-50.
- Schafer S, Viswanathan S, Widjaja AA, Lim WW, Moreno-Moral A, DeLaughter DM, et al. IL-11 is a crucial determinant of cardiovascular fibrosis. Nature. 2017;552(7683):110-5.
- Gao C, Wang Y. mRNA Metabolism in Cardiac Development and Disease: Life After Transcription. Physiol Rev. 2020;100(2):673-94.
- Volkers M, Preiss T, Hentze MW. RNA-binding proteins in cardiovascular biology and disease: the beat goes on. Nat Rev Cardiol. 2024.
- Hedaya OM, Venkata Subbaiah KC, Jiang F, Xie LH, Wu J, Khor ES, et al. Secondary structures that regulate mRNA translation provide insights for ASO-mediated modulation of cardiac hypertrophy. Nat Commun. 2023;14(1):6166.
- Yao P, Potdar AA, Arif A, Ray PS, Mukhopadhyay R, Willard B, et al. Coding Region Polyadenylation Generates a Truncated tRNA Synthetase that Counters Translation Repression. Cell. 2012;149(1):88-100.
- Jiang F, Hedaya OM, Khor E, Wu J, Auguste M, Yao P. RNA binding protein PRRC2B mediates translation of specific mRNAs and regulates cell cycle progression. Nucleic Acids Res. 2023;51(11):5831-46.
- Anastasakis DG, Apostolidi M, Garman KA, Polash AH, Umar MI, Meng Q, et al. Nuclear PKM2 binds pre-mRNA at folded G-quadruplexes and reveals their gene regulatory role. Mol Cell. 2024;84(19):3775-89 e6.
- Kim B, Kim VN. fCLIP-seq for transcriptomic footprinting of dsRNA-binding proteins: Lessons from DROSHA. Methods. 2019;152:3-11.
- Danan C, Manickavel S, Hafner M. PAR-CLIP: A Method for Transcriptome-Wide Identification of RNA Binding Protein Interaction Sites. Methods Mol Biol. 2016;1358:153-73.
- Panda AC, Martindale JL, Gorospe M. Affinity Pulldown of Biotinylated RNA for Detection of Protein-RNA Complexes. Bio Protoc. 2016;6(24).
- Sears RM, May DG, Roux KJ. BioID as a Tool for Protein-Proximity Labeling in Living Cells. Methods Mol Biol. 2019;2012:299-313.
- Mellacheruvu D, Wright Z, Couzens AL, Lambert JP, St-Denis NA, Li T, et al. The CRAPome: a contaminant repository for affinity purification-mass spectrometry data. Nat Methods. 2013;10(8):730-6.
- de la Parra C, Ernlund A, Alard A, Ruggles K, Ueberheide B, Schneider RJ. A widespread alternate form of cap-dependent mRNA translation initiation. Nat Commun. 2018;9(1):3068.
- Liakath-Ali K, Mills EW, Sequeira I, Lichtenberger BM, Pisco AO, Sipila KH, et al. An evolutionarily conserved ribosome-rescue pathway maintains epidermal homeostasis. Nature. 2018;556(7701):376-80.
- Wu J, Xie HL, Jiang F, Mickelson DJ, Myers JR, Tang WH, et al. EPRS regulates proline-rich pro-fibrotic protein synthesis during cardiac fibrosis. bioRxiv. 2019.
- Zhou P, Zhang Y, Ma Q, Gu F, Day DS, He A, et al. Interrogating translational efficiency and lineage-specific transcriptomes using ribosome affinity purification. Proc Natl Acad Sci U S A. 2013;110(38):15395-400.
- Sanz E, Yang L, Su T, Morris DR, McKnight GS, Amieux PS. Cell-type-specific isolation of ribosome-associated mRNA from complex tissues. Proc Natl Acad Sci U S A. 2009;106(33):13939-44.
- Busch JD, Cipullo M, Atanassov I, Bratic A, Silva Ramos E, Schondorf T, et al. MitoRibo-Tag Mice Provide a Tool for In Vivo Studies of Mitoribosome Composition. Cell Rep. 2019;29(6):1728-38 e9.
- van Heesch S, Witte F, Schneider-Lunitz V, Schulz JF, Adami E, Faber AB, et al. The Translational Landscape of the Human Heart. Cell. 2019;178(1):242-60.e29.
- Schafer S, Viswanathan S, Widjaja AA, Lim W-W, Moreno-Moral A, DeLaughter DM, et al. IL-11 is a crucial determinant of cardiovascular fibrosis. Nature. 2017;552(7683):110-5.
- Chothani S, Schäfer S, Adami E, Viswanathan S, Widjaja AA, Langley SR, et al. Widespread Translational Control of Fibrosis in the Human Heart by RNA-Binding Proteins. Circulation. 2019;140(11):937-51.
- Lewis YE, Moskovitz A, Mutlak M, Heineke J, Caspi LH, Kehat I. Localization of transcripts, translation, and degradation for spatiotemporal sarcomere maintenance. J Mol Cell Cardiol. 2018;116:16-28.
- Scarborough EA, Uchida K, Vogel M, Erlitzki N, Iyer M, Phyo SA, et al. Microtubules orchestrate local translation to enable cardiac growth. Nat Commun. 2021;12(1):1547.
- Bogdanov V, Soltisz AM, Moise N, Sakuta G, Orengo BH, Janssen PML, et al. Distributed synthesis of sarcolemmal and sarcoplasmic reticulum membrane proteins in cardiac myocytes. Basic Res Cardiol. 2021;116(1):63.
- Eichel CA, Rios-Perez EB, Liu F, Jameson MB, Jones DK, Knickelbine JJ, et al. A microtranslatome coordinately regulates sodium and potassium currents in the human heart. Elife. 2019;8.
- Tominaga M, Hamanoue S, Goto H, Saito T, Nagai JI, Masuno M, et al. Diamond-Blackfan anemia caused by chromosome 1p22 deletion encompassing RPL5. Hum Genome Var. 2019;6:36.
- Ulirsch JC, Verboon JM, Kazerounian S, Guo MH, Yuan D, Ludwig LS, et al. The Genetic Landscape of Diamond-Blackfan Anemia. Am J Hum Genet. 2019;104(2):356.
- Vlachos A, Osorio DS, Atsidaftos E, Kang J, Lababidi ML, Seiden HS, et al. Increased Prevalence of Congenital Heart Disease in Children With Diamond Blackfan Anemia Suggests Unrecognized Diamond Blackfan Anemia as a Cause of Congenital Heart Disease in the General Population: A Report of the Diamond Blackfan Anemia Registry. Circ Genom Precis Med. 2018;11(5):e002044.
- Liu YL, Shibuya A, Glader B, Wilkes MC, Barna M, Sakamoto KM. Animal models of Diamond-Blackfan anemia: updates and challenges. Haematologica. 2023;108(5):1222-31.
- Grimes KM, Prasad V, Huo J, Kuwabara Y, Vanhoutte D, Baldwin TA, et al. Rpl3l gene deletion in mice reduces heart weight over time. Front Physiol. 2023;14:1054169.
- Shiraishi C, Matsumoto A, Ichihara K, Yamamoto T, Yokoyama T, Mizoo T, et al. RPL3L-containing ribosomes determine translation elongation dynamics required for cardiac function. Nat Commun. 2023;14(1):2131.
- Milenkovic I, Santos Vieira HG, Lucas MC, Ruiz-Orera J, Patone G, Kesteven S, et al. Dynamic interplay between RPL3- and RPL3L-containing ribosomes modulates mitochondrial activity in the mammalian heart. Nucleic Acids Res. 2023;51(11):5301-24.
- Wu J, Subbaiah KCV, Hedaya O, Chen S, Munger J, Tang WHW, et al. FAM210A regulates mitochondrial translation and maintains cardiac mitochondrial homeostasis. Cardiovasc Res. 2023;119(14):2441-57.
- Michael, R. Murphy MG, Teresa M. Lee, Joshua M. Fisher, Megha V. Patel, Parul Jayakar, Amanda Buchanan, Rajesh K. Soni, Yue Yin, Feiyue Yang, Muredach P. Reilly, Wendy K. Chung, Xuebing Wu. Recessive but damaging alleles of muscle-specific ribosomal protein gene RPL3L drive neonatal dilated cardiomyopathy. In: Columbia University Irving Medical Center, editor. BioRxiv2025.
- Methawasin M, Zhang Y, Gregorich ZR, He Y, Liu C, Muldoon J, et al. Reducing Granules Without Splicing Restoration Alleviates RBM20 Cardiomyopathy. Circ Res. 2025;136(10):1134-46.
- Witte F, Ruiz-Orera J, Mattioli CC, Blachut S, Adami E, Schulz JF, et al. A trans locus causes a ribosomopathy in hypertrophic hearts that affects mRNA translation in a protein length-dependent fashion. Genome Biol. 2021;22(1):191.
- Khalyfa A, Bourbeau D, Chen E, Petroulakis E, Pan J, Xu S, et al. Characterization of elongation factor-1A (eEF1A-1) and eEF1A-2/S1 protein expression in normal and wasted mice. J Biol Chem. 2001;276(25):22915-22.
- Cao S, Smith LL, Padilla-Lopez SR, Guida BS, Blume E, Shi J, et al. Homozygous EEF1A2 mutation causes dilated cardiomyopathy, failure to thrive, global developmental delay, epilepsy and early death. Human Molecular Genetics. 2017;26(18):3545-52.
- Feng W, Wang L, Veevers J, Liu C, Huang T, Chen J. Loss of eEF1A2 (Eukaryotic Elongation Factor 1 A2) in Murine Myocardium Results in Dilated Cardiomyopathy. Circ Heart Fail. 2021;14(10):e008665.
- Boczonadi V, Horvath R. Mitochondria: impaired mitochondrial translation in human disease. Int J Biochem Cell Biol. 2014;48(100):77-84.
- Euro L, Konovalova S, Asin-Cayuela J, Tulinius M, Griffin H, Horvath R, et al. Structural modeling of tissue-specific mitochondrial alanyl-tRNA synthetase (AARS2) defects predicts differential effects on aminoacylation. Frontiers in Genetics. 2015;6.
- Götz A, Tyynismaa H, Euro L, Ellonen P, Hyötyläinen T, Ojala T, et al. Exome Sequencing Identifies Mitochondrial Alanyl-tRNA Synthetase Mutations in Infantile Mitochondrial Cardiomyopathy. The American Journal of Human Genetics. 2011;88(5):635-42.
- Sommerville EW, Zhou X-L, Oláhová M, Jenkins J, Euro L, Konovalova S, et al. Instability of the mitochondrial alanyl-tRNA synthetase underlies fatal infantile-onset cardiomyopathy. Human Molecular Genetics. 2018;28(2):258-68.
- Rudler DL, Hughes LA, Perks KL, Richman TR, Kuznetsova I, Ermer JA, et al. Fidelity of translation initiation is required for coordinated respiratory complex assembly. Science Advances. 2019;5(12):eaay2118.
- Gebauer F, Schwarzl T, Valcarcel J, Hentze MW. RNA-binding proteins in human genetic disease. Nat Rev Genet. 2021;22(3):185-98.
- Sun J, Wang L, Matthews RC, Walcott GP, Lu YA, Wei Y, et al. CCND2 Modified mRNA Activates Cell Cycle of Cardiomyocytes in Hearts With Myocardial Infarction in Mice and Pigs. Circ Res. 2023;133(6):484-504.
- Bohlen J, Roiuk M, Neff M, Teleman AA. PRRC2 proteins impact translation initiation by promoting leaky scanning. Nucleic Acids Res. 2023;51(7):3391-409.
- Das D, Khor ES, Jiang F, He J, Kawakami Y, Wainwright L, et al. Loss-of-function of RNA-binding protein PRRC2B causes translational defects and congenital cardiovascular malformation. medRxiv. 2024.
- Goldberg N, Bril D, Eisenstein M, Olender T, Savidor A, Bialik S, et al. The RNA-binding protein PRRC2B preserves 5’ TOP mRNA during starvation to maintain ribosome biogenesis during nutrient recovery. bioRxiv. 2024:2024.12.04.626744.
- Rao A, Lyu B, Jahan I, Lubertozzi A, Zhou G, Tedeschi F, et al. The translation initiation factor homolog eif4e1c regulates cardiomyocyte metabolism and proliferation during heart regeneration. Development. 2023;150(20).
- Jobava R, Mao Y, Guan B-J, Hu D, Krokowski D, Chen C-W, et al. Adaptive translational pausing is a hallmark of the cellular response to severe environmental stress. Molecular Cell. 2021;81(20):4191-208.e8.
- Santos-Ribeiro D, Godinas L, Pilette C, Perros F. The integrated stress response system in cardiovascular disease. Drug Discovery Today. 2018;23(4):920-9.
- Estrada K, Styrkarsdottir U, Evangelou E, Hsu YH, Duncan EL, Ntzani EE, et al. Genome-wide meta-analysis identifies 56 bone mineral density loci and reveals 14 loci associated with risk of fracture. Nat Genet. 2012;44(5):491-501.
- Tanaka K-i, Xue Y, Nguyen-Yamamoto L, Morris JA, Kanazawa I, Sugimoto T, et al. FAM210A is a novel determinant of bone and muscle structure and strength. Proceedings of the National Academy of Sciences. 2018;115(16):E3759-E68.
- Wu J, Subbaiah KCV, Hedaya O, Chen S, Munger J, Tang WHW, et al. FAM210A regulates mitochondrial translation and maintains cardiac mitochondrial homeostasis. Cardiovasc Res. 2023.
- van Rooij E, Sutherland LB, Thatcher JE, DiMaio JM, Naseem RH, Marshall WS, et al. Dysregulation of microRNAs after myocardial infarction reveals a role of miR-29 in cardiac fibrosis. Proc Natl Acad Sci U S A. 2008;105(35):13027-32.
- Wu J, Venkata Subbaiah KC, Jiang F, Hedaya O, Mohan A, Yang T, et al. MicroRNA-574 regulates FAM210A expression and influences pathological cardiac remodeling. EMBO Mol Med. 2020:e12710.
- Hollinger J, Wu J, Awayda KM, O'Connell MR, Yao P. Expression and purification of the mitochondrial transmembrane protein FAM210A in Escherichia coli. Protein Expr Purif. 2023;210:106322.
- Mohamed TMA, Ang YS, Radzinsky E, Zhou P, Huang Y, Elfenbein A, et al. Regulation of Cell Cycle to Stimulate Adult Cardiomyocyte Proliferation and Cardiac Regeneration. Cell. 2018;173(1):104-16 e12.
- Aballo TJ, Roberts DS, Bayne EF, Zhu W, Walcott G, Mahmoud AI, et al. Integrated proteomics reveals alterations in sarcomere composition and developmental processes during postnatal swine heart development. J Mol Cell Cardiol. 2023;176:33-40.
- Zhao M, Nakada Y, Wei Y, Bian W, Chu Y, Borovjagin AV, et al. Cyclin D2 Overexpression Enhances the Efficacy of Human Induced Pluripotent Stem Cell-Derived Cardiomyocytes for Myocardial Repair in a Swine Model of Myocardial Infarction. Circulation. 2021;144(3):210-28.
- Heallen T, Zhang M, Wang J, Bonilla-Claudio M, Klysik E, Johnson RL, et al. Hippo pathway inhibits Wnt signaling to restrain cardiomyocyte proliferation and heart size. Science. 2011;332(6028):458-61.
- Bartsch D, Kalamkar K, Ahuja G, Lackmann JW, Hescheler J, Weber T, et al. mRNA translational specialization by RBPMS presets the competence for cardiac commitment in hESCs. Sci Adv. 2023;9(13):eade1792.
- Gao F, Liang T, Lu YW, Pu L, Fu X, Dong X, et al. Reduced Mitochondrial Protein Translation Promotes Cardiomyocyte Proliferation and Heart Regeneration. Circulation. 2023;148(23):1887-906.
- Stoehr A, Kennedy L, Yang Y, Patel S, Lin Y, Linask KL, et al. The ribosomal prolyl-hydroxylase OGFOD1 decreases during cardiac differentiation and modulates translation and splicing. JCI Insight. 2019;5(13).
- Porrello ER, Mahmoud AI, Simpson E, Hill JA, Richardson JA, Olson EN, et al. Transient regenerative potential of the neonatal mouse heart. Science. 2011;331(6020):1078-80.
- Wang Z, Cui M, Shah AM, Ye W, Tan W, Min YL, et al. Mechanistic basis of neonatal heart regeneration revealed by transcriptome and histone modification profiling. Proc Natl Acad Sci U S A. 2019;116(37):18455-65.
- Aurora AB, Porrello ER, Tan W, Mahmoud AI, Hill JA, Bassel-Duby R, et al. Macrophages are required for neonatal heart regeneration. J Clin Invest. 2014;124(3):1382-92.
- Huang H, Weng H, Sun W, Qin X, Shi H, Wu H, et al. Recognition of RNA N(6)-methyladenosine by IGF2BP proteins enhances mRNA stability and translation. Nat Cell Biol. 2018;20(3):285-95.
- Li S, Shen S, Xu H, Cai S, Yuan X, Wang C, et al. IGF2BP3 promotes adult myocardial regeneration by stabilizing MMP3 mRNA through interaction with m6A modification. Cell Death Discov. 2023;9(1):164.
- Caudron-Herger M, Jansen RE, Wassmer E, Diederichs S. RBP2GO: a comprehensive pan-species database on RNA-binding proteins, their interactions and functions. Nucleic Acids Res. 2021;49(D1):D425-D36.
- Nakano K, Sadahiro T, Fujita R, Isomi M, Abe Y, Yamada Y, et al. Development of adeno-associated viral vectors targeting cardiac fibroblasts for efficient in vivo cardiac reprogramming. Stem Cell Reports. 2024;19(10):1389-98.
- Tani H, Sadahiro T, Yamada Y, Isomi M, Yamakawa H, Fujita R, et al. Direct Reprogramming Improves Cardiac Function and Reverses Fibrosis in Chronic Myocardial Infarction. Circulation. 2023;147(3):223-38.
- Xie Y, Wang Q, Yang Y, Near D, Wang H, Colon M, et al. Translational landscape of direct cardiac reprogramming reveals a role of Ybx1 in repressing cardiac fate acquisition. Nat Cardiovasc Res. 2023;2(11):1060-77.
- Wang L, Liu Z, Yin C, Asfour H, Chen O, Li Y, et al. Stoichiometry of Gata4, Mef2c, and Tbx5 influences the efficiency and quality of induced cardiac myocyte reprogramming. Circ Res. 2015;116(2):237-44.
- Kim S, Coulombe PA. Emerging role for the cytoskeleton as an organizer and regulator of translation. Nat Rev Mol Cell Biol. 2010;11(1):75-81.
- Hannan RD, Jenkins A, Jenkins AK, Brandenburger Y. Cardiac hypertrophy: a matter of translation. Clin Exp Pharmacol Physiol. 2003;30(8):517-27.
- Chorghade S, Seimetz J, Emmons R, Yang J, Bresson SM, Lisio M, et al. Poly(A) tail length regulates PABPC1 expression to tune translation in the heart. Elife. 2017;6.
- Gudbjarnason S, Telerman M, Chiba C, Wolf PL, Bing RJ. Myocardial Protein Synthesis in Cardiac Hypertrophy. J Lab Clin Med. 1964;63:244-53.
- Gonzalez-Teran B, Lopez JA, Rodriguez E, Leiva L, Martinez-Martinez S, Bernal JA, et al. p38gamma and delta promote heart hypertrophy by targeting the mTOR-inhibitory protein DEPTOR for degradation. Nat Commun. 2016;7:10477.
- Dorn GW, 2nd, Force T. Protein kinase cascades in the regulation of cardiac hypertrophy. J Clin Invest. 2005;115(3):527-37.
- Sciarretta S, Volpe M, Sadoshima J. Mammalian target of rapamycin signaling in cardiac physiology and disease. Circ Res. 2014;114(3):549-64.
- Laplante M, Sabatini DM. mTOR signaling in growth control and disease. Cell. 2012;149(2):274-93.
- Brandenburger Y, Arthur JF, Woodcock EA, Du XJ, Gao XM, Autelitano DJ, et al. Cardiac hypertrophy in vivo is associated with increased expression of the ribosomal gene transcription factor UBF. FEBS Lett. 2003;548(1-3):79-84.
- Luyken J, Hannan RD, Cheung JY, Rothblum LI. Regulation of rDNA transcription during endothelin-1-induced hypertrophy of neonatal cardiomyocytes. Hyperphosphorylation of upstream binding factor, an rDNA transcription factor. Circ Res. 1996;78(3):354-61.
- Hannan RD, Luyken J, Rothblum LI. Regulation of ribosomal DNA transcription during contraction-induced hypertrophy of neonatal cardiomyocytes. J Biol Chem. 1996;271(6):3213-20.
- Pestova TV, Kolupaeva VG, Lomakin IB, Pilipenko EV, Shatsky IN, Agol VI, et al. Molecular mechanisms of translation initiation in eukaryotes. Proc Natl Acad Sci U S A. 2001;98(13):7029-36.
- Gebauer F, Hentze MW. Molecular mechanisms of translational control. Nat Rev Mol Cell Biol. 2004;5(10):827-35.
- Lazaris-Karatzas A, Montine KS, Sonenberg N. Malignant transformation by a eukaryotic initiation factor subunit that binds to mRNA 5' cap. Nature. 1990;345(6275):544-7.
- Ventoso I, Blanco R, Perales C, Carrasco L. HIV-1 protease cleaves eukaryotic initiation factor 4G and inhibits cap-dependent translation. Proceedings of the National Academy of Sciences. 2001;98(23):12966.
- Wada H, Ivester CT, Carabello BA, Cooper G, McDermott PJ. Translational Initiation Factor eIF-4E: A LINK BETWEEN CARDIAC LOAD AND PROTEIN SYNTHESIS. Journal of Biological Chemistry. 1996;271(14):8359-64.
- Tuxworth WJ, Wada H, Ishibashi Y, McDermott PJ. Role of load in regulating eIF-4F complex formation in adult feline cardiocytes. American Journal of Physiology-Heart and Circulatory Physiology. 1999;277(4):H1273-H82.
- Makhlouf AA, Namboodiri AMS, McDermott PJ. Transcriptional regulation of the rat eIF4E gene in cardiac muscle cells: the role of specific elements in the promoter region. Gene. 2001;267(1):1-12.
- Nagatomo Y, Carabello BA, Hamawaki M, Nemoto S, Matsuo T, McDermott PJ. Translational mechanisms accelerate the rate of protein synthesis during canine pressure-overload hypertrophy. American Journal of Physiology-Heart and Circulatory Physiology. 1999;277(6):H2176-H84.
- Makhlouf AA, McDermott PJ. Increased expression of eukaryotic initiation factor 4E during growth of neonatal rat cardiocytes in vitro. American Journal of Physiology-Heart and Circulatory Physiology. 1998;274(6):H2133-H42.
- Gingras A-C, Raught B, Sonenberg N. eIF4 Initiation Factors: Effectors of mRNA Recruitment to Ribosomes and Regulators of Translation. Annual Review of Biochemistry. 1999;68(1):913-63.
- Saghir AN, Tuxworth WJ, Jr., Hagedorn CH, McDermott PJ. Modifications of eukaryotic initiation factor 4F (eIF4F) in adult cardiocytes by adenoviral gene transfer: differential effects on eIF4F activity and total protein synthesis rates. Biochem J. 2001;356(Pt 2):557-66.
- Hardt SE, Tomita H, Katus HA, Sadoshima J. Phosphorylation of Eukaryotic Translation Initiation Factor 2Bε by Glycogen Synthase Kinase-3β Regulates β-Adrenergic Cardiac Myocyte Hypertrophy. Circulation Research. 2004;94(7):926-35.
- Lu Z, Xu X, Fassett J, Kwak D, Liu X, Hu X, et al. Loss of the Eukaryotic Initiation Factor 2α Kinase General Control Nonderepressible 2 Kinase Protects Mice From Pressure Overload-Induced Congestive Heart Failure Without Affecting Ventricular Hypertrophy. Hypertension. 2013;63.
- Kimball, SR. Eukaryotic initiation factor eIF2. The International Journal of Biochemistry & Cell Biology. 1999;31(1):25-9.
- Rani S, Sreenivasaiah PK, Cho C, Kim DH. Salubrinal Alleviates Pressure Overload-Induced Cardiac Hypertrophy by Inhibiting Endoplasmic Reticulum Stress Pathway. Mol Cells. 2017;40(1):66-72.
- James CC, Smyth JW. Alternative mechanisms of translation initiation: An emerging dynamic regulator of the proteome in health and disease. Life Sci. 2018;212:138-44.
- Martínez-Salas E, Piñeiro D, fernandez sanchez N. Alternative Mechanisms to Initiate Translation in Eukaryotic mRNAs. Comparative and functional genomics. 2012;2012:391546.
- Basheer WA, Xiao S, Epifantseva I, Fu Y, Kleber AG, Hong T, et al. GJA1-20k Arranges Actin to Guide Cx43 Delivery to Cardiac Intercalated Discs. Circulation research. 2017;121(9):1069-80.
- Peters Nicholas S, Coromilas J, Severs Nicholas J, Wit Andrew L. Disturbed Connexin43 Gap Junction Distribution Correlates With the Location of Reentrant Circuits in the Epicardial Border Zone of Healing Canine Infarcts That Cause Ventricular Tachycardia. Circulation. 1997;95(4):988-96.
- Smyth JW, Shaw RM. Autoregulation of connexin43 gap junction formation by internally translated isoforms. Cell Rep. 2013;5(3):611-8.
- Ul-Hussain M, Olk S, Schoenebeck B, Wasielewski B, Meier C, Prochnow N, et al. Internal ribosomal entry site (IRES) activity generates endogenous carboxyl-terminal domains of Cx43 and is responsive to hypoxic conditions. The Journal of biological chemistry. 2014;289(30):20979-90.
- Schiavi A, Hudder A, Werner R. Connexin43 mRNA contains a functional internal ribosome entry site. FEBS Letters. 1999;464(3):118-22.
- Salat-Canela C, Sesé M, Peula C, Ramón y Cajal S, Aasen T. Internal translation of the connexin 43 transcript. Cell Communication and Signaling. 2014;12(1):31.
- Knight JRP, Garland G, Pöyry T, Mead E, Vlahov N, Sfakianos A, et al. Control of translation elongation in health and disease. Disease Models & Mechanisms. 2020;13(3):dmm043208.
- Voorhees RM, Ramakrishnan V. Structural Basis of the Translational Elongation Cycle. Annual Review of Biochemistry. 2013;82(1):203-36.
- Hannan R, Jenkins A, Jenkins A, Brandenburger Y. Cardiac hypertrophy: A matter of translation. Clinical and Experimental Pharmacology and Physiology. 2003;30(8):517-27.
- Faller WJ, Jackson TJ, Knight JRP, Ridgway RA, Jamieson T, Karim SA, et al. mTORC1-mediated translational elongation limits intestinal tumour initiation and growth. Nature. 2015;517(7535):497-500.
- Delaidelli A, Jan A, Herms J, Sorensen PH. Translational control in brain pathologies: biological significance and therapeutic opportunities. Acta Neuropathologica. 2019;137(4):535-55.
- Kameshima S, Okada M, Ikeda S, Watanabe Y, Yamawaki H. Coordination of changes in expression and phosphorylation of eukaryotic elongation factor 2 (eEF2) and eEF2 kinase in hypertrophied cardiomyocytes. Biochemistry and Biophysics Reports. 2016;7.
- Everett AD, Stoops TD, Nairn AC, Brautigan D. Angiotensin II regulates phosphorylation of translation elongation factor-2 in cardiac myocytes. American Journal of Physiology-Heart and Circulatory Physiology. 2001;281(1):H161-H7.
- Wang L, Proud CG. Regulation of the phosphorylation of elongation factor 2 by MEK-dependent signalling in adult rat cardiomyocytes. FEBS Letters. 2002;531(2):285-9.
- McLeod LE, Wang L, Proud CG. beta-Adrenergic agonists increase phosphorylation of elongation factor 2 in cardiomyocytes without eliciting calcium-independent eEF2 kinase activity. FEBS Letters. 2001;489(2-3):225-8.
- Kaul G, Pattan G, Rafeequi T. Eukaryotic elongation factor-2 (eEF2): its regulation and peptide chain elongation. Cell Biochemistry and Function. 2011;29(3):227-34.
- Grund A, Szaroszyk M, Korf-Klingebiel M, Malek Mohammadi M, Trogisch F, Schrameck U, et al. TIP 30 counteracts cardiac hypertrophy and failure by inhibiting translational elongation. EMBO Molecular Medicine. 2019;11.
- Pittman Y, Kandl K, Lewis M, Valente L, Kinzy T. Coordination of Eukaryotic Translation Elongation Factor 1A (eEF1A) Function in Actin Organization and Translation Elongation by the Guanine Nucleotide Exchange Factor eEF1B. The Journal of biological chemistry. 2009;284:4739-47.
- He H, Chen M, Scheffler NK, Gibson Bradford W, Spremulli Linda L, Gottlieb Roberta A. Phosphorylation of Mitochondrial Elongation Factor Tu in Ischemic Myocardium. Circulation Research. 2001;89(5):461-7.
- Borutaite V, Mildaziene V, Brown GC, Brand MD. Control and kinetic analysis of ischemia-damaged heart mitochondria: which parts of the oxidative phosphorylation system are affected by ischemia? Biochimica et Biophysica Acta (BBA) - Molecular Basis of Disease. 1995;1272(3):154-8.
- Cai YC, Bullard J, Thompson N, Spremulli L. Interaction of Mitochondrial Elongation Factor Tu with Aminoacyl-tRNA and Elongation Factor Ts. The Journal of biological chemistry. 2000;275:20308-14.
- Lippmann C, Lindschau C, Buchner K, Erdmann VA, editors. Phosphorylation of Elongation Factor Tu in Vitro and in Vivo1991; Berlin, Heidelberg: Springer Berlin Heidelberg.
- Brandenburger Y, Jenkins A, Autelitano DJ, Hannan RD. Increased expression of UBF is a critical determinant for rRNA synthesis and hypertrophic growth of cardiac myocytes. The FASEB Journal. 2001;15(11):2051-3.
- Hannan RD, Luyken J, Rothblum LI. Regulation of rDNA Transcription Factors during Cardiomyocyte Hypertrophy Induced by Adrenergic Agents. Journal of Biological Chemistry. 1995;270(14):8290-7.
- Hannan RD, Luyken J, Rothblum LI. Regulation of Ribosomal DNA Transcription during Contraction-induced Hypertrophy of Neonatal Cardiomyocytes. Journal of Biological Chemistry. 1996;271(6):3213-20.
- Luyken J, Hannan RD, Cheung JY, Rothblum LI. Regulation of rDNA Transcription During Endothelin-1Induced Hypertrophy of Neonatal Cardiomyocytes. Circulation Research. 1996;78(3):354-61.
- Sadoshima J, Izumo S. Rapamycin Selectively Inhibits Angiotensin II–Induced Increase in Protein Synthesis in Cardiac Myocytes In Vitro. Circulation Research. 1995;77(6):1040-52.
- Oh H, Fujio Y, Kunisada K, Hirota H, Matsui H, Kishimoto T, et al. Activation of Phosphatidylinositol 3-Kinase through Glycoprotein 130 Induces Protein Kinase B and p70 S6 Kinase Phosphorylation in Cardiac Myocytes. Journal of Biological Chemistry. 1998;273(16):9703-10.
- Chothani S, Schafer S, Adami E, Viswanathan S, Widjaja AA, Langley SR, et al. Widespread Translational Control of Fibrosis in the Human Heart by RNA-Binding Proteins. Circulation. 2019;140(11):937-51.
- Volkers M, Toko H, Doroudgar S, Din S, Quijada P, Joyo AY, et al. Pathological hypertrophy amelioration by PRAS40-mediated inhibition of mTORC1. Proc Natl Acad Sci U S A. 2013;110(31):12661-6.
- Ma XM, Blenis J. Molecular mechanisms of mTOR-mediated translational control. Nat Rev Mol Cell Biol. 2009;10(5):307-18.
- Ben-Sahra I, Howell JJ, Asara JM, Manning BD. Stimulation of de novo pyrimidine synthesis by growth signaling through mTOR and S6K1. Science. 2013;339(6125):1323-8.
- Robitaille AM, Christen S, Shimobayashi M, Cornu M, Fava LL, Moes S, et al. Quantitative phosphoproteomics reveal mTORC1 activates de novo pyrimidine synthesis. Science. 2013;339(6125):1320-3.
- Hannan KM, Brandenburger Y, Jenkins A, Sharkey K, Cavanaugh A, Rothblum L, et al. mTOR-dependent regulation of ribosomal gene transcription requires S6K1 and is mediated by phosphorylation of the carboxy-terminal activation domain of the nucleolar transcription factor UBF. Mol Cell Biol. 2003;23(23):8862-77.
- Jackson RJ, Hellen CU, Pestova TV. The mechanism of eukaryotic translation initiation and principles of its regulation. Nat Rev Mol Cell Biol. 2010;11(2):113-27.
- Zhu Y, Pires KM, Whitehead KJ, Olsen CD, Wayment B, Zhang YC, et al. Mechanistic target of rapamycin (Mtor) is essential for murine embryonic heart development and growth. PLoS One. 2013;8(1):e54221.
- Mazelin L, Panthu B, Nicot AS, Belotti E, Tintignac L, Teixeira G, et al. mTOR inactivation in myocardium from infant mice rapidly leads to dilated cardiomyopathy due to translation defects and p53/JNK-mediated apoptosis. J Mol Cell Cardiol. 2016;97:213-25.
- Shende P, Plaisance I, Morandi C, Pellieux C, Berthonneche C, Zorzato F, et al. Cardiac raptor ablation impairs adaptive hypertrophy, alters metabolic gene expression, and causes heart failure in mice. Circulation. 2011;123(10):1073-82.
- Zhang D, Contu R, Latronico MV, Zhang J, Rizzi R, Catalucci D, et al. MTORC1 regulates cardiac function and myocyte survival through 4E-BP1 inhibition in mice. J Clin Invest. 2010;120(8):2805-16.
- Shioi T, McMullen JR, Tarnavski O, Converso K, Sherwood MC, Manning WJ, et al. Rapamycin attenuates load-induced cardiac hypertrophy in mice. Circulation. 2003;107(12):1664-70.
- McMullen JR, Sherwood MC, Tarnavski O, Zhang L, Dorfman AL, Shioi T, et al. Inhibition of mTOR signaling with rapamycin regresses established cardiac hypertrophy induced by pressure overload. Circulation. 2004;109(24):3050-5.
- Wu X, Cao Y, Nie J, Liu H, Lu S, Hu X, et al. Genetic and pharmacological inhibition of Rheb1-mTORC1 signaling exerts cardioprotection against adverse cardiac remodeling in mice. Am J Pathol. 2013;182(6):2005-14.
- Varma E, Burghaus J, Schwarzl T, Sekaran T, Gupta P, Gorska AA, et al. Translational control of Ybx1 expression regulates cardiac function in response to pressure overload in vivo. Basic Res Cardiol. 2023;118(1):25.
- Kmietczyk V, Oelschlager J, Gupta P, Varma E, Hartl S, Furkel J, et al. Ythdf2 regulates cardiac remodeling through its mRNA target transcripts. J Mol Cell Cardiol. 2023;181:57-66.
- Berulava T, Buchholz E, Elerdashvili V, Pena T, Islam MR, Lbik D, et al. Changes in m6A RNA methylation contribute to heart failure progression by modulating translation. Eur J Heart Fail. 2020;22(1):54-66.
- Rockey DC, Bell PD, Hill JA. Fibrosis--a common pathway to organ injury and failure. N Engl J Med. 2015;372(12):1138-49.
- Davis J, Molkentin JD. Myofibroblasts: Trust your heart and let fate decide. Journal of Molecular and Cellular Cardiology. 2014;70:9-18.
- Akhurst RJ, Hata A. Targeting the TGFβ signalling pathway in disease. Nature Reviews Drug Discovery. 2012;11(10):790-811.
- Obana M, Maeda M, Takeda K, Hayama A, Mohri T, Yamashita T, et al. Therapeutic Activation of Signal Transducer and Activator of Transcription 3 by Interleukin-11 Ameliorates Cardiac Fibrosis After Myocardial Infarction. Circulation. 2010;121(5):684-91.
- Widjaja AA, Lim WW, Viswanathan S, Chothani S, Corden B, Dasan CM, et al. Inhibition of IL-11 signalling extends mammalian healthspan and lifespan. Nature. 2024;632(8023):157-65.
- Schneider-Lunitz V, Ruiz-Orera J, Hubner N, van Heesch S. Multifunctional RNA-binding proteins influence mRNA abundance and translational efficiency of distinct sets of target genes. PLoS Comput Biol. 2021;17(12):e1009658.
- Doroudgar S, Hofmann C, Boileau E, Malone B, Riechert E, Gorska AA, et al. Monitoring Cell-Type-Specific Gene Expression Using Ribosome Profiling In Vivo During Cardiac Hemodynamic Stress. Circ Res. 2019;125(4):431-48.
- Yao P, Fox PL. Aminoacyl-tRNA synthetases in medicine and disease. EMBO Mol Med. 2013;5(3):332-43.
- Sampath P, Mazumder B, Seshadri V, Gerber CA, Chavatte L, Kinter M, et al. Noncanonical function of glutamyl-prolyl-tRNA synthetase: gene-specific silencing of translation. Cell. 2004;119(2):195-208.
- Ray PS, Jia J, Yao P, Majumder M, Hatzoglou M, Fox PL. A stress-responsive RNA switch regulates VEGFA expression. Nature. 2009;457(7231):915-9.
- Yao P, Potdar AA, Ray PS, Eswarappa SM, Flagg AC, Willard B, et al. The HILDA complex coordinates a conditional switch in the 3'-untranslated region of the VEGFA mRNA. PLoS Biol. 2013;11(8):e1001635.
- Mendes MI, Gutierrez Salazar M, Guerrero K, Thiffault I, Salomons GS, Gauquelin L, et al. Bi-allelic Mutations in EPRS, Encoding the Glutamyl-Prolyl-Aminoacyl-tRNA Synthetase, Cause a Hypomyelinating Leukodystrophy. Am J Hum Genet. 2018;102(4):676-84.
- Da M, Feng Y, Xu J, Hu Y, Lin Y, Ni B, et al. Association of aminoacyl-tRNA synthetases gene polymorphisms with the risk of congenital heart disease in the Chinese Han population. PLoS One. 2014;9(10):e110072.
- Rau CD, Romay MC, Tuteryan M, Wang JJ, Santolini M, Ren S, et al. Systems Genetics Approach Identifies Gene Pathways and Adamts2 as Drivers of Isoproterenol-Induced Cardiac Hypertrophy and Cardiomyopathy in Mice. Cell Syst. 2017;4(1):121-8 e4.
- Galindo CL, Skinner MA, Errami M, Olson LD, Watson DA, Li J, et al. Transcriptional profile of isoproterenol-induced cardiomyopathy and comparison to exercise-induced cardiac hypertrophy and human cardiac failure. BMC physiology. 2009;9:23.
- Harigaya Y, Parker R. No-go decay: a quality control mechanism for RNA in translation. Wiley Interdiscip Rev RNA. 2010;1(1):132-41.
- Wu J, Hollinger J, Bonanno E, Jiang F, Yao P. Cardiomyocyte-Specific Loss of Glutamyl-prolyl-tRNA Synthetase Leads to Disturbed Protein Homeostasis and Dilated Cardiomyopathy. Cells. 2023;13(1).
- Doerfel LK, Wohlgemuth I, Kothe C, Peske F, Urlaub H, Rodnina MV. EF-P is essential for rapid synthesis of proteins containing consecutive proline residues. Science. 2013;339(6115):85-8.
- Schuller AP, Wu CC, Dever TE, Buskirk AR, Green R. eIF5A Functions Globally in Translation Elongation and Termination. Mol Cell. 2017;66(2):194-205 e5.
- Lassak J, Wilson DN, Jung K. Stall no more at polyproline stretches with the translation elongation factors EF-P and IF-5A. Mol Microbiol. 2016;99(2):219-35.
- Nakanishi S, Cleveland JL. Targeting the polyamine-hypusine circuit for the prevention and treatment of cancer. Amino Acids. 2016;48(10):2353-62.
- Imam S, Mirmira RG, Jaume JC. Eukaryotic translation initiation factor 5A inhibition alters physiopathology and immune responses in a "humanized" transgenic mouse model of type 1 diabetes. Am J Physiol Endocrinol Metab. 2014;306(7):E791-8.
- Subbaiah KCV, Wu J, Tang WHW, Yao P. Ciclopirox Inhibition of eIF5A Hypusination Attenuates Fibroblast Activation and Cardiac Fibrosis. J Cardiovasc Dev Dis. 2023;10(2).
- Goldfarb L, Dalakas M. Tragedy in a heartbeat: malfunctioning desmin causes skeletal and cardiac muscle disease. The Journal of Clinical Investigation. 2011;121:455-.
- McLendon P, Robbins J. Desmin-related cardiomyopathy: An unfolding story. American journal of physiology Heart and circulatory physiology. 2011;301:H1220-8.
- Park SG, Schimmel P, Kim S. Aminoacyl tRNA synthetases and their connections to disease. Proceedings of the National Academy of Sciences. 2008;105(32):11043.
- Liu Y, Satz J, Vo M-N, Nangle L, Schimmel P, Ackerman S. Deficiencies in tRNA synthetase editing activity cause cardioproteinopathy. Proc Natl Acad Sci U S A. 2014;111.
- Ling J, Reynolds N, Ibba M. Aminoacyl-tRNA Synthesis and Translational Quality Control. Annual review of microbiology. 2009;63:61-78.
- Yao P, Zhu B, Jaeger S, Eriani G, Wang ED. Recognition of tRNALeu by Aquifex aeolicus leucyl-tRNA synthetase during the aminoacylation and editing steps. Nucleic Acids Res. 2008;36(8):2728-38.
- Beebe K, Pouplana L, Schimmel P. Elucidation of tRNA-dependent editing by a class II tRNA synthetase and significance for cell viability. The EMBO journal. 2003;22:668-75.
- Lee j-w, Beebe K, Nangle L, Jang J, Guess C, Cook S, et al. Editing-defective tRNA synthetase causes protein misfolding and neurodegeneration. Nature. 2006;443:50-5.
- Nangle L, Motta C, Schimmel P. Global Effects of Mistranslation from an Editing Defect in Mammalian Cells. Chemistry & biology. 2006;13:1091-100.
- Huang Q, Yao P, Eriani G, Wang ED. In vivo identification of essential nucleotides in tRNALeu to its functions by using a constructed yeast tRNALeu knockout strain. Nucleic Acids Res. 2012;40(20):10463-77.
- Zuhlke V, Du Mesnil de R, Gudbjarnason S, Bing RJ. Inhibition of protein synthesis in cardiac hypertrophy and its relation to myocardial failure. Circ Res. 1966;18(5):558-72.
- Keller TL, Zocco D, Sundrud MS, Hendrick M, Edenius M, Yum J, et al. Halofuginone and other febrifugine derivatives inhibit prolyl-tRNA synthetase. Nat Chem Biol. 2012;8(3):311-7.
- Zhou H, Sun L, Yang XL, Schimmel P. ATP-directed capture of bioactive herbal-based medicine on human tRNA synthetase. Nature. 2013;494(7435):121-4.
- Qin P, Arabacilar P, Bernard RE, Bao W, Olzinski AR, Guo Y, et al. Activation of the Amino Acid Response Pathway Blunts the Effects of Cardiac Stress. J Am Heart Assoc. 2017;6(5).
- Sundrud MS, Koralov SB, Feuerer M, Calado DP, Kozhaya AE, Rhule-Smith A, et al. Halofuginone inhibits TH17 cell differentiation by activating the amino acid starvation response. Science. 2009;324(5932):1334-8.
- Zhang T, Wu Y, Hu Z, Xing W, Kun LV, Wang D, et al. Small-Molecule Integrated Stress Response Inhibitor Reduces Susceptibility to Postinfarct Atrial Fibrillation in Rats via the Inhibition of Integrated Stress Responses. J Pharmacol Exp Ther. 2021;378(3):197-206.
- Onat UI, Yildirim AD, Tufanli O, Cimen I, Kocaturk B, Veli Z, et al. Intercepting the Lipid-Induced Integrated Stress Response Reduces Atherosclerosis. J Am Coll Cardiol. 2019;73(10):1149-69.
- Hedaya OM, Jiang F, Baliga U, Ivanov A, Chen S, Schwartz JL, et al. Upstream open reading frame inactivation augments GATA4 translation and cardiomyocyte hypertrophy in mice. bioRxiv. 2025.
- Xiang Y, Huang W, Tan L, Chen T, He Y, Irving PS, et al. Pervasive downstream RNA hairpins dynamically dictate start-codon selection. Nature. 2023;621(7978):423-30.
- Guenther UP, Weinberg DE, Zubradt MM, Tedeschi FA, Stawicki BN, Zagore LL, et al. The helicase Ded1p controls use of near-cognate translation initiation codons in 5' UTRs. Nature. 2018;559(7712):130-4.
- Cheng W, Wang S, Zhang Z, Morgens DW, Hayes LR, Lee S, et al. CRISPR-Cas9 Screens Identify the RNA Helicase DDX3X as a Repressor of C9ORF72 (GGGGCC)n Repeat-Associated Non-AUG Translation. Neuron. 2019;104(5):885-98 e8.
- Aghajanian H, Kimura T, Rurik JG, Hancock AS, Leibowitz MS, Li L, et al. Targeting cardiac fibrosis with engineered T cells. Nature. 2019;573(7774):430-3.
- Rurik JG, Tombacz I, Yadegari A, Mendez Fernandez PO, Shewale SV, Li L, et al. CAR T cells produced in vivo to treat cardiac injury. Science. 2022;375(6576):91-6.
- Chen H, Liu D, Aditham A, Guo J, Huang J, Kostas F, et al. Chemical and topological design of multicapped mRNA and capped circular RNA to augment translation. Nat Biotechnol. 2024.
- Anttila V, Saraste A, Knuuti J, Hedman M, Jaakkola P, Laugwitz KL, et al. Direct intramyocardial injection of VEGF mRNA in patients undergoing coronary artery bypass grafting. Mol Ther. 2023;31(3):866-74.
- Liang XH, Shen W, Sun H, Migawa MT, Vickers TA, Crooke ST. Translation efficiency of mRNAs is increased by antisense oligonucleotides targeting upstream open reading frames. Nat Biotechnol. 2016;34(8):875-80.
- Chen Q, Yu T, Gong J, Shan H. Advanced Nanomedicine Delivery Systems for Cardiovascular Diseases: Viral and Non-Viral Strategies in Targeted Therapy. Molecules. 2025;30(4).
- Philippou S, Mastroyiannopoulos NP, Tomazou M, Oulas A, Ackers-Johnson M, Foo RS, et al. Selective Delivery to Cardiac Muscle Cells Using Cell-Specific Aptamers. Pharmaceuticals (Basel). 2023;16(9).
- Narayan C, Lin LH, Barros MN, Gilbert TC, Brown CR, Reddin D, et al. Identification of In Vivo Internalizing Cardiac-Specific RNA Aptamers. bioRxiv. 2024.
- Albers S, Allen EC, Bharti N, Davyt M, Joshi D, Perez-Garcia CG, et al. Engineered tRNAs suppress nonsense mutations in cells and in vivo. Nature. 2023;618(7966):842-8.
- Chen R, Wang SK, Belk JA, Amaya L, Li Z, Cardenas A, et al. Engineering circular RNA for enhanced protein production. Nat Biotechnol. 2023;41(2):262-72.
- Carrieri C, Cimatti L, Biagioli M, Beugnet A, Zucchelli S, Fedele S, et al. Long non-coding antisense RNA controls Uchl1 translation through an embedded SINEB2 repeat. Nature. 2012;491(7424):454-7.
- Arnoldi M, Zarantonello G, Espinoza S, Gustincich S, Di Leva F, Biagioli M. Design and Delivery of SINEUP: A New Modular Tool to Increase Protein Translation. Methods Mol Biol. 2022;2434:63-87.
- Ford PW, Narasimhan M, Bennett EJ. Ubiquitin-dependent translation control mechanisms: Degradation and beyond. Cell Rep. 2024;43(12):115050.
- Santos-Ribeiro D, Godinas L, Pilette C, Perros F. The integrated stress response system in cardiovascular disease. Drug Discov Today. 2018;23(4):920-9.
- Sciarretta S, Forte M, Frati G, Sadoshima J. New Insights Into the Role of mTOR Signaling in the Cardiovascular System. Circ Res. 2018;122(3):489-505.
- McGirr T, Onar O, Jafarnejad SM. Dysregulated ribosome quality control in human diseases. FEBS J. 2025;292(5):936-59.
- Ford PW, Garshott DM, Narasimhan M, Ge X, Jordahl EM, Subramanya S, et al. RNF10 and RIOK3 facilitate 40S ribosomal subunit degradation upon 60S biogenesis disruption or amino acid starvation. Cell Rep. 2025;44(3):115371.
- Garzia A, Meyer C, Tuschl T. The E3 ubiquitin ligase RNF10 modifies 40S ribosomal subunits of ribosomes compromised in translation. Cell Rep. 2021;36(5):109468.
- Clemens, MJ. Initiation factor eIF2 alpha phosphorylation in stress responses and apoptosis. Prog Mol Subcell Biol. 2001;27:57-89.
- Vanhoutte D, Schips TG, Vo A, Grimes KM, Baldwin TA, Brody MJ, et al. Thbs1 induces lethal cardiac atrophy through PERK-ATF4 regulated autophagy. Nat Commun. 2021;12(1):3928.
- Zhang S, Tian W, Duan X, Zhang Q, Cao L, Liu C, et al. Melatonin attenuates diabetic cardiomyopathy by increasing autophagy of cardiomyocytes via regulation of VEGF-B/GRP78/PERK signaling pathway. Cardiovasc Diabetol. 2024;23(1):19.
- Li RJ, He KL, Li X, Wang LL, Liu CL, He YY. Salubrinal protects cardiomyocytes against apoptosis in a rat myocardial infarction model via suppressing the dephosphorylation of eukaryotic translation initiation factor 2alpha. Mol Med Rep. 2015;12(1):1043-9.
- Kim J, Kundu M, Viollet B, Guan KL. AMPK and mTOR regulate autophagy through direct phosphorylation of Ulk1. Nat Cell Biol. 2011;13(2):132-41.
- Li Y, Chen C, Yao F, Su Q, Liu D, Xue R, et al. AMPK inhibits cardiac hypertrophy by promoting autophagy via mTORC1. Arch Biochem Biophys. 2014;558:79-86.
- He C, Zhu H, Li H, Zou MH, Xie Z. Dissociation of Bcl-2-Beclin1 complex by activated AMPK enhances cardiac autophagy and protects against cardiomyocyte apoptosis in diabetes. Diabetes. 2013;62(4):1270-81.
- Matsui Y, Takagi H, Qu X, Abdellatif M, Sakoda H, Asano T, et al. Distinct roles of autophagy in the heart during ischemia and reperfusion: roles of AMP-activated protein kinase and Beclin 1 in mediating autophagy. Circ Res. 2007;100(6):914-22.
- Xie W, Du L. Diabetes is an inflammatory disease: evidence from traditional Chinese medicines. Diabetes Obes Metab. 2011;13(4):289-301.
- Kanamori H, Naruse G, Yoshida A, Minatoguchi S, Watanabe T, Kawaguchi T, et al. Metformin Enhances Autophagy and Provides Cardioprotection in delta-Sarcoglycan Deficiency-Induced Dilated Cardiomyopathy. Circ Heart Fail. 2019;12(4):e005418.
- Li C, Li S, Zhang G, Li Q, Song W, Wang X, et al. IRE1alpha Mediates the Hypertrophic Growth of Cardiomyocytes Through Facilitating the Formation of Initiation Complex to Promote the Translation of TOP-Motif Transcripts. Circulation. 2024;150(13):1010-29.
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



