Submitted:
17 January 2025
Posted:
17 January 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Experimental Fish
2.2. Sample Processing and Collection
2.3. Total RNA Extraction and cDNA Synthesis
2.4. Amplification of the CDS Region of the Opnod1, Opnod2 and Optbk1 Genes
2.5. Sequence Analysis of Opnod1, Opnod2 and Optbk1 Genes
2.6. Sequence Analysis of Opnod1, Opnod2 and Optbk1 Genes
2.7. In Vitro Stimulation of Kidney Cells of O. punctatus with Poly I:C and LPS
3. Results
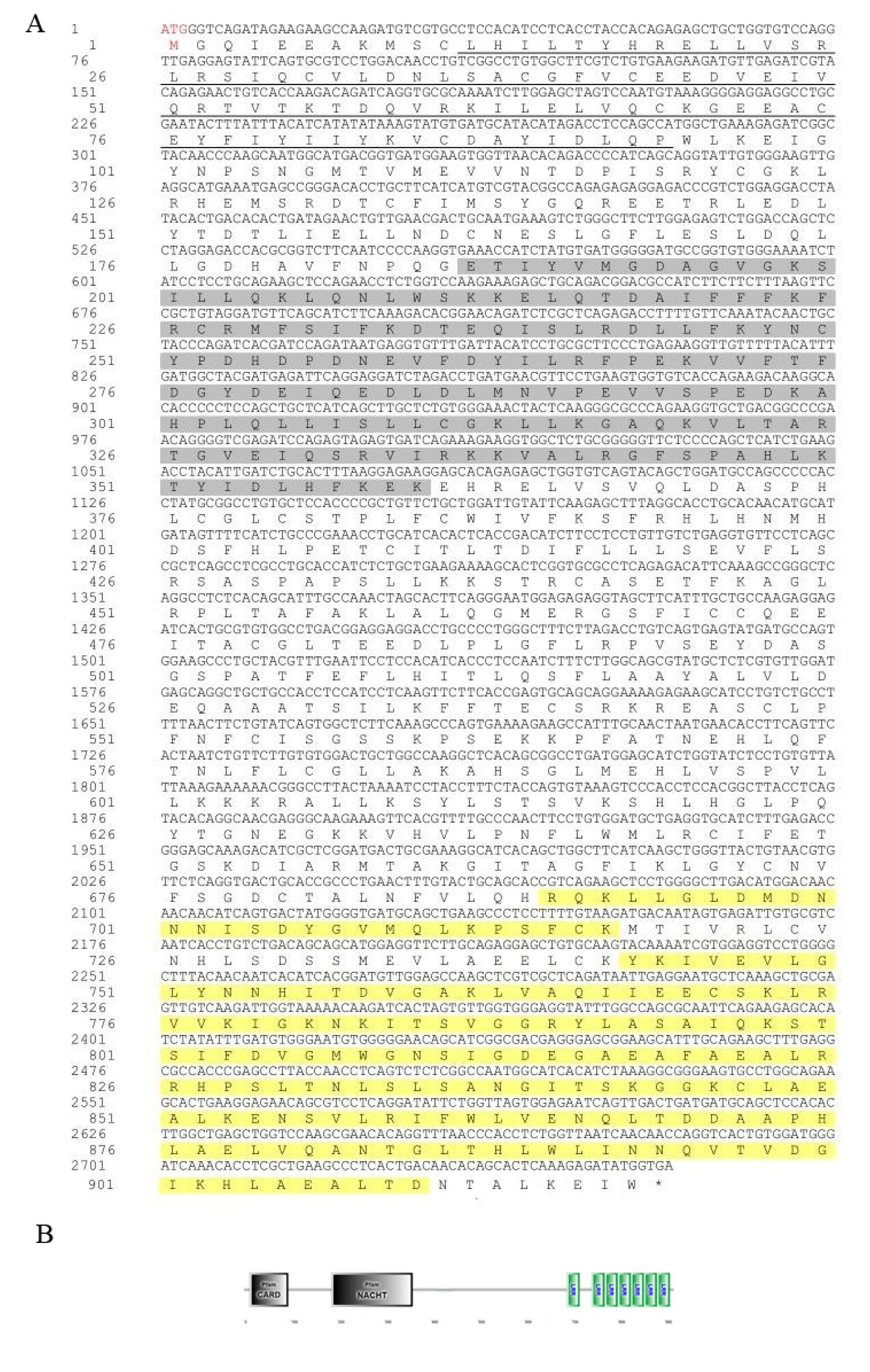
3.1. Sequence Characteristics of Opnod1, Opnod2 and Optbk1 Genes
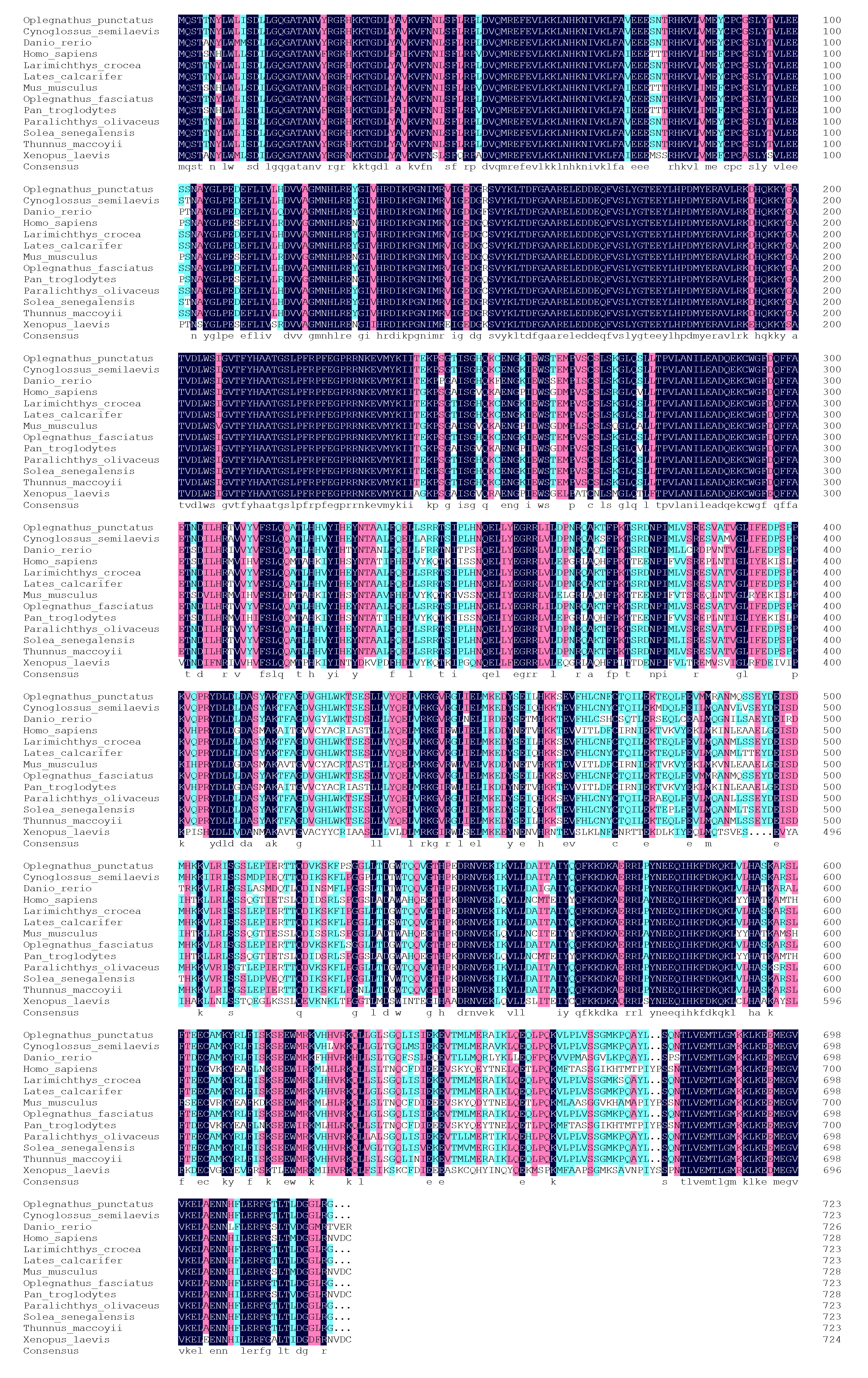
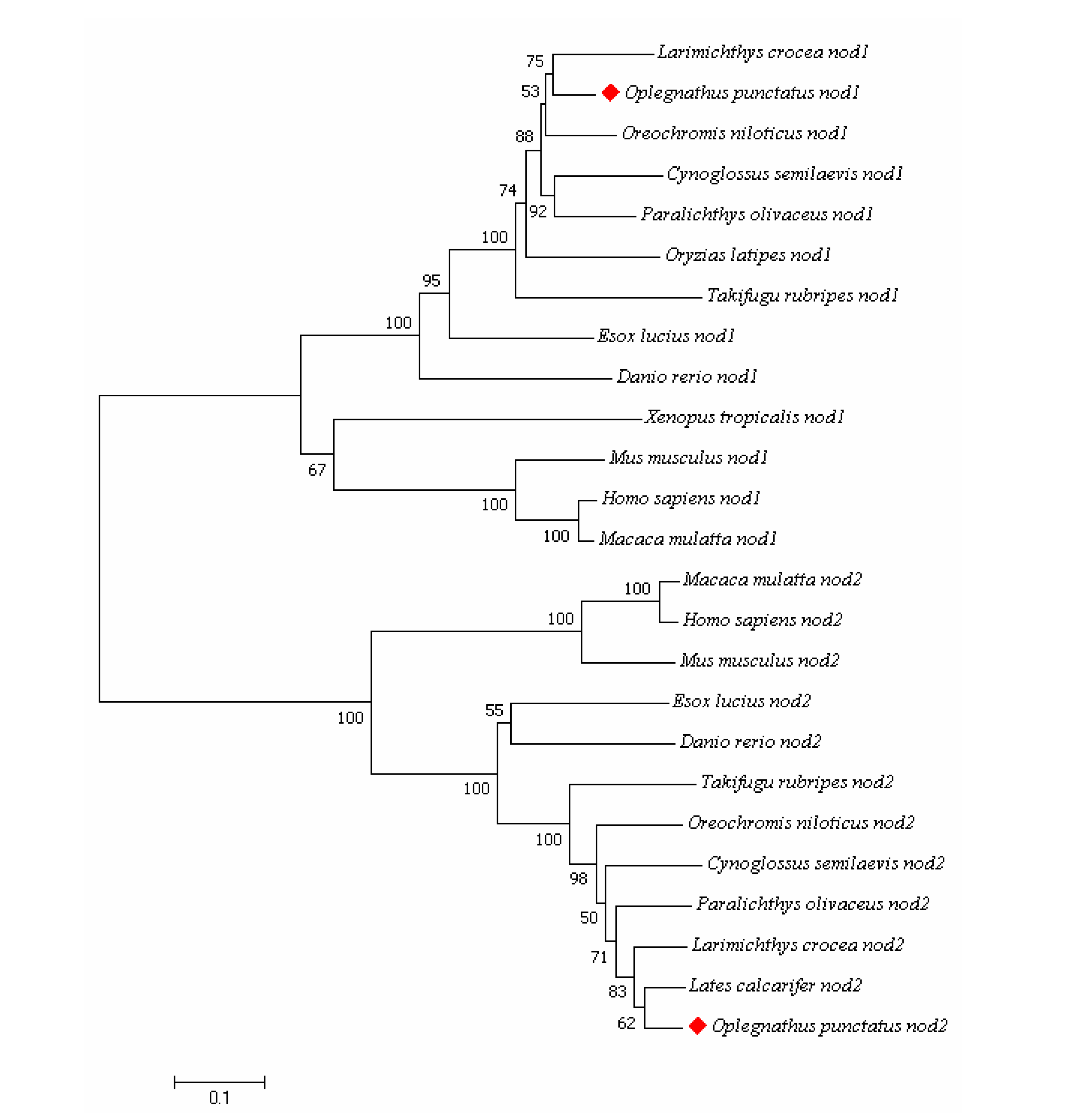
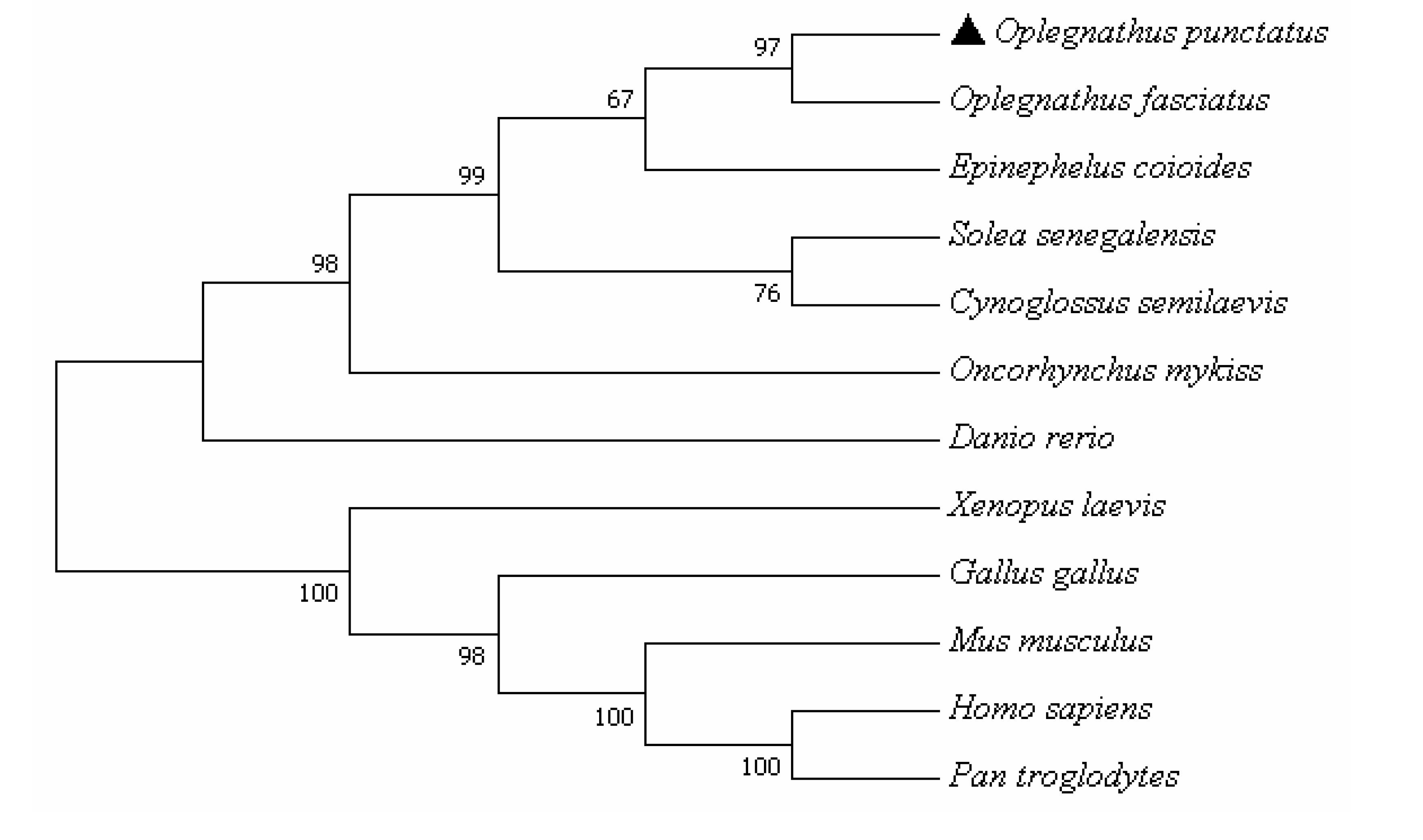
3.2. Amino Acid Multiple Sequence Alignment and Phylogenetic Tree Analysis
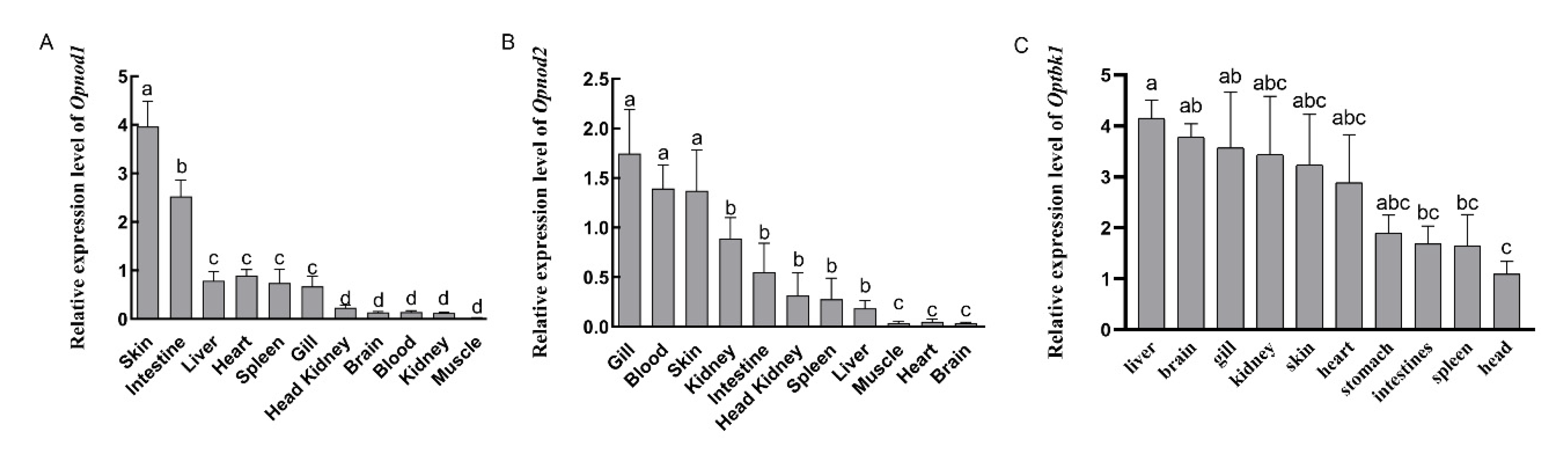
3.3. Expression Patterns of Opnod1, Opnod2 and Optbk1 Genes in Healthy Individuals
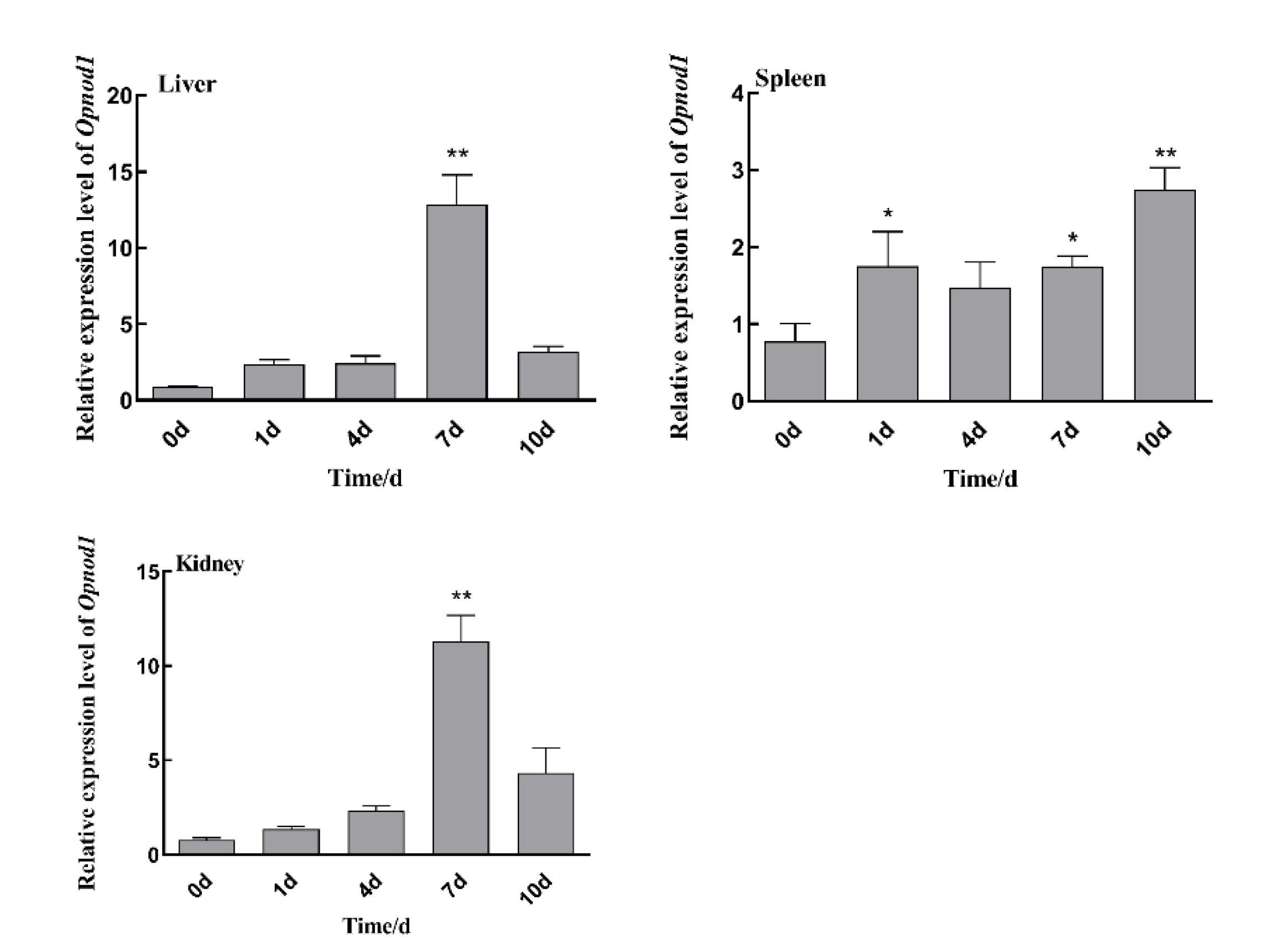
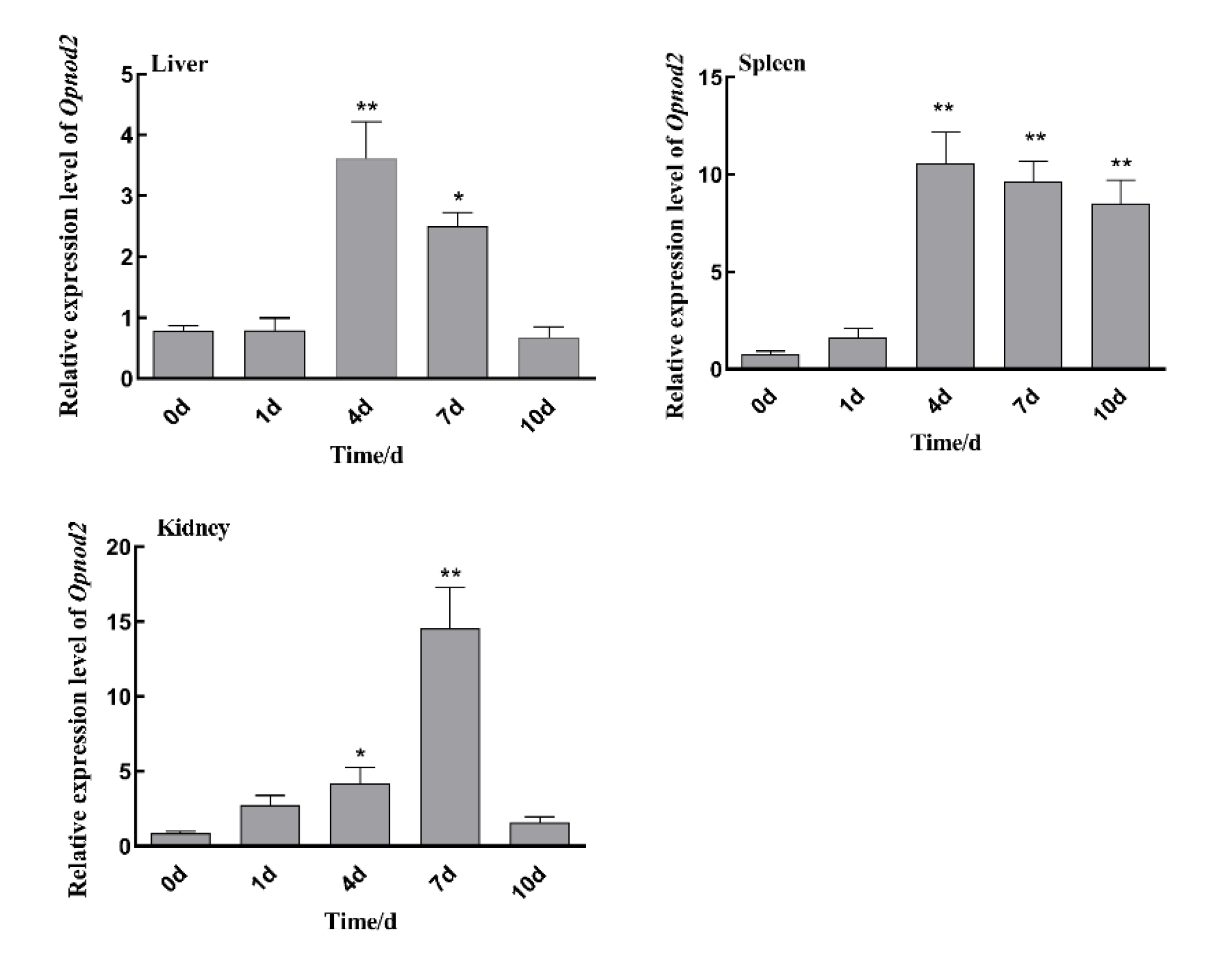
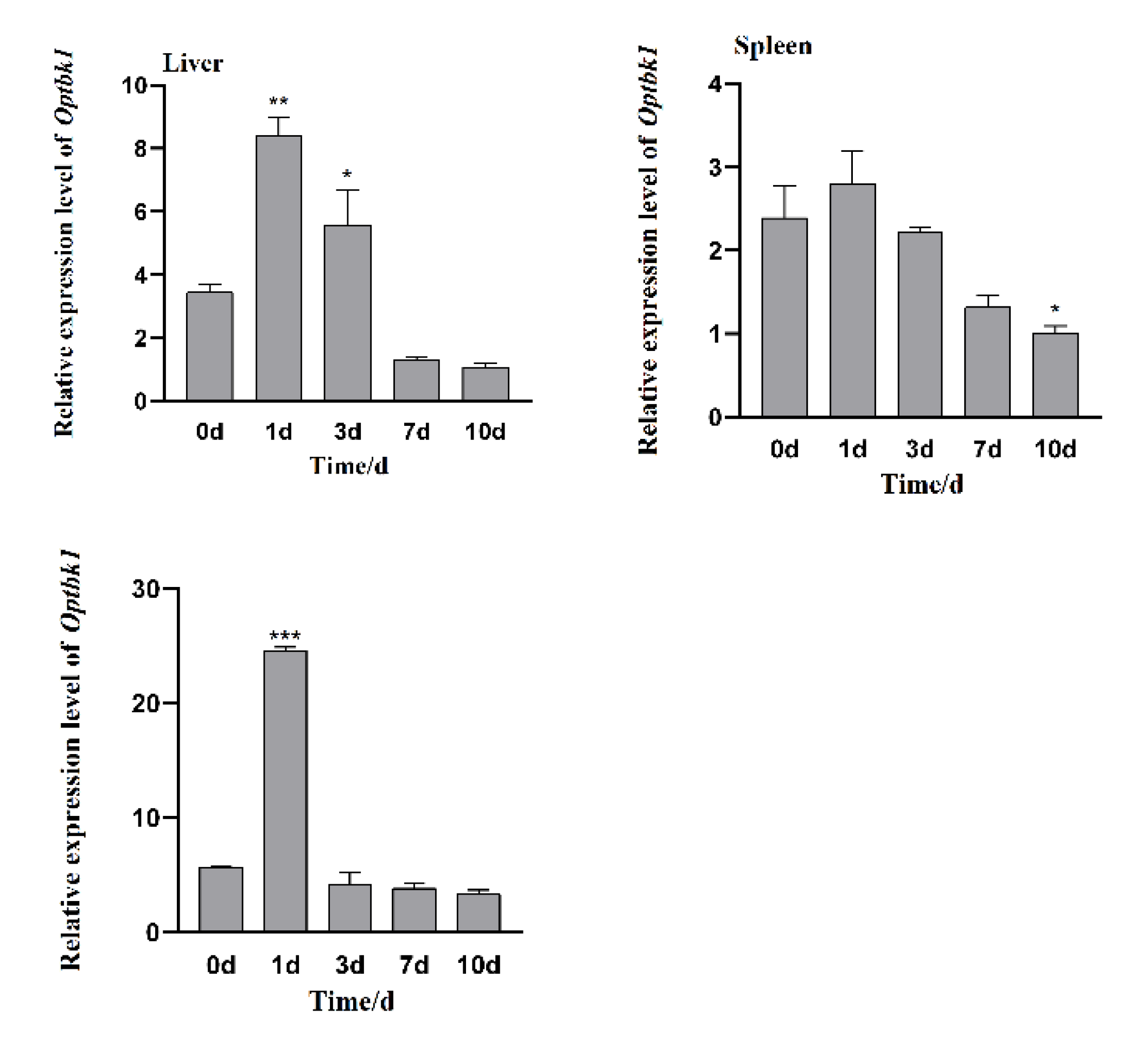
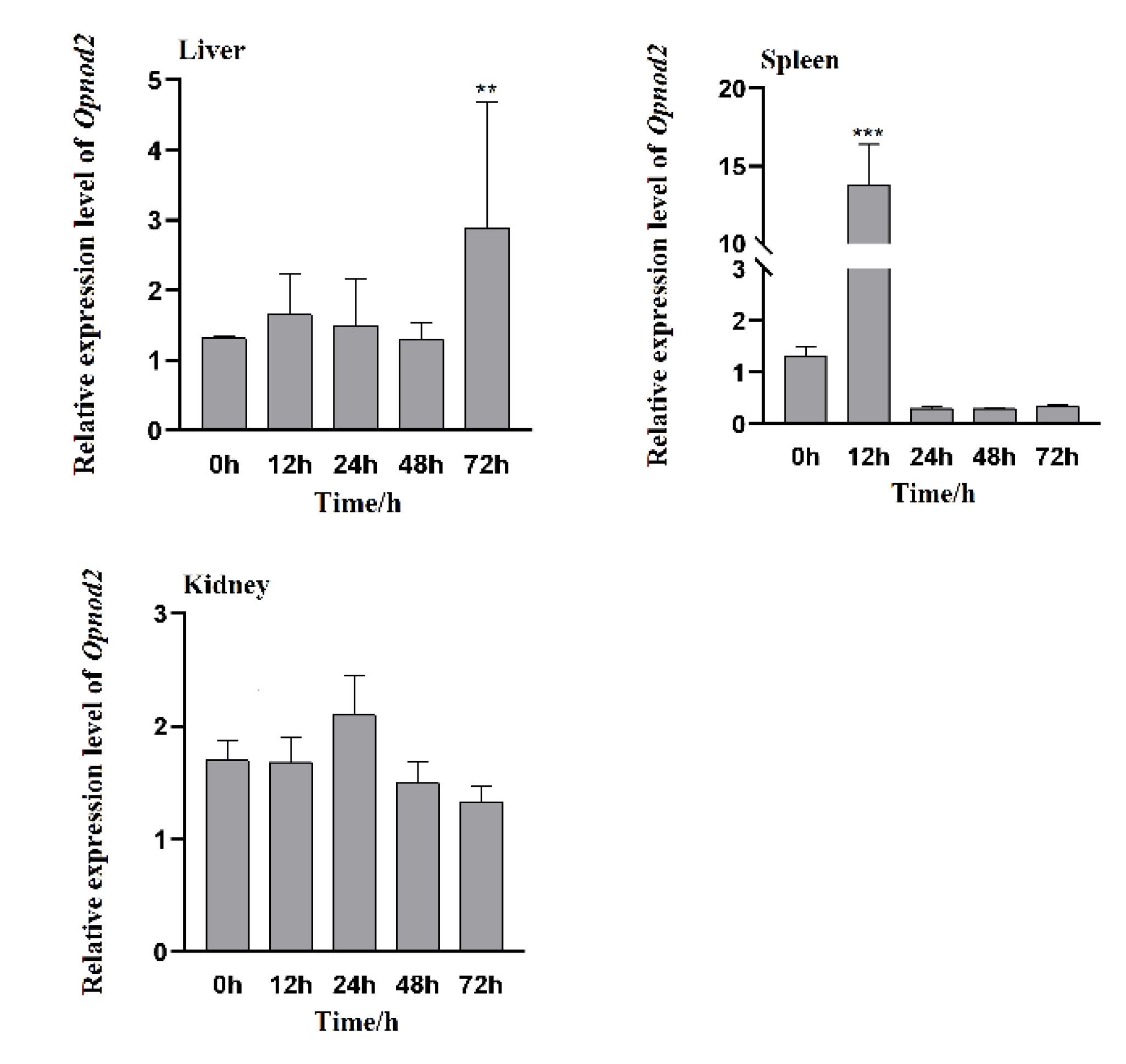
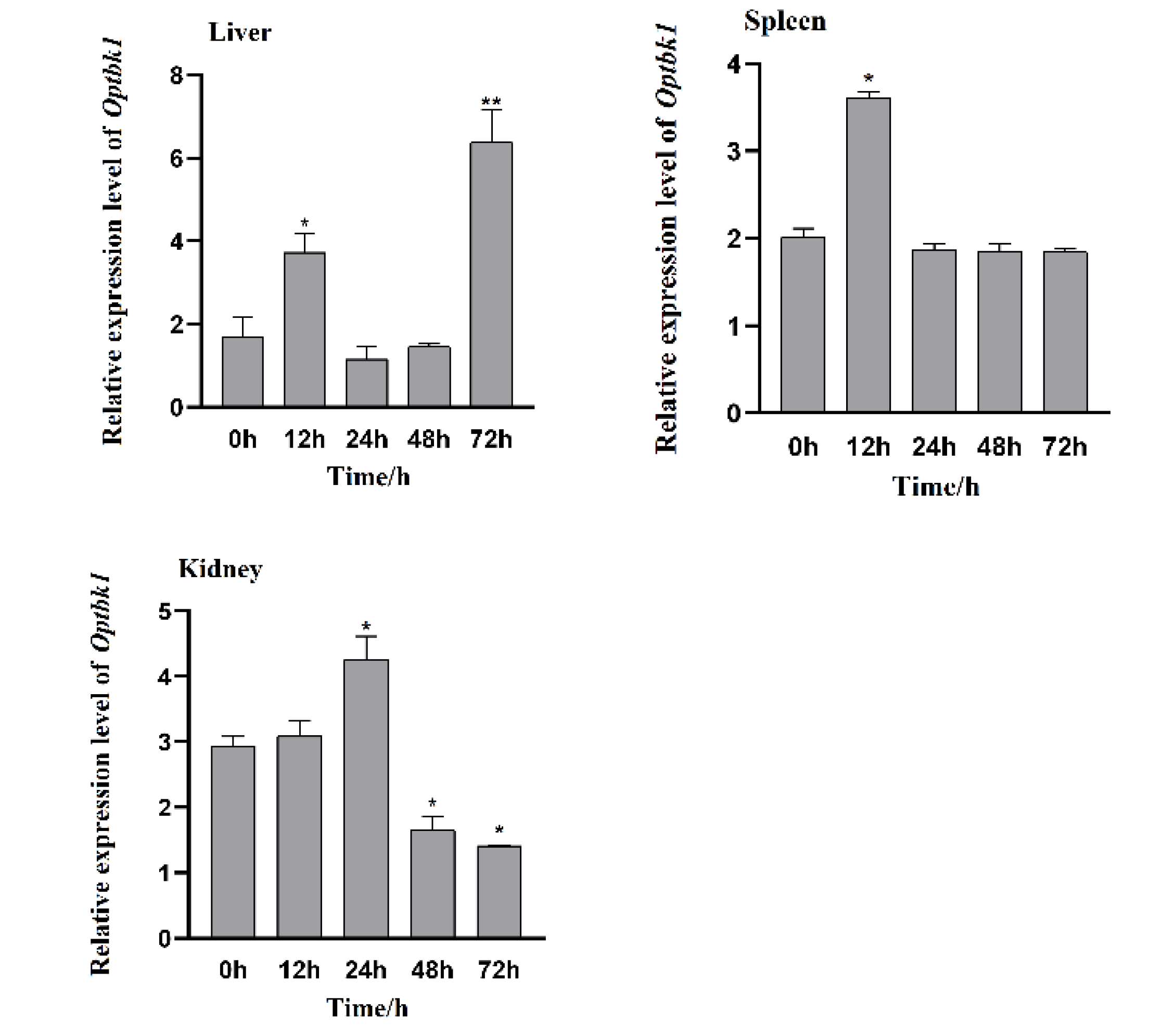
3.4. Changes in Expression of Opnod1, Opnod2 and Optbk1 Genes After Iridovirus Stimulation
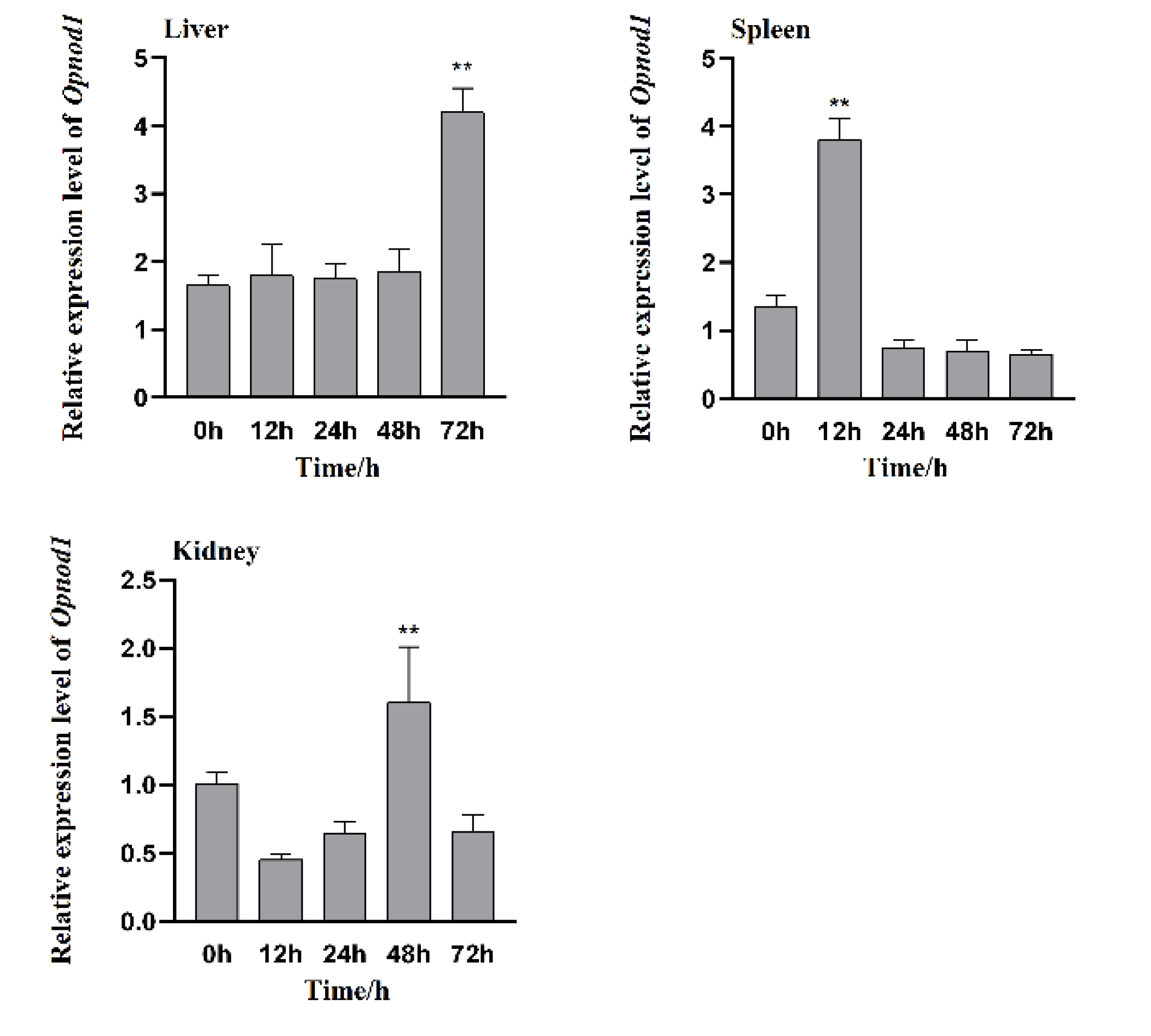
3.5. Changes in the Expression of Opnod1, Opnod2 and Optbk1 Genes After Stimulation by Vibrio harveyi
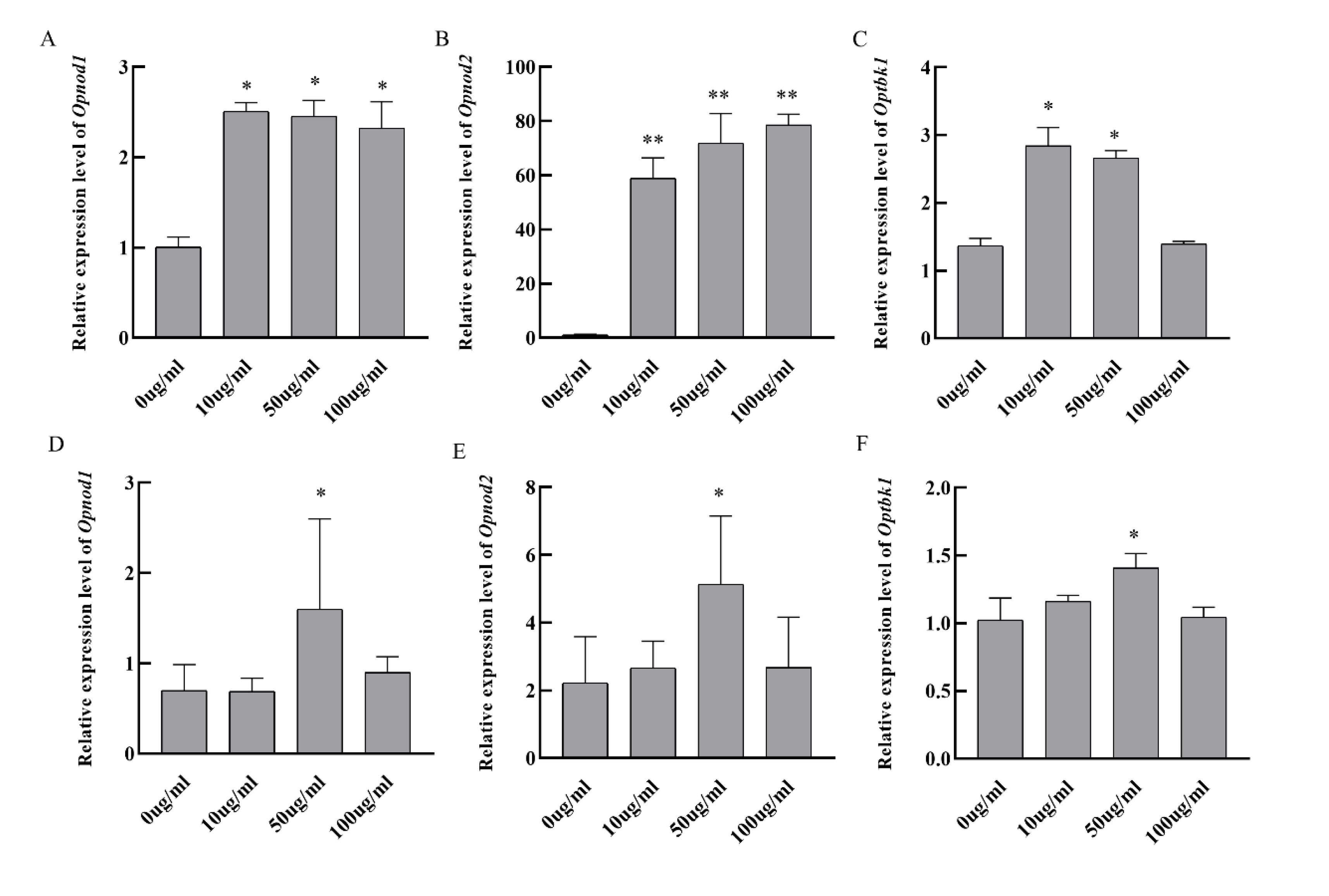
3.6. In Vitro Stimulation of Grouper Kidney Cells
4. Discussion
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Netea, MG. Training innate immunity: the changing concept of immunological memory in innate host defence. EUR J CLIN INVEST. 2013 43:881-4.
- Gomez GD, Balcazar JL. A review on the interactions between gut microbiota and innate immunity of fish. FEMS Immunol Med Microbiol. 2008 52:145-54.
- Takeuchi O, Akira S. Pattern recognition receptors and inflammation. CELL. 2010 140:805-20.
- Liao Z, Su J. Progresses on three pattern recognition receptor families (TLRs, RLRs and NLRs) in teleost. DEV COMP IMMUNOL. 2021 122:104131.
- Sahoo, BR. Structure of fish Toll-like receptors (TLR) and NOD-like receptors (NLR). INT J BIOL MACROMOL. 2020 161:1602-17.
- Rietdijk ST, Burwell T, Bertin J, Coyle AJ. Sensing intracellular pathogens-NOD-like receptors. CURR OPIN PHARMACOL. 2008 8:261-6.
- Zhang L, Gao Z, Yu L, Zhang B, Wang J, Zhou J. Nucleotide-binding and oligomerization domain (NOD)-like receptors in teleost fish: Current knowledge and future perspectives. J FISH DIS. 2018 41:1317-30.
- Sha Z, Abernathy JW, Wang S, Li P, Kucuktas H, Liu H, et al. NOD-like subfamily of the nucleotide-binding domain and leucine-rich repeat containing family receptors and their expression in channel catfish. DEV COMP IMMUNOL. 2009 33:991-9.
- Gu T, Lu L, Wang J, Tian L, Wei W, Wu X, et al. The NOD1 and NOD2 in mandarinfish (Siniperca chuatsi): molecular characterization, tissue distribution, and expression analysis. BMC GENET. 2018 19:61.
- Chamaillard M, Hashimoto M, Horie Y, Masumoto J, Qiu S, Saab L, et al. An essential role for NOD1 in host recognition of bacterial peptidoglycan containing diaminopimelic acid. NAT IMMUNOL. 2003 4:702-7.
- Chang M, Wang T, Nie P, Zou J, Secombes CJ. Cloning of two rainbow trout nucleotide-binding oligomerization domain containing 2 (NOD2) splice variants and functional characterization of the NOD2 effector domains. FISH SHELLFISH IMMUN. 2011 30:118-27.
- Motta V, Soares F, Sun T, Philpott DJ. NOD-like receptors: versatile cytosolic sentinels. PHYSIOL REV. 2015 95:149-78.
- Philpott DJ, Sorbara MT, Robertson SJ, Croitoru K, Girardin SE. NOD proteins: regulators of inflammation in health and disease. NAT REV IMMUNOL. 2014 14:9-23.
- Basu M, Paichha M, Lenka SS, Chakrabarty R, Samanta M. Hypoxic stress: impact on the modulation of TLR2, TLR4, NOD1 and NOD2 receptor and their down-stream signalling genes expression in catla (Catla catla). MOL BIOL REP. 2016 43:1-9.
- Sabbah A, Chang TH, Harnack R, Frohlich V, Tominaga K, Dube PH, et al. Activation of innate immune antiviral responses by Nod2. NAT IMMUNOL. 2009 10:1073-80.
- Chen WQ, Xu QQ, Chang MX, Nie P, Peng KM. Molecular characterization and expression analysis of nuclear oligomerization domain proteins NOD1 and NOD2 in grass carp Ctenopharyngodon idella. FISH SHELLFISH IMMUN. 2010 28:18-29.
- Li J, Gao Y, Xu T. Comparative genomic and evolution of vertebrate NOD1 and NOD2 genes and their immune response in miiuy croaker. FISH SHELLFISH IMMUN. 2015 46:387-97.
- Zou PF, Chang MX, Li Y, Xue NN, Li JH, Chen SN, et al. NOD2 in zebrafish functions in antibacterial and also antiviral responses via NF-kappaB, and also MDA5, RIG-I and MAVS. FISH SHELLFISH IMMUN. 2016 55:173-85.
- Gao FY, Pang JC, Lu MX, Yang XL, Zhu HP, Ke XL, et al. Molecular characterization, expression and functional analysis of NOD1, NOD2 and NLRC3 in Nile tilapia (Oreochromis niloticus). FISH SHELLFISH IMMUN. 2018 73:207-19.
- Langevin C, Aleksejeva E, Passoni G, Palha N, Levraud JP, Boudinot P. The antiviral innate immune response in fish: evolution and conservation of the IFN system. J MOL BIOL. 2013 425:4904-20.
- Benavides A, Gutierrez D, Epuyao N, Modak B, Imarai M, Valenzuela B. Alpinone: A positive regulator molecule of immune antiviral response in Atlantic salmon kidney cells. DEV COMP IMMUNOL. 2022 126:104262.
- Li S, Lu LF, LaPatra SE, Chen DD, Zhang YA. Zebrafish STAT6 negatively regulates IFNphi1 production by attenuating the kinase activity of TANK-binding kinase 1. DEV COMP IMMUNOL. 2017 67:189-201.
- Feng X, Su J, Yang C, Yan N, Rao Y, Chen X. Molecular characterizations of grass carp (Ctenopharyngodon idella) TBK1 gene and its roles in regulating IFN-I pathway. DEV COMP IMMUNOL. 2014 45:278-90.
- Yan C, Xiao J, Li J, Chen H, Liu J, Wang C, et al. TBK1 of black carp plays an important role in host innate immune response against SVCV and GCRV. FISH SHELLFISH IMMUN. 2017 69:108-18.
- Wu M, Zhao X, Gong XY, Wang Y, Gui JF, Zhang YB. FTRCA1, a Species-Specific Member of finTRIM Family, Negatively Regulates Fish IFN Response through Autophage-Lysosomal Degradation of TBK1. J IMMUNOL. 2019 202:2407-20.
- Dong C, Weng S, Luo Y, Huang M, Ai H, Yin Z, et al. A new marine megalocytivirus from spotted knifejaw, Oplegnathus punctatus, and its pathogenicity to freshwater mandarinfish, Siniperca chuatsi. VIRUS RES. 2010 147:98-106.
- Liao J, Kang S, Zhang L, Zhang D, Xu Z, Qin Q, et al. Isolation and identification of a megalocytivirus strain (SKIV-TJ) from cultured spotted knifejaw (Oplegnathus punctatus) in China and its pathogenicity analysis. FISH SHELLFISH IMMUN. 2023 141:109034.
- Huang X, Wei J, Zheng Q, Zhang Y, Zhu W, Liu J, et al. Isolation, identification and genomic analysis of an ISKNV-type megalocytivirus from spotted knifejaw (Oplegnathus punctatus). AQUACULTURE. 2021 532:736032.
- Wang J, Chen Z, Xu W, Li Y, Lu S, Wang L, et al. Transcriptomic analysis reveals the gene expression profiles in the spleen of spotted knifejaw (Oplegnathus punctatus) infected by Vibrio harveyi. DEV COMP IMMUNOL. 2022 133:104432.
- Hou QH, Yi SB, Ding X, Zhang HX, Sun Y, Zhang Y, et al. Differential expression analysis of nuclear oligomerization domain proteins NOD1 and NOD2 in orange-spotted grouper (Epinephelus coioides). FISH SHELLFISH IMMUN. 2012 33:1102-11.
- Laing KJ, Purcell MK, Winton JR, Hansen JD. A genomic view of the NOD-like receptor family in teleost fish: identification of a novel NLR subfamily in zebrafish. BMC EVOL BIOL. 2008 8:42.
- Swain B, Basu M, Lenka SS, Das S, Jayasankar P, Samanta M. Characterization and Inductive Expression Analysis of Interferon Gamma-Related Gene in the Indian Major Carp, Rohu (Labeo rohita). DNA CELL BIOL. 2015 34:367-78.
- Basu M, Swain B, Sahoo BR, Maiti NK, Samanta M. Induction of toll-like receptor (TLR) 2, and MyD88-dependent TLR- signaling in response to ligand stimulation and bacterial infections in the Indian major carp, mrigal (Cirrhinus mrigala). MOL BIOL REP. 2012 39:6015-28.
- Caruso R, Nunez G. Innate Immunity: ER Stress Recruits NOD1 and NOD2 for Delivery of Inflammation. CURR BIOL. 2016 26:R508-11.
- Feng H, Liu H, Kong R, Wang L, Wang Y, Hu W, et al. Expression profiles of carp IRF-3/-7 correlate with the up-regulation of RIG-I/MAVS/TRAF3/TBK1, four pivotal molecules in RIG-I signaling pathway. FISH SHELLFISH IMMUN. 2011 30:1159-69.
- Zhang DL, Yu DH, Chen J, Fan S, Wang ZY. Expression profiles and interaction suggest TBK1 can be regulated by Nrdp1 in response to immune stimulation in large yellow croaker Larimichthys crocea. FISH SHELLFISH IMMUN. 2015 46:745-52.
- Chen J, Li ZC, Lu LF, Li P, Li XY, Li S. Functional Characterization of Dark Sleeper (Odontobutis obscura) TBK1 on IFN Regulation. FRONT IMMUNOL. 2019 10:985.





| Primers Primer | Sequence(5′-3′) | Use Application |
|---|---|---|
| Opnod1-ORF-F | ATGGGTCAGATAGAAGAAGCCAAG | ORF verification |
| Opnod1-ORF-R | TCACCATATCTCTTTGAGTGCTGTG | |
| Opnod2-ORF-F | ATGTTTGTCCAGGAGCTTGTGCTG | |
| Opnod2-ORF-R | TCAGAAGACCAGTCTTGATTCACG | |
| Optbk1-ORF-F | ATGCAGAGCACCACTAACTACCTG | |
| Optbk1-ORF-R | TCAGCCTCTCAGACCTCCGTCCAG | |
| Opnod1-qRT-F | GTTGGTGGGAGGTATTTGG | qRT-PCR |
| Opnod1-qRT-R | GTTGGTAAGGCTCGGGTG | |
| Opnod2-qRT-F | GGGGCAATAAGATAGGCG | |
| Opnod2-qRT-R | TGACGATGTTGGCGAGGG | |
| Optbk1-qRT-F | AGGACGACGAGCACTTTGTG | |
| Optbk1-qRT-R | CGTATTTCTTCTGGTGGTCTTTT | |
| β-actin-F | GCTGTGCTGTCCCTGT | |
| β-actin-R | GAGTAGCCACGCTCTGTC |
| Species | GenBank Access Number | Similarity/% | |
|---|---|---|---|
| Nod 1 | Oreochromis niloticus | XP_005472430.1 | 86.27 |
| Larimichthys crocea | XP_019134818.2 | 85.07 | |
| Paralichthys olivaceus | XP_019946646.1 | 84.39 | |
| Cynoglossus semilaevis | XP_008322367.1 | 81.05 | |
| Oryzias latipes | XP_020565632.1 | 78.45 | |
| Takifugu rubripes | XP_003965935.3 | 74.4 | |
| Esox lucius | XP_010883447.1 | 71.35 | |
| Danio rerio | XP_002665106.3 | 65.25 | |
| Macaca mulatta | XP_028701734.1 | 50.71 | |
| Mus musculus | NP_001164478.1 | 50.33 | |
| Homo sapiens | XP_011513383.1 | 49.6 | |
| Xenopus laevis | XP_031759856.1 | 48.74 | |
| Nod2 | Lates calcarifer | XP_018522174 | 89.59 |
| Larimichthys crocea | XP_010727419.3 | 85.64 | |
| Oreochromis niloticus | XP_003437591.1 | 83.82 | |
| Paralichthys olivaceus | XP_019935411.1 | 83.22 | |
| Cynoglossus semilaevis | XP_008335431.1 | 79.27 | |
| Takifugu rubripes | XP_029701512.1 | 77.25 | |
| Esox lucius | XP_010894874.4 | 67.4 | |
| Danio rerio | NP_001314973.1 | 64.18 | |
| Macaca mulatta | XP_014981593.2 | 46.26 | |
| Homo sapiens | NP_071445.1 | 46.26 | |
| Mus musculus | AAN84594.1 | 45.54 | |
| Tbk1 | Oplegnathus fasciatus | AHX37216.1 | 99.86 |
| Thunnus maccoyii | XP_042259009.1 | 98.47 | |
| Larimichthys crocea | AKM77645.1 | 98.06 | |
| Epinephelus coioides | ATI15615.1 | 97.93 | |
| Lates calcarifer | XP_018530412.1 | 97.92 | |
| Paralichthys olivaceus | XP_019966450.1 | 97.09 | |
| Solea senegalensis | XP_043878151.1 | 96.26 | |
| Cynoglossus semilaevis | XP_008313509.1 | 95.29 | |
| Danio rerio | NP_001038213.2 | 85.48 | |
| Mus musculus | NP_062760.3 | 71.78 | |
| Homo sapiens | NP_037386.1 | 71.65 | |
| Pan troglodytes | XP_509194.2 | 71.64 | |
| Xenopus laevis | NP_001086516.1 | 64.03 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




