Submitted:
26 September 2024
Posted:
27 September 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
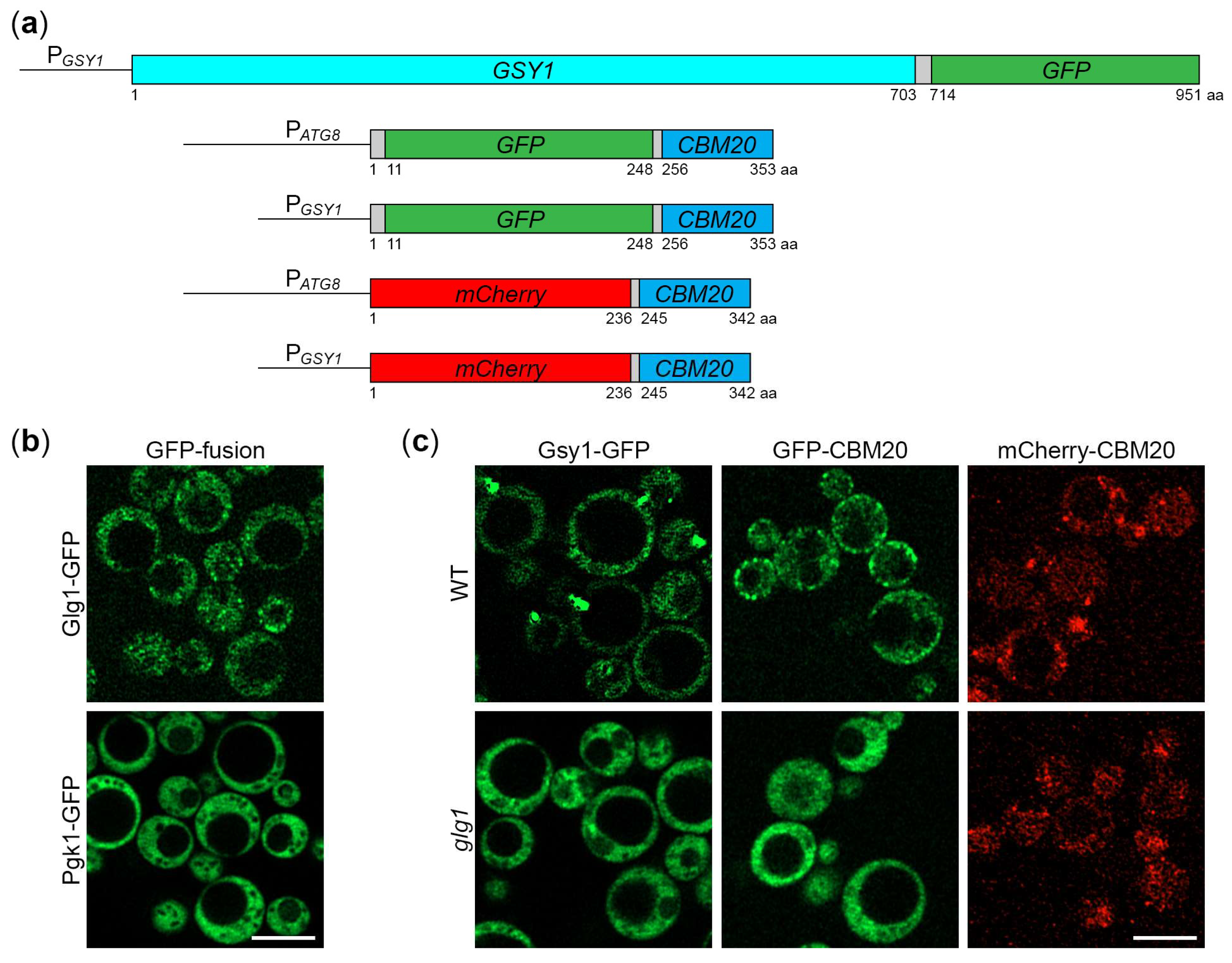
2.1. K. phaffii Gsy1-GFP Marks Glycogen Granules
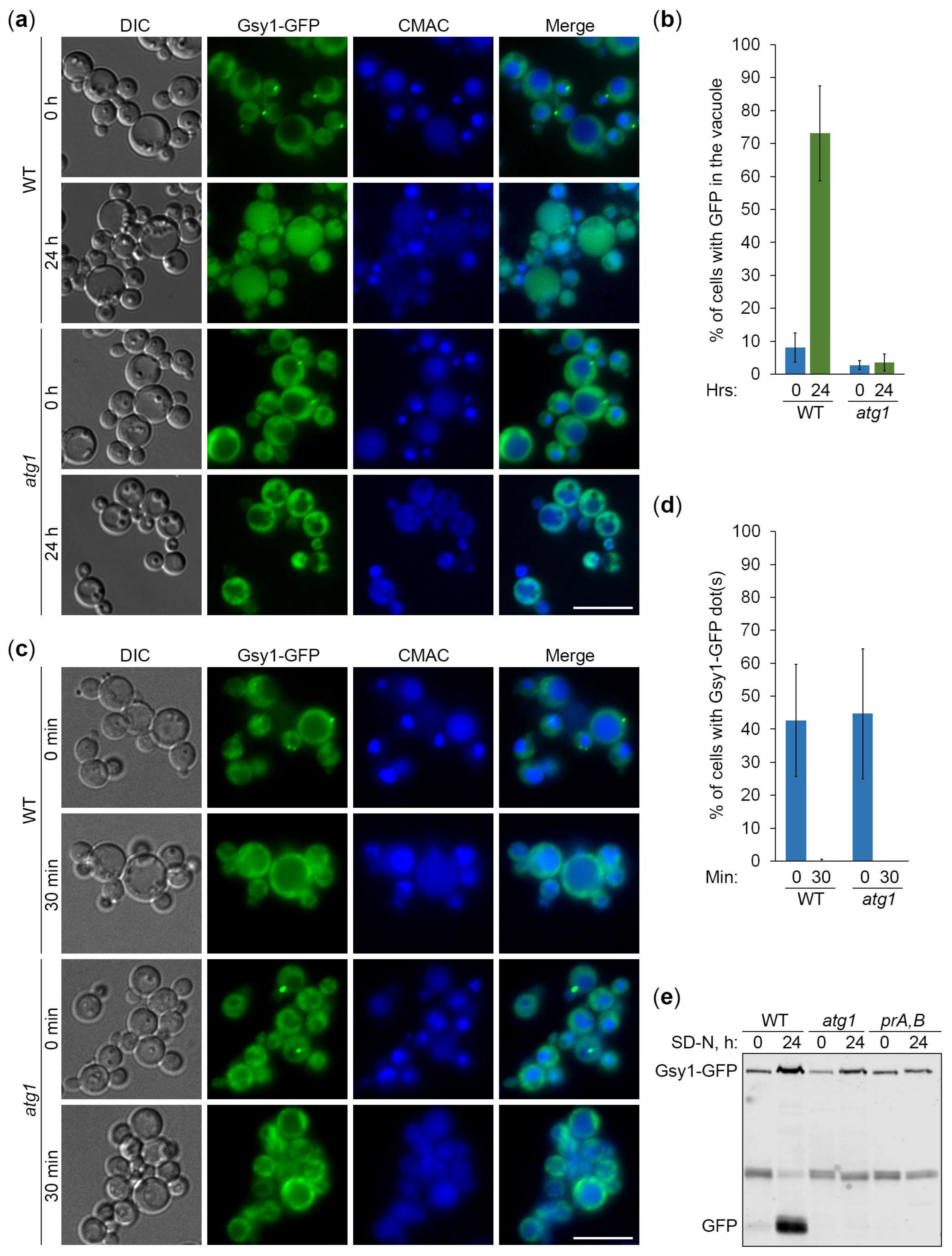
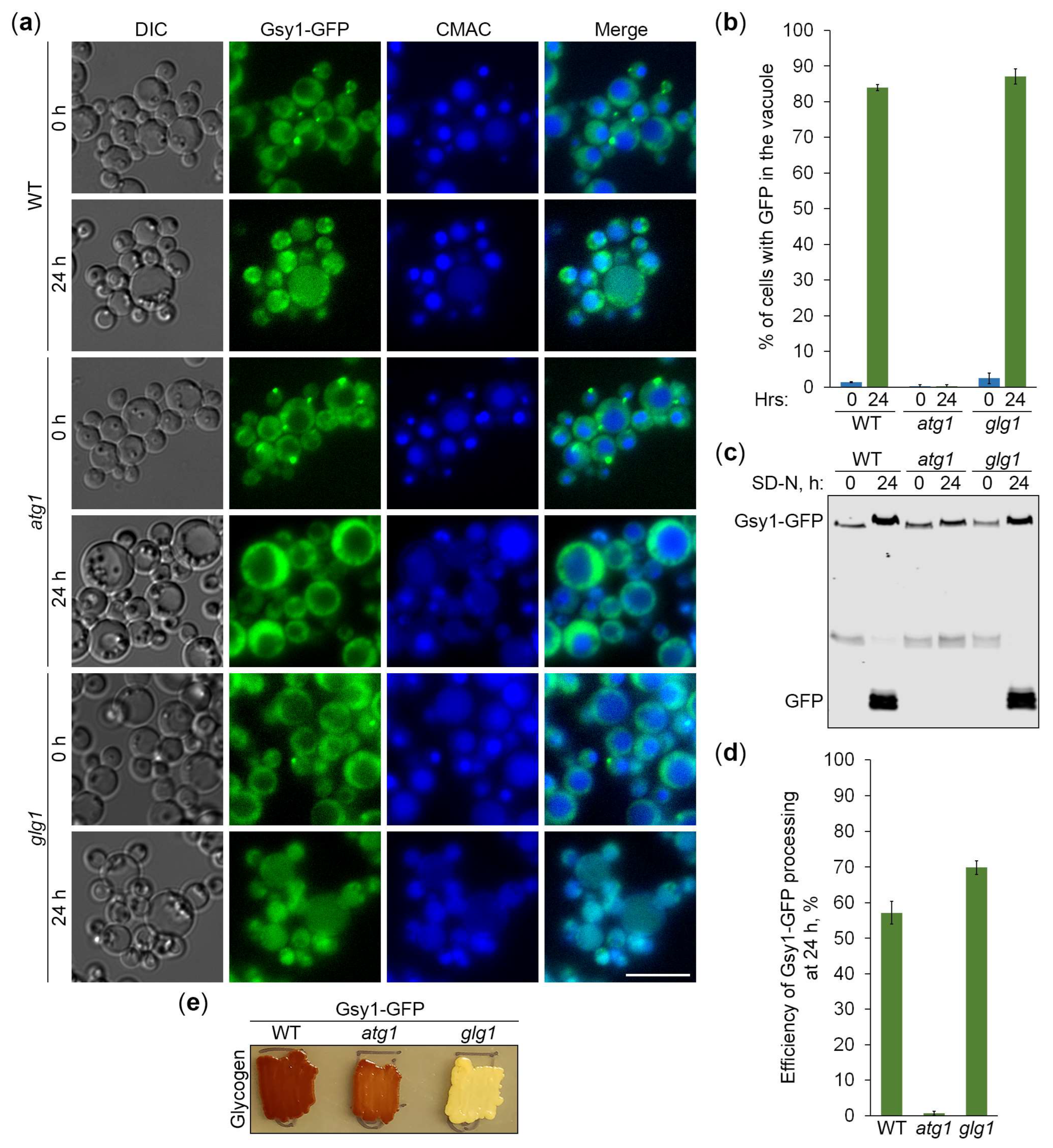
2.2. Gsy1-GFP Reports About Autophagy of Glycogen Granules
2.3. Gsy1-GFP Marked Glycogen Is Neutral Cargo of Bulk Autophagy in K. phaffii
2.4. CBM20 Fusion Proteins Mark Glycogen Granules in K. phaffii
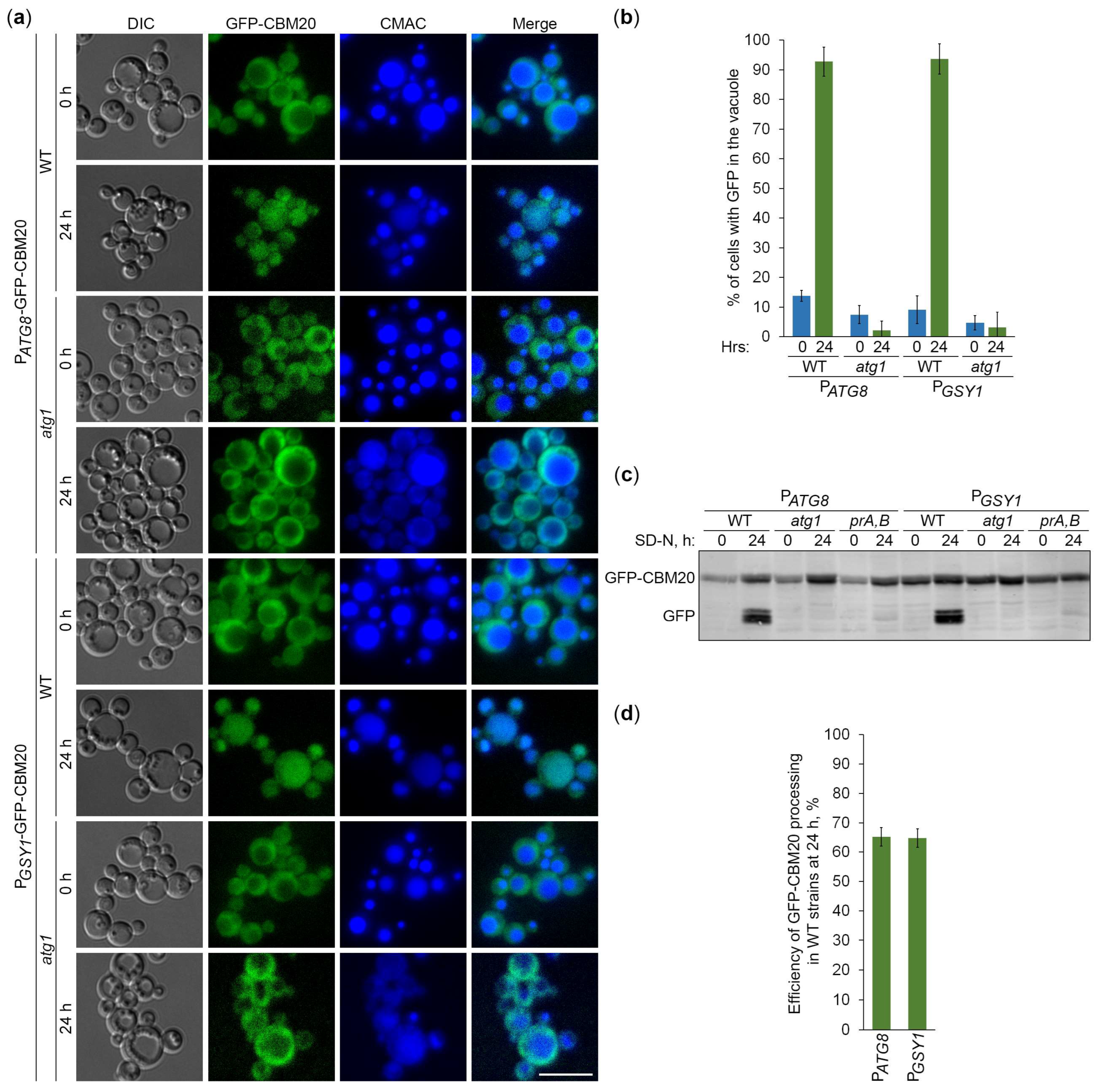
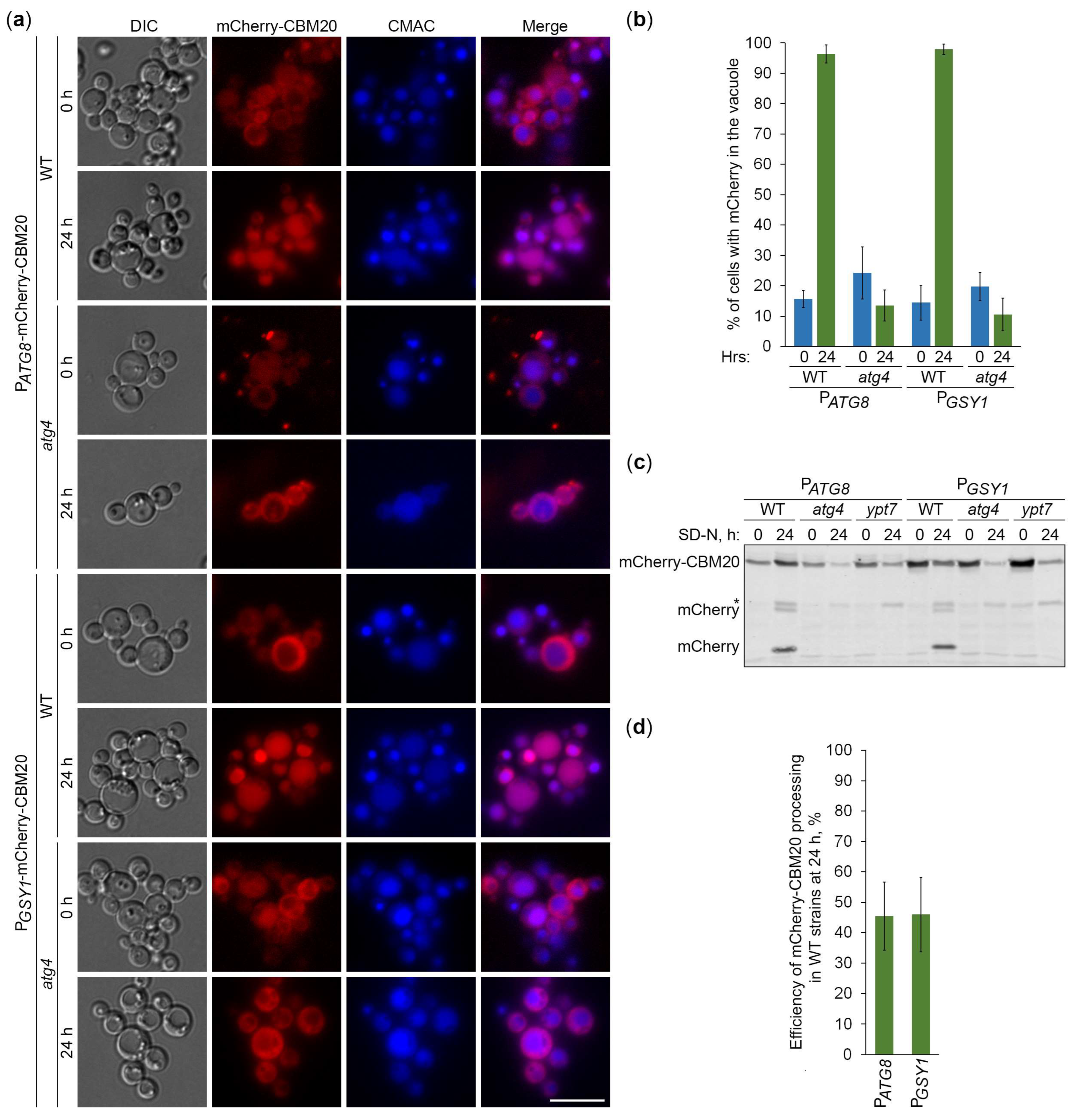
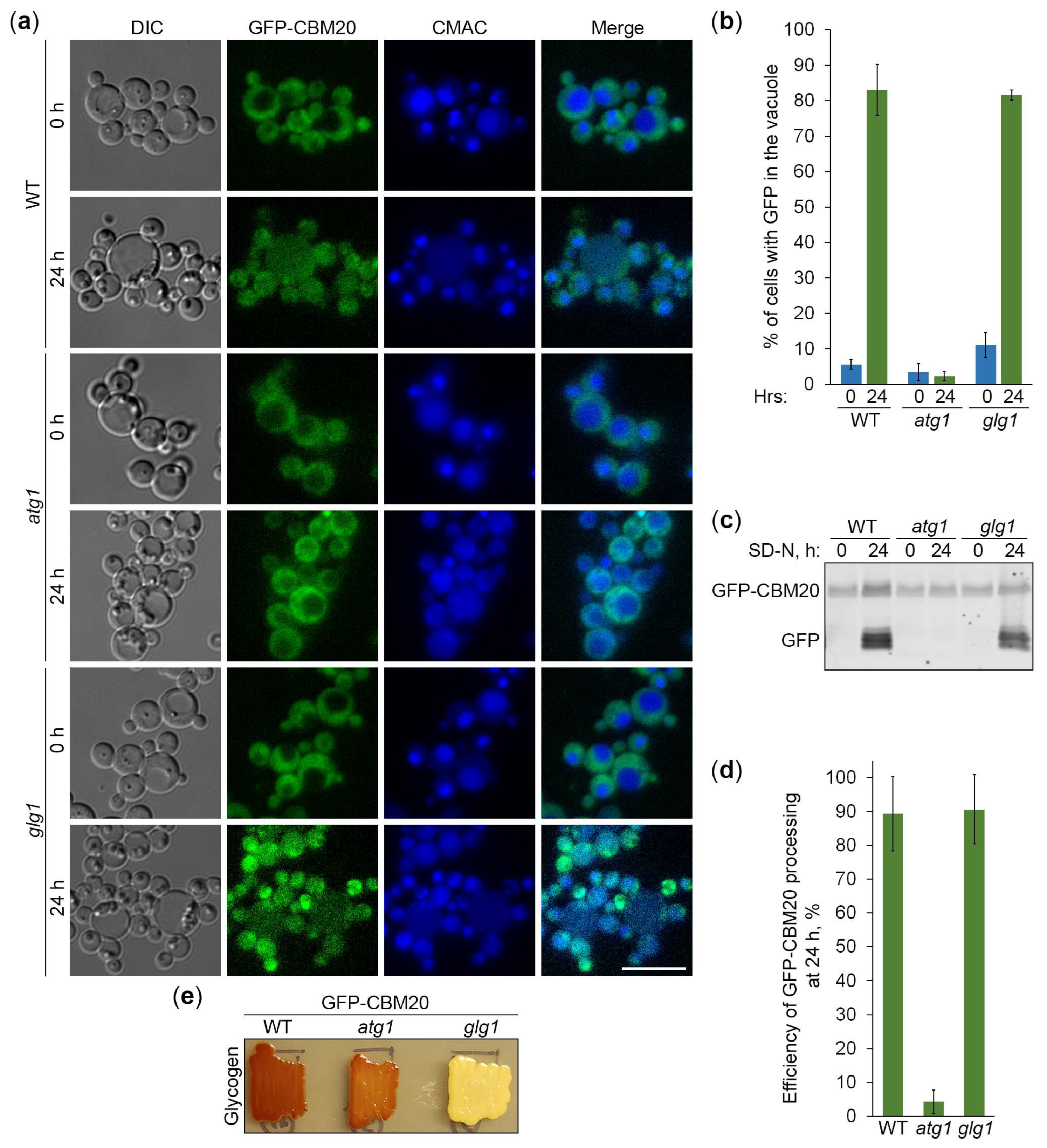
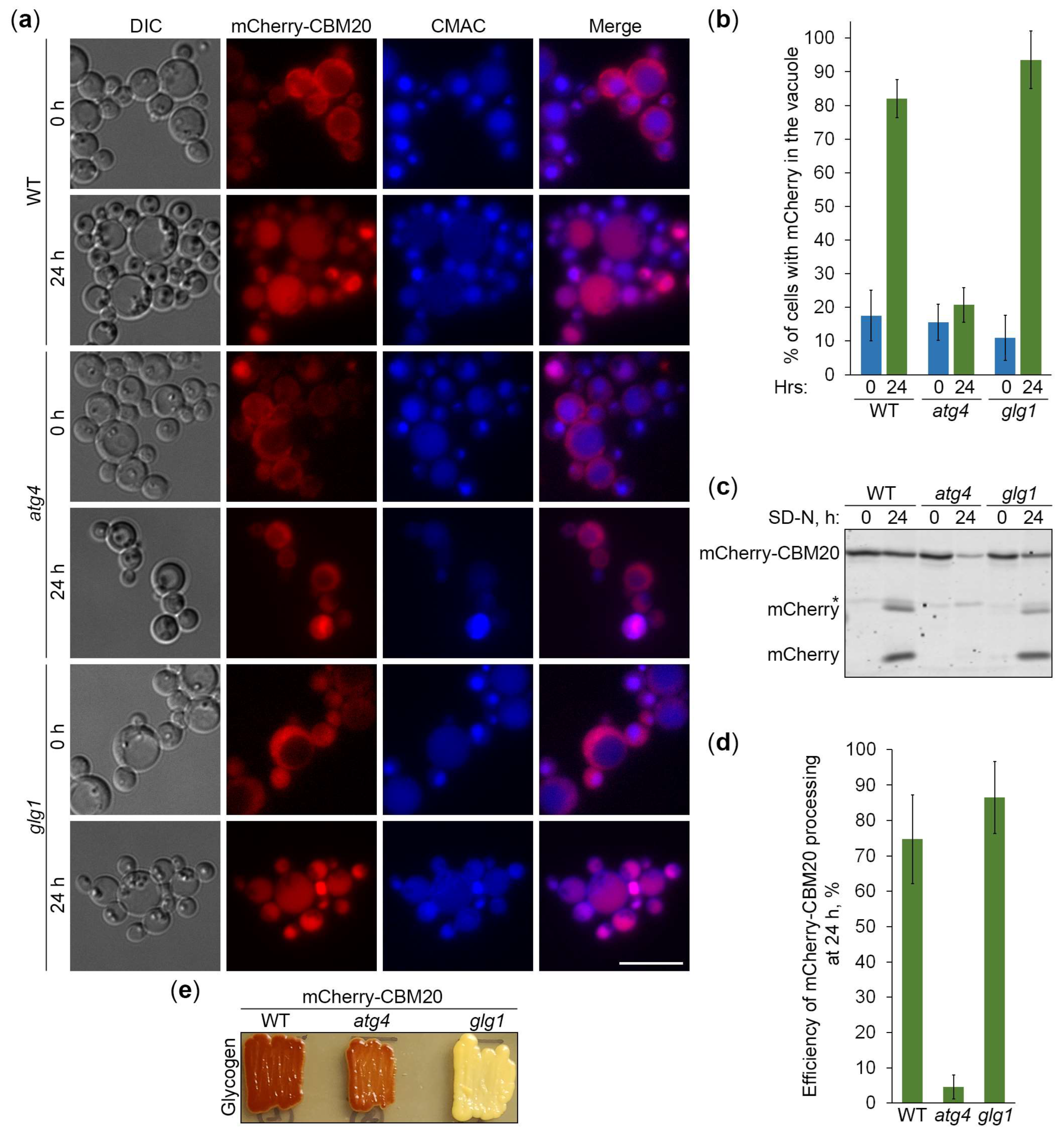
2.5. CBM20 Fusions Report About Autophagy of Glycogen Granules
2.6. Glycogen Marked with CBM20 Fusions Is Neutral Cargo of Bulk Autophagy in K. phaffii
3. Discussion
4. Materials and Methods
4.1. Strains and Plasmids
| Mutant | Strain | Background | Genotype and Plasmid | Source |
|---|---|---|---|---|
| WT | PPY12h | PPY12h | arg4 his4 | [30] |
| WT | SRK147 | PPY12h | his4::pRK22 (PGLG1-GLG1-GFP, HIS4) | [18] |
| WT | SNW78 | PPY12h | his4::pNW10 (PGLG1-PGK1-GFP, HIS4) | This study |
| WT | SRK152 | PPY12h | his4::pRK23 (PGSY1-GSY1-GFP, HIS4) | This study |
| WT | SRK176 | PPY12h | his4::pRK29 (PATG8-GFP-CBM20, HIS4) | This study |
| WT | SRK213 | PPY12h | his4::pRK34 (PGSY1-GFP-CBM20, HIS4) | This study |
| WT | SNW57 | PPY12h | arg4::pRK28 (PATG8-mCherry-CBM20, ARG4) | This study |
| WT | SNW59 | PPY12h | arg4::pRK32 (PGSY1-mCherry-CBM20, ARG4) | This study |
| atg1 | R12 | GS115 | atg1-1::ZeocinR his4 | [21] |
| atg1 | SRK154 | R12 | his4::pRK23 (PGSY1-GSY1-GFP, HIS4) | This study |
| atg1 | SRK178 | R12 | his4::pRK29 (PATG8-GFP-CBM20, HIS4) | This study |
| atg1 | SRK215 | R12 | his4::pRK34 (PGSY1-GFP-CBM20, HIS4) | This study |
| atg4 | PPM408 | PPY12h | atg4::ZeocinR arg4 his4 | [24] |
| atg4 | SNW55 | PPM408 | arg4::pRK28 (PATG8-mCherry-CBM20, ARG4) | This study |
| atg4 | SNW62 | PPM408 | arg4::pRK32 (PGSY1-mCherry-CBM20, ARG4) | This study |
| prA,B | SMD1163 | GS115 | pep4 prb1 his4 | [22] |
| prA,B | SRK157 | SMD1163 | his4::pRK23 (PGSY1-GSY1-GFP, HIS4) | This study |
| prA,B | SRK180 | SMD1163 | his4::pRK29 (PATG8-GFP-CBM20, HIS4) | This study |
| prA,B | SRK217 | SMD1163 | his4::pRK34 (PGSY1-GFP-CBM20, HIS4) | This study |
| ypt7 | SRRM197 | PPY12h | Δypt7::GeneticinR arg4 his4 | [25] |
| ypt7 | SPB1 | SRRM197 | arg4::pRK28 (PATG8-mCherry-CBM20, ARG4) | This study |
| ypt7 | SPB2 | SRRM197 | arg4::pRK32 (PGSY1-mCherry-CBM20, ARG4) | This study |
| glg1 | SNW49 | PPY12h | Δglg1::ZeocinR (pNW9) | [18] |
| glg1 | SNW51 | SNW49 | his4::pRK23 (PGSY1-GSY1-GFP, HIS4) | This study |
| glg1 | SNW53 | SNW49 | his4::pRK34 (PGSY1-GFP-CBM20, HIS4) | This study |
| glg1 | SNW64 | SNW49 | arg4::pRK32 (PGSY1-mCherry-CBM20, ARG4) | This study |
4.2. Immunoblotting
4.3. Epifluorescence Microscopy
4.4. Confocal Microscopy
4.5. Glycogen Staining
4.6. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Prats, C.; Graham, T.E.; Shearer, J. The dynamic life of the glycogen granule. J Biol Chem 2018, 293, 7089-7098. [CrossRef]
- Liu, Q.H.; Tang, J.W.; Wen, P.B.; Wang, M.M.; Zhang, X.; Wang, L. From Prokaryotes to Eukaryotes: Insights Into the Molecular Structure of Glycogen Particles. Front Mol Biosci 2021, 8, 673315. [CrossRef]
- Skurat, A.V.; Dietrich, A.D.; Roach, P.J. Interaction between glycogenin and glycogen synthase. Arch Biochem Biophys 2006, 456, 93-97. [CrossRef]
- François, J.; Parrou, J.L. Reserve carbohydrates metabolism in the yeast Saccharomyces cerevisiae. FEMS Microbiology Reviews 2001, 25, 125-145. [CrossRef]
- Wilson, W.A.; Roach, P.J.; Montero, M.; Baroja-Fernandez, E.; Munoz, F.J.; Eydallin, G.; Viale, A.M.; Pozueta-Romero, J. Regulation of glycogen metabolism in yeast and bacteria. FEMS Microbiol Rev 2010, 34, 952-985. [CrossRef]
- Roach, P.J.; Depaoli-Roach, A.A.; Hurley, T.D.; Tagliabracci, V.S. Glycogen and its metabolism: some new developments and old themes. Biochem J 2012, 441, 763-787. [CrossRef]
- Schiaffino, S.; Hanzlikova, V. Autophagic degradation of glycogen in skeletal muscles of the newborn rat. J Cell Biol 1972, 52, 41-51. [CrossRef]
- Raben, N.; Schreiner, C.; Baum, R.; Takikita, S.; Xu, S.; Xie, T.; Myerowitz, R.; Komatsu, M.; Van der Meulen, J.H.; Nagaraju, K.; et al. Suppression of autophagy permits successful enzyme replacement therapy in a lysosomal storage disorder--murine Pompe disease. Autophagy 2010, 6, 1078-1089. [CrossRef]
- Klionsky, D.J.; Petroni, G.; Amaravadi, R.K.; Baehrecke, E.H.; Ballabio, A.; Boya, P.; Bravo-San Pedro, J.M.; Cadwell, K.; Cecconi, F.; Choi, A.M.K.; et al. Autophagy in major human diseases. EMBO J 2021, 40, e108863. [CrossRef]
- Pant, D.C.; Nazarko, T.Y. Selective autophagy: the rise of the zebrafish model. Autophagy 2021, 17, 3297-3305. [CrossRef]
- Nazarko, T.Y.; Farre, J.C.; Polupanov, A.S.; Sibirny, A.A.; Subramani, S. Autophagy-related pathways and specific role of sterol glucoside in yeasts. Autophagy 2007, 3, 263-265. [CrossRef]
- Wang, Z.; Wilson, W.A.; Fujino, M.A.; Roach, P.J. Antagonistic Controls of Autophagy and Glycogen Accumulation by Snf1p, the Yeast Homolog of AMP-Activated Protein Kinase, and the Cyclin-Dependent Kinase Pho85p. Molecular and Cellular Biology 2001, 21, 5742-5752. [CrossRef]
- Wilson, W.A.; Wang, Z.; Roach, P.J. Systematic Identification of the Genes Affecting Glycogen Storage in the Yeast Saccharomyces cerevisiae: Implication of the Vacuole as a Determinant of Glycogen Level*S. Molecular & Cellular Proteomics 2002, 1, 232-242. [CrossRef]
- Jiang, S.; Heller, B.; Tagliabracci, V.S.; Zhai, L.; Irimia, J.M.; DePaoli-Roach, A.A.; Wells, C.D.; Skurat, A.V.; Roach, P.J. Starch binding domain-containing protein 1/genethonin 1 is a novel participant in glycogen metabolism. J Biol Chem 2010, 285, 34960-34971. [CrossRef]
- Jiang, S.; Wells, C.D.; Roach, P.J. Starch-binding domain-containing protein 1 (Stbd1) and glycogen metabolism: Identification of the Atg8 family interacting motif (AIM) in Stbd1 required for interaction with GABARAPL1. Biochemical and Biophysical Research Communications 2011, 413, 420-425. [CrossRef]
- Yi, H.; Fredrickson, K.B.; Das, S.; Kishnani, P.S.; Sun, B. Stbd1 is highly elevated in skeletal muscle of Pompe disease mice but suppression of its expression does not affect lysosomal glycogen accumulation. Mol Genet Metab 2013, 109, 312-314. [CrossRef]
- Sun, T.; Yi, H.; Yang, C.; Kishnani, P.S.; Sun, B. Starch Binding Domain-containing Protein 1 Plays a Dominant Role in Glycogen Transport to Lysosomes in Liver. J Biol Chem 2016, 291, 16479-16484. [CrossRef]
- Wijewantha, N.V.; Kumar, R.; Nazarko, T.Y.J.C. Glycogen Granules Are Degraded by Non-Selective Autophagy in Nitrogen-Starved Komagataella phaffii. 2024, 13, 467. [CrossRef]
- Nazarko, T.Y. Autophagy of Glycogen Is Non-Selective in Komagataella phaffii. Autophagy Rep 2024, 3. [CrossRef]
- Isoda, T.; Takeda, E.; Hosokawa, S.; Hotta-Ren, S.; Ohsumi, Y. Atg45 is an autophagy receptor for glycogen, a non-preferred cargo of bulk autophagy in yeast. iScience 2024, 27, 109810. [CrossRef]
- Stromhaug, P.E.; Bevan, A.; Dunn, W.A., Jr. GSA11 encodes a unique 208-kDa protein required for pexophagy and autophagy in Pichia pastoris. J Biol Chem 2001, 276, 42422-42435. [CrossRef]
- Tuttle, D.L.; Dunn, W.A., Jr. Divergent modes of autophagy in the methylotrophic yeast Pichia pastoris. J Cell Sci 1995, 108 ( Pt 1), 25-35. [CrossRef]
- Skurat, A.V.; Segvich, D.M.; DePaoli-Roach, A.A.; Roach, P.J. Novel method for detection of glycogen in cells. Glycobiology 2017, 27, 416-424. [CrossRef]
- Mukaiyama, H.; Oku, M.; Baba, M.; Samizo, T.; Hammond, A.T.; Glick, B.S.; Kato, N.; Sakai, Y. Paz2 and 13 other PAZ gene products regulate vacuolar engulfment of peroxisomes during micropexophagy. Genes Cells 2002, 7, 75-90. [CrossRef]
- Manjithaya, R.; Anjard, C.; Loomis, W.F.; Subramani, S. Unconventional secretion of Pichia pastoris Acb1 is dependent on GRASP protein, peroxisomal functions, and autophagosome formation. J Cell Biol 2010, 188, 537-546. [CrossRef]
- Cregg, J.M.; Russell, K.A. Transformation. Methods Mol Biol 1998, 103, 27-39. [CrossRef]
- Kumar, R.; Shroff, A.; Nazarko, T.Y. Komagataella phaffii Cue5 Piggybacks on Lipid Droplets for Its Vacuolar Degradation during Stationary Phase Lipophagy. Cells 2022, 11. [CrossRef]
- Farre, J.C.; Mathewson, R.D.; Manjithaya, R.; Subramani, S. Roles of Pichia pastoris Uvrag in vacuolar protein sorting and the phosphatidylinositol 3-kinase complex in phagophore elongation in autophagy pathways. Autophagy 2010, 6, 86-99. [CrossRef]
- Sears, I.B.; O'Connor, J.; Rossanese, O.W.; Glick, B.S. A versatile set of vectors for constitutive and regulated gene expression in Pichia pastoris. Yeast 1998, 14, 783-790. [CrossRef]
- Gould, S.J.; McCollum, D.; Spong, A.P.; Heyman, J.A.; Subramani, S. Development of the yeast Pichia pastoris as a model organism for a genetic and molecular analysis of peroxisome assembly. Yeast 1992, 8, 613-628. [CrossRef]
- Baerends, R.J.; Faber, K.N.; Kram, A.M.; Kiel, J.A.; van der Klei, I.J.; Veenhuis, M. A stretch of positively charged amino acids at the N terminus of Hansenula polymorpha Pex3p is involved in incorporation of the protein into the peroxisomal membrane. J Biol Chem 2000, 275, 9986-9995. [CrossRef]
- Corpet, F. Multiple sequence alignment with hierarchical clustering. Nucleic Acids Res 1988, 16, 10881-10890. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




