3. Results and Discussion
The computational study involved the docking of all datasets, as described, within the open pore's active region of GABA-AT, sodium channel, JNK, and Anti-Oxidant. Following the process detailed in the new section, we determined the predicted binding energies of the active site of these inhibitors, as presented in
Table 2. The computation of these energies for binding and docking was carried out using AutoDock4.
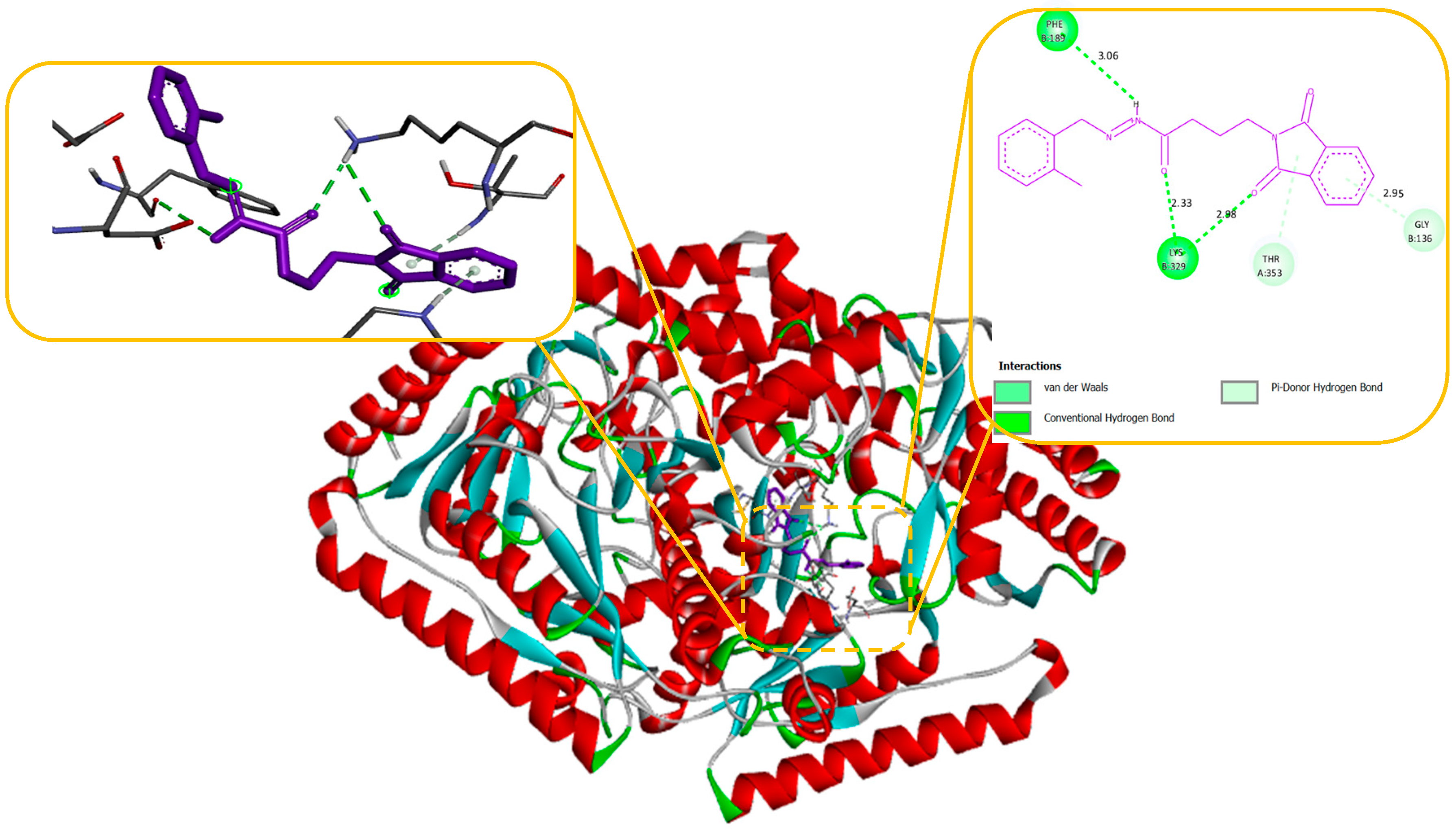
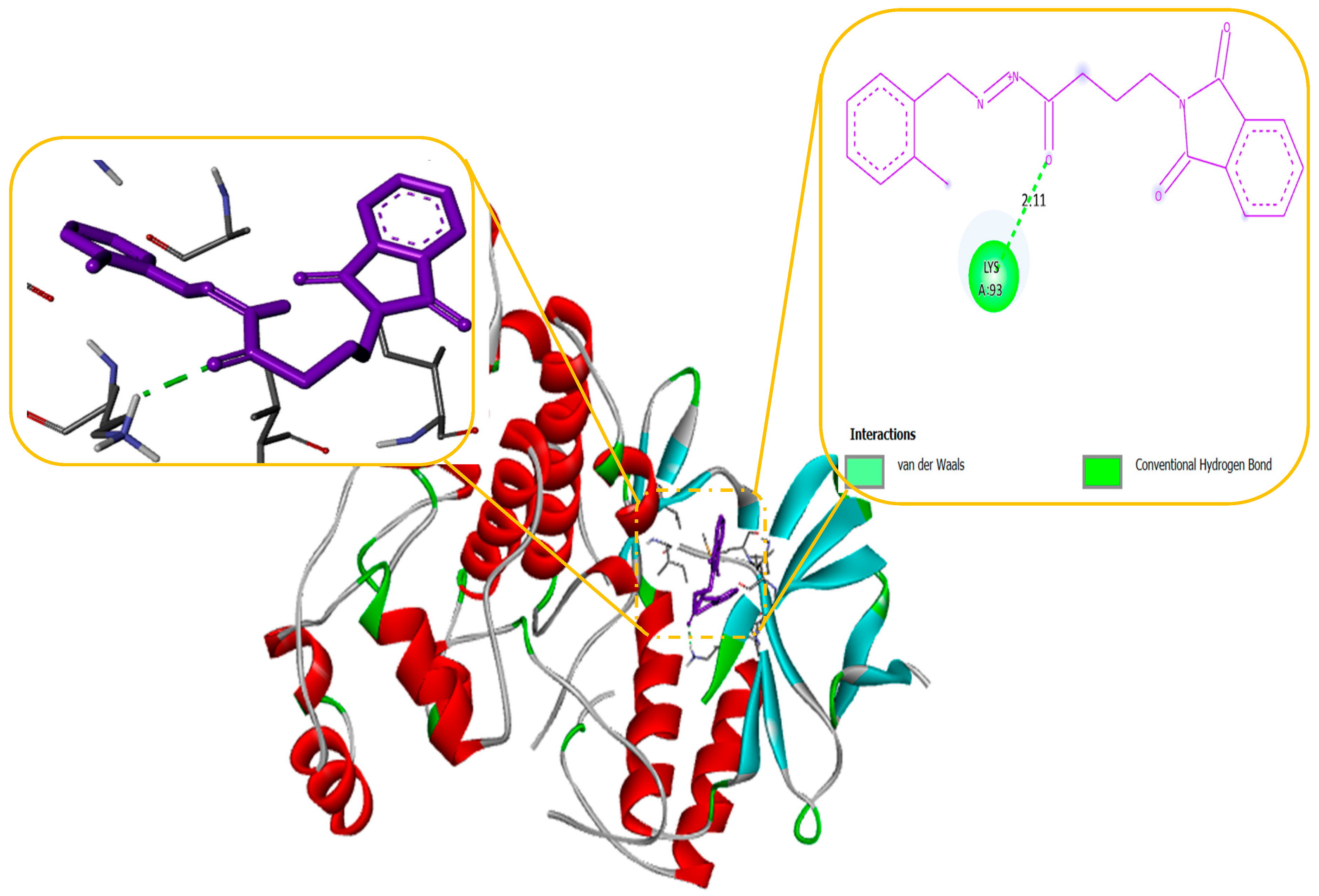
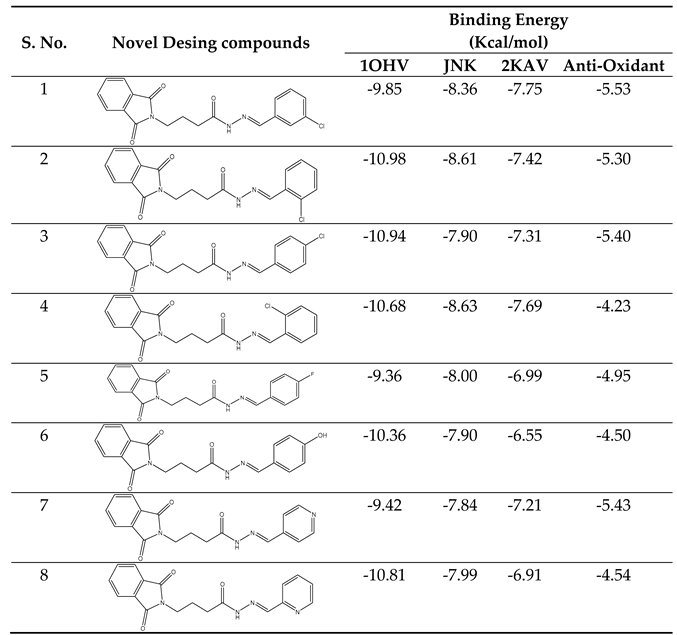
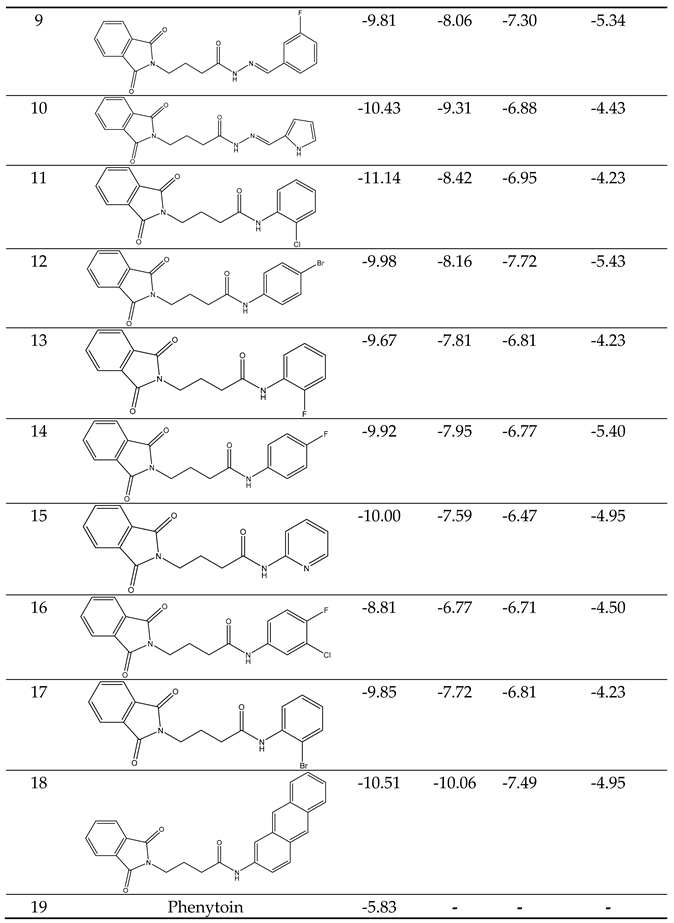
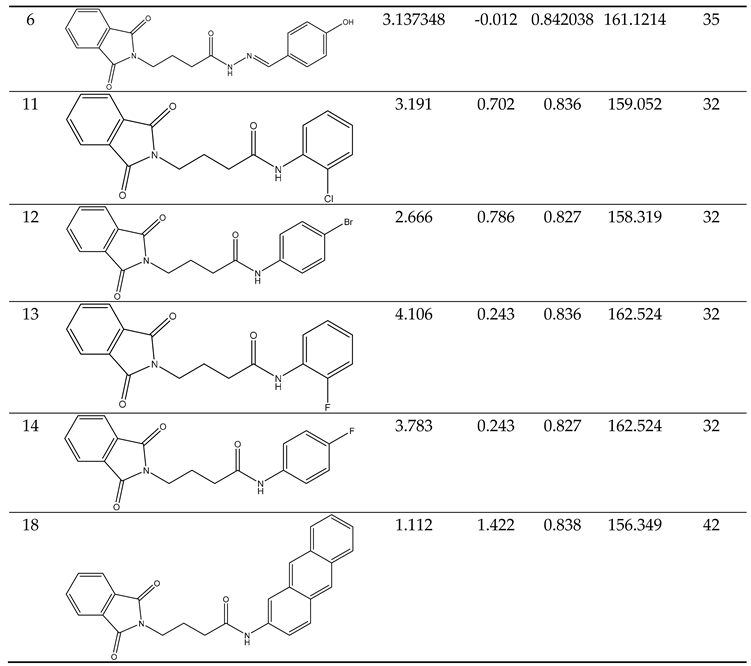
Our docking results show that compounds 2, 3, 4, 6, 8, 10, 11, 15, and 18 have binding energies that are, respectively -10.98, -10.94, 10.68, -10.36, 10.81, -10.43, -11.14, -10.00 and -10.51 kcal/mol. These values indicate that these compounds are more potent than phenytoin, which has a binding energy of -5.83 kcal/mol, serving as a Reference drug. Our docking studies interaction patterns of compound 2 between GABA-AT, sodium channel, JNK, and Anti-Oxidant with N-Phenyl Phthalimide derivatives (
Figure 3,
Figure 4,
Figure 5 and
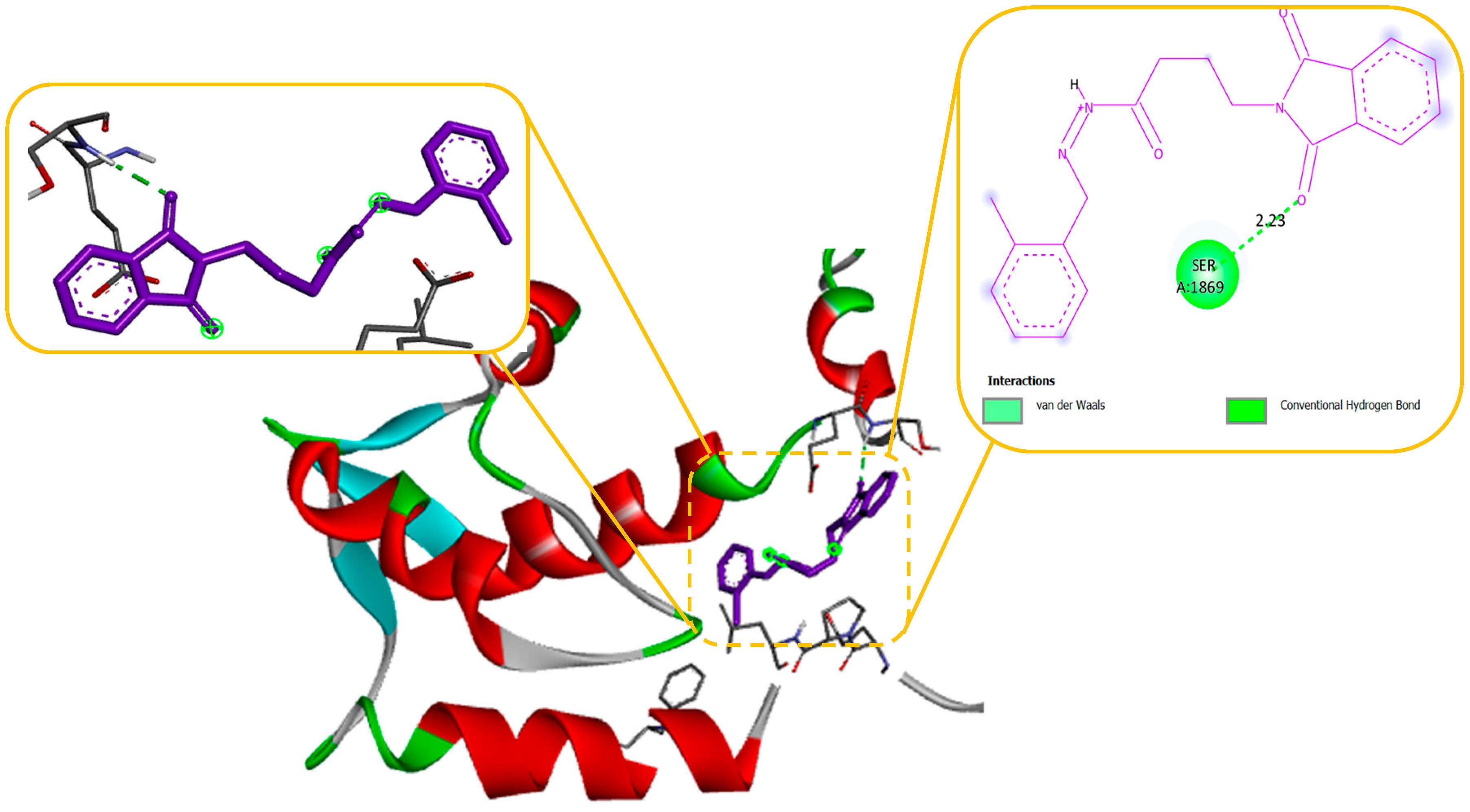
Figure 6) with their 2D and 3D interaction. Compound 2 primarily cooperated with NaV1.2 domain IV-S6, as depicted in
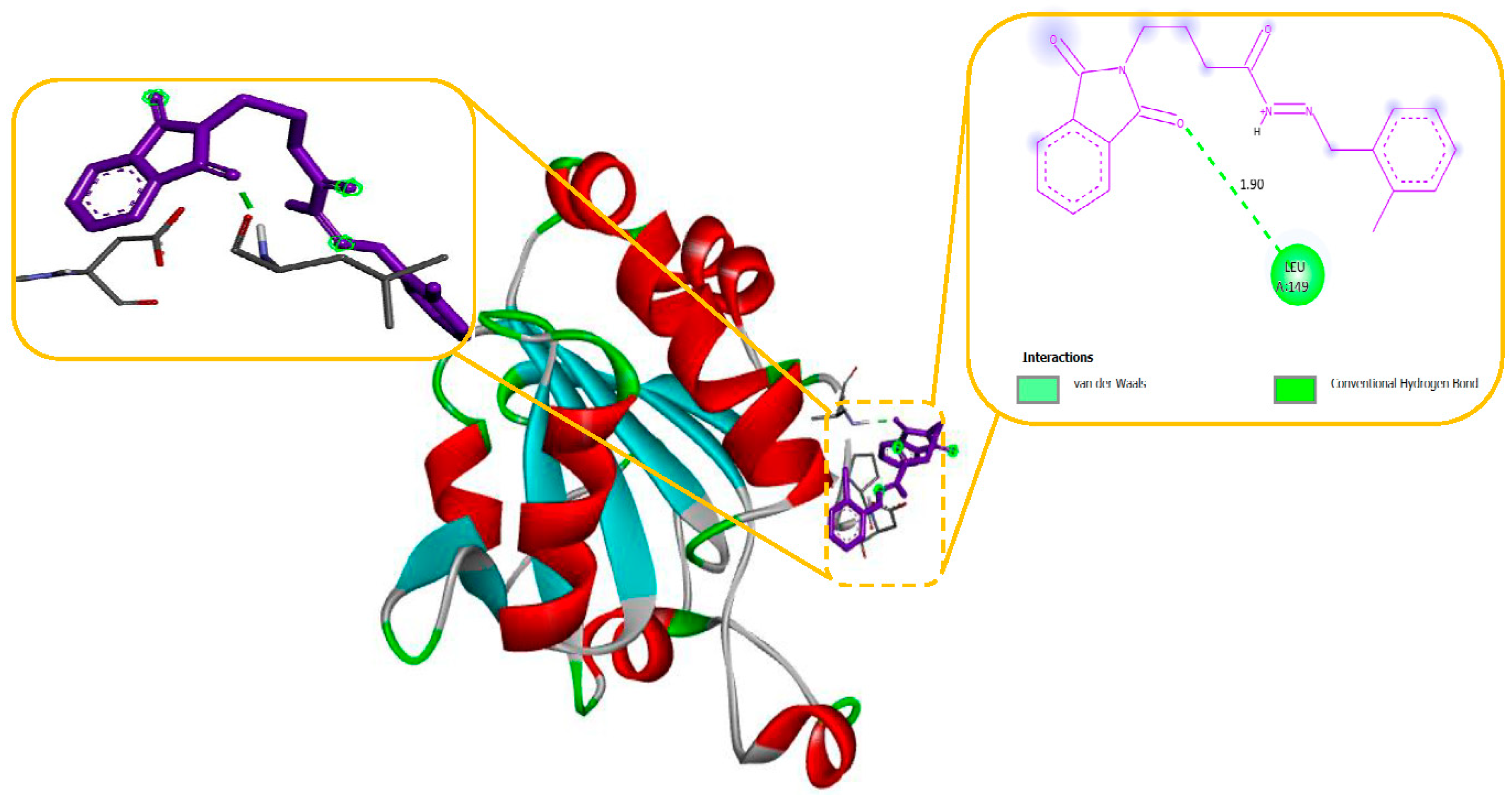
Figure 4. The N-Phenyl Phthalimide derivatives, on the other hand, demonstrated a predominant contact with domain II-S6. This interaction involved the formation of hydrogen bonds and additional hydrophobic interactions, primarily within the inner pore of the channel, with domains I, II, IV, and sometimes with domain III, as illustrated in
Figure 4.
Molecular modelling further revealed specific hydrogen bonding interactions in compound 11. For instance, the oxygen atom in the NO
2 group established hydrogen bonds with LYS 329 (Figure 7) and also the OH group with the PHE 189 and ASP298 by docking with GABA-AT. Similar hydrogen bonds were created between the GLU1788 and the NH atom in compound 11 (
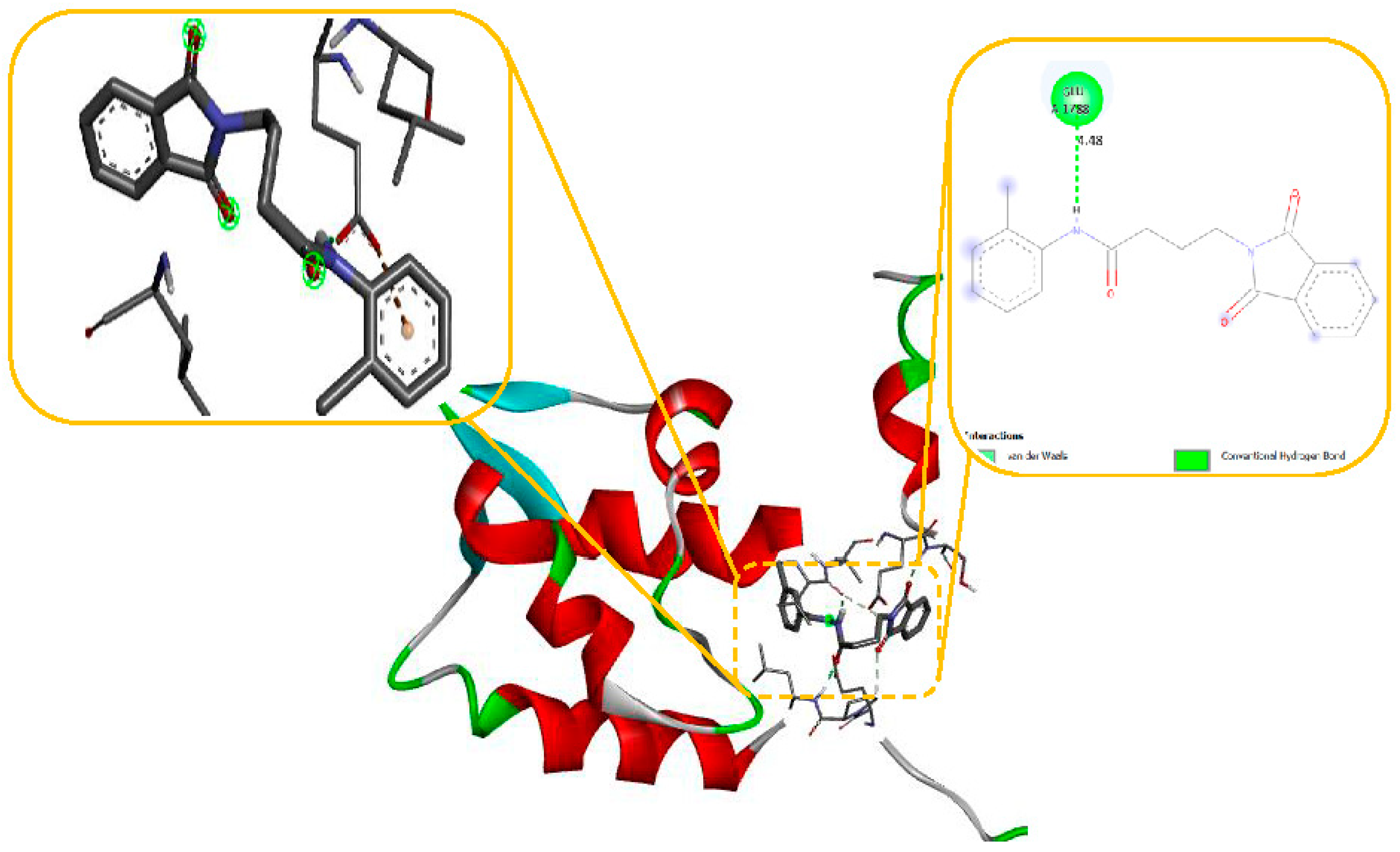
Figure 8) while docking with Sodithe um Channel. Additionally, according to a docking prediction shown in
Figure 9, a hydrogen bond between the oxygen atom of the compound 11 group and the hydrogen atom of the MET149 group could possibly form inside the active site with JNK receptor.
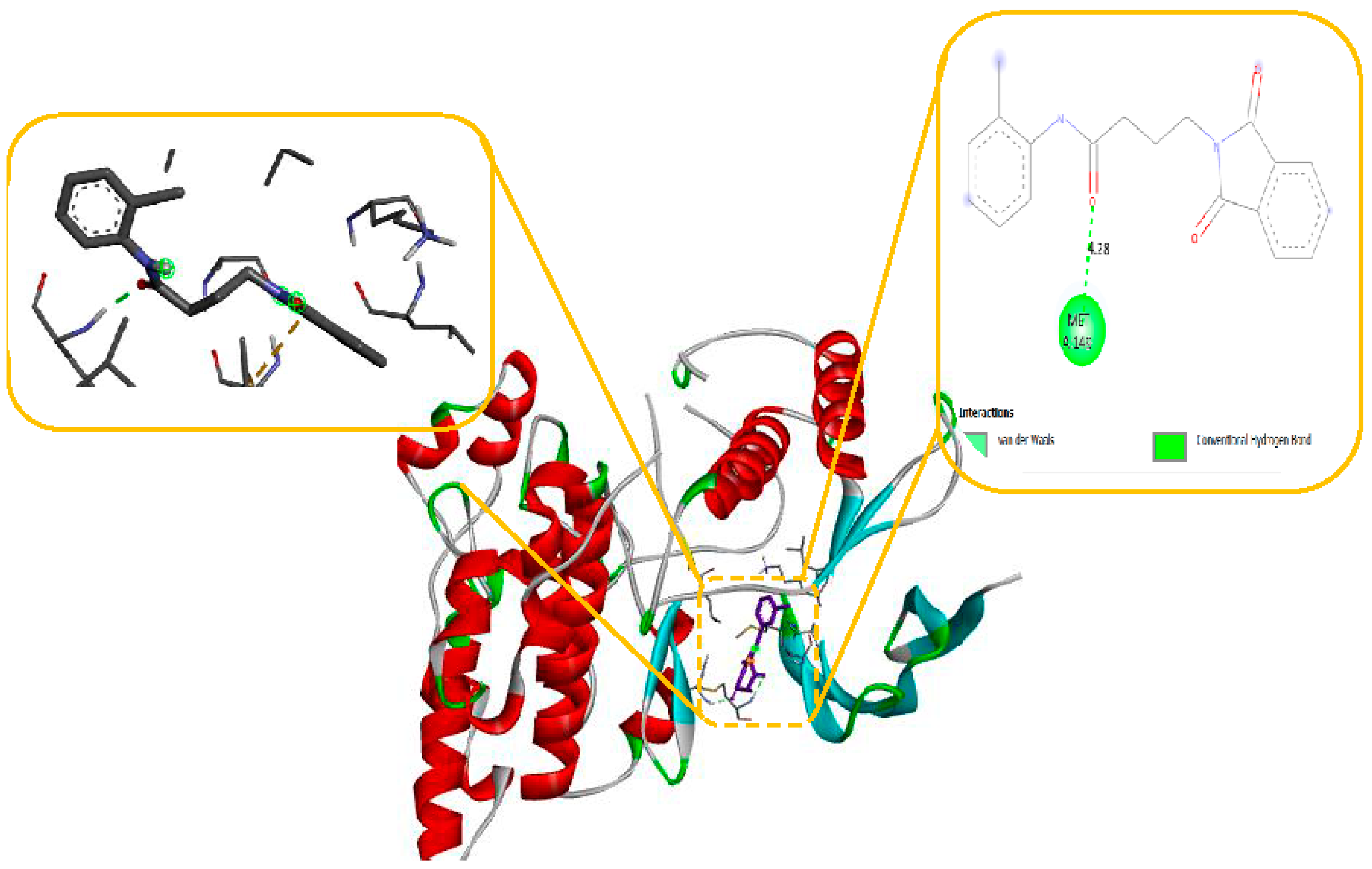
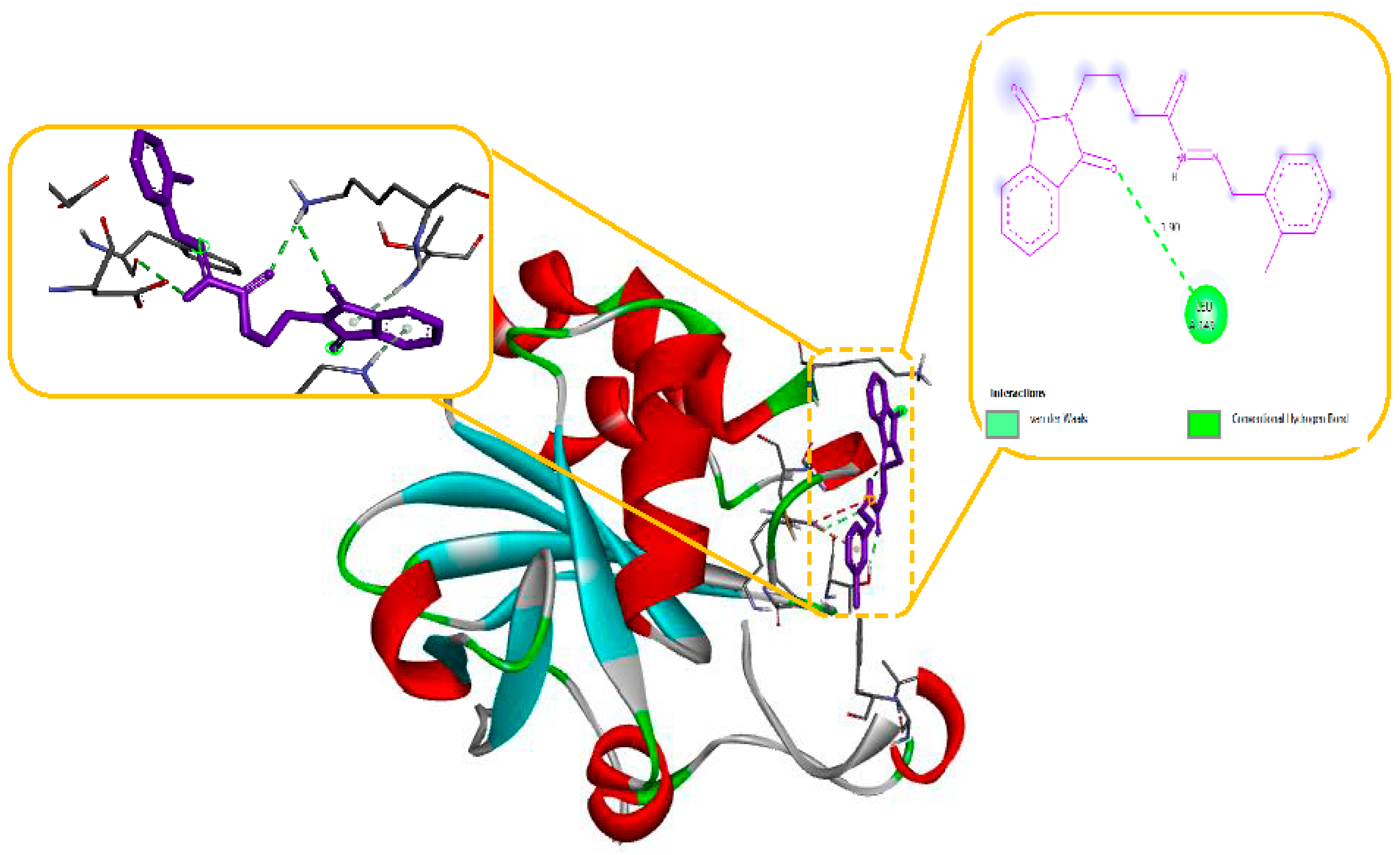
As seen in
Figure 10, the N-phenyl ring of LEU 149 and the aromatic ring of compound 11 effectively interacted via a Hydrogen bonding interaction in the domain with Anti-oxidant Receptor.
Despite the promising Predicated binding energies outlined in
Table 3, the investigated substances' activities.
The dataset comprised a total of 11 molecules, and the application of the Modified-K-Medoids1.2 algorithm in our study resulted in the creation of training. The compounds and associated activity values for the compounds in the dataset have been described in
Table 4.
Values in parenthesis in Eq. (1) represent standard deviation. The ratio of training set compounds to model descriptors, expressed as a Topliss ratio [
23]. This demonstrated that the model followed the general semi-empirical QSAR rule [
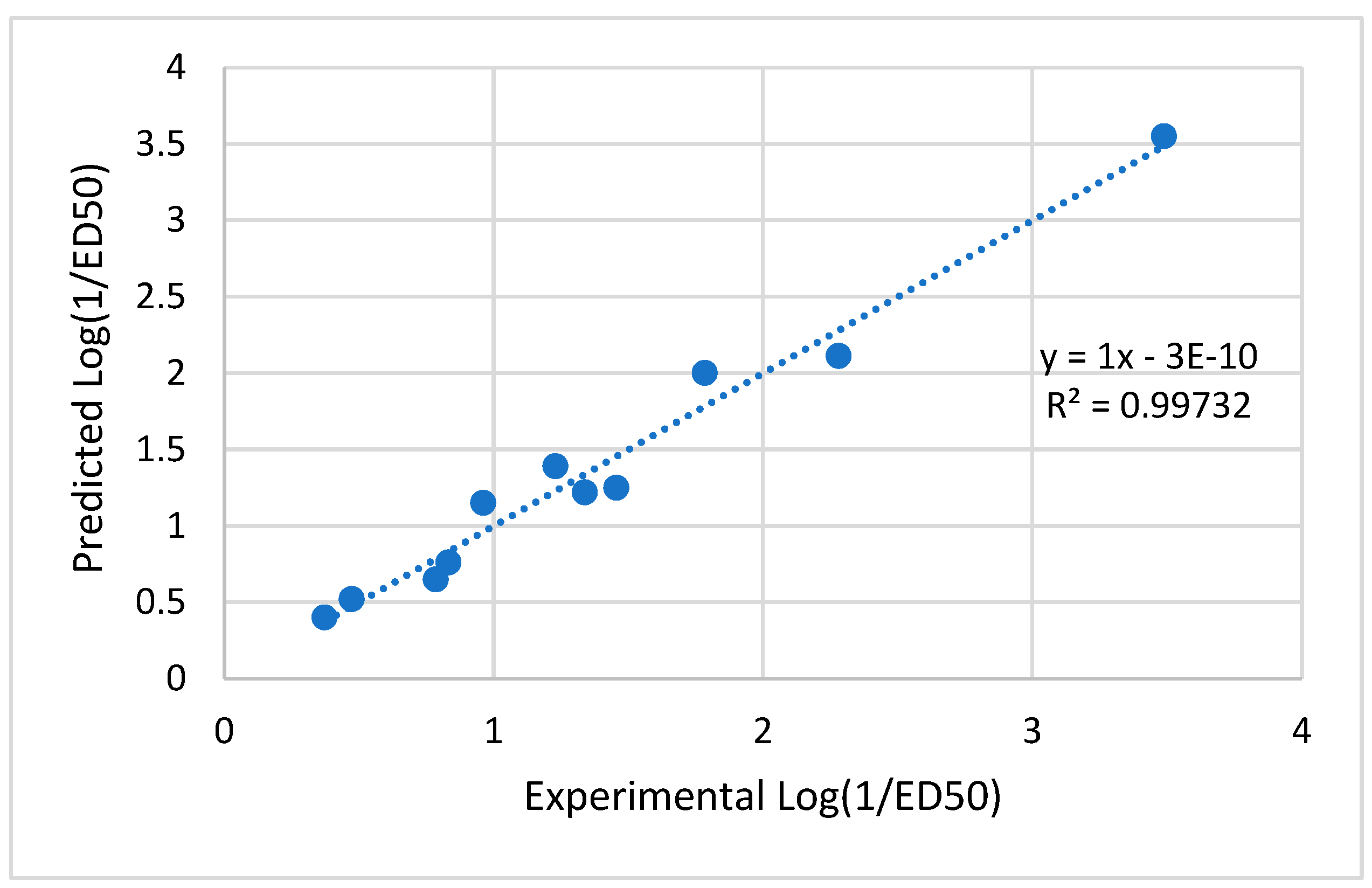
24]. These indicated that the model had no multi-co-linearity issues and that descriptors had made a major contribution to the model at a 95 percent level. Moreover, in the plots depicting the predicted versus experimenting with the data from the test set and the training set, the correlation coefficient (r
2) values exceeded 0.6 (refer to
Figure 11). This finding suggests that there is a linear link the values of the Predicted and experimental activity. The model created for the study seems to have good quality based on the results so far [
25,
26].
External and Internal Validation
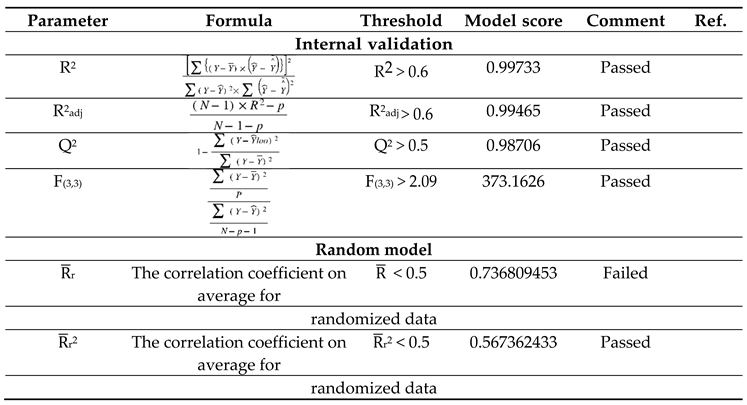
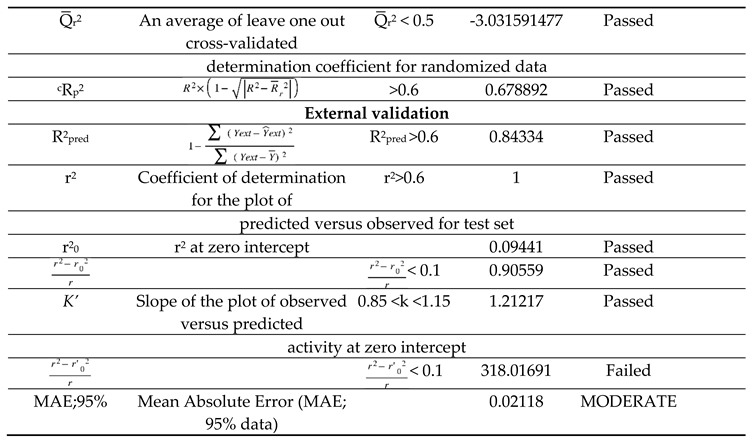
The study's validation model was displayed in
Table 5 validation parameters. Q
2, R
2, R
2 adj, R
2 cross-validated, and R
2 predicted for test set for MLR determination. R
2 pred, the Y-randomization parameters, and the correlation coefficient between the test set's predicted activity and actual activity. [
27] R
2 were greater than 0.5. Model's F3,3 was significantly higher than the calculated F-value for 3, 3 degrees of freedom at the 95% level of significance (373.162). These results demonstrated that the proposed model was highly capable of internal and external prediction, that it is not a random event, and that it can explain the variation in molecular activity levels of the dataset with 95% accuracy. [
28,
29] The model complied with all of the mean absolute error (MAE) specifications for a predictive model proposed by Roy et al. [
30] (
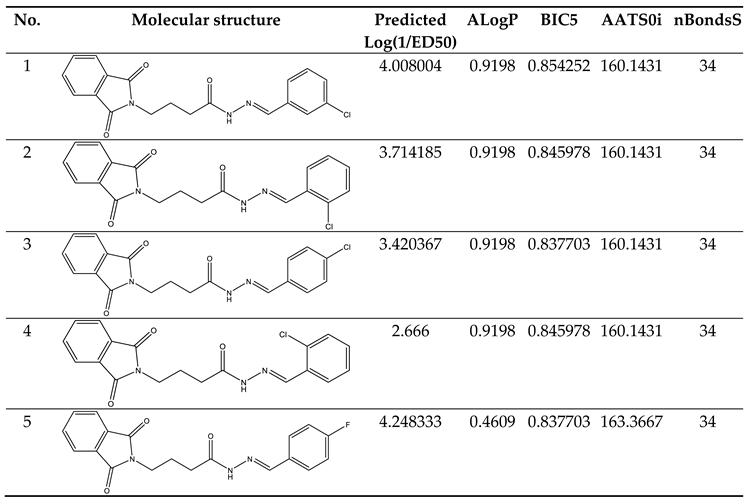
Table 5). Due to its robustness, dependability, and applicability to forecast anticonvulsant action for both known and hypothetical compounds, the study's presented model had good internal and external as well as internal prediction abilities. With the help of Prediction Reliability Indicator 1.0, compounds designed were tested for prediction power some molecules predicted good anticonvulsant activity using the default setting (
Table 6). Hence, the designed compounds 2, and compound 11 were found to be good predicted anticonvulsant activity.
In this context, the variables are defined as follows:
- Y represents the training set's observed activity value.
- Ȳ stands for the average of the training set's observed activity values.
- Predicted activity for training is consistent with the predicted activity levels for the training set.
- loo represents using leave-one-out cross-validation, the training set's expected activity.
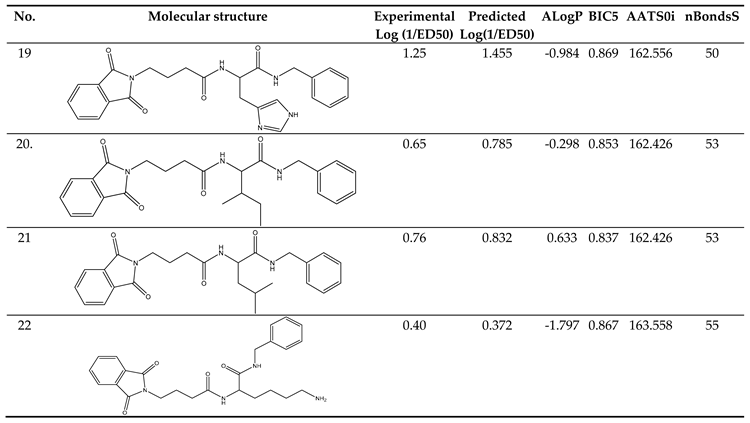
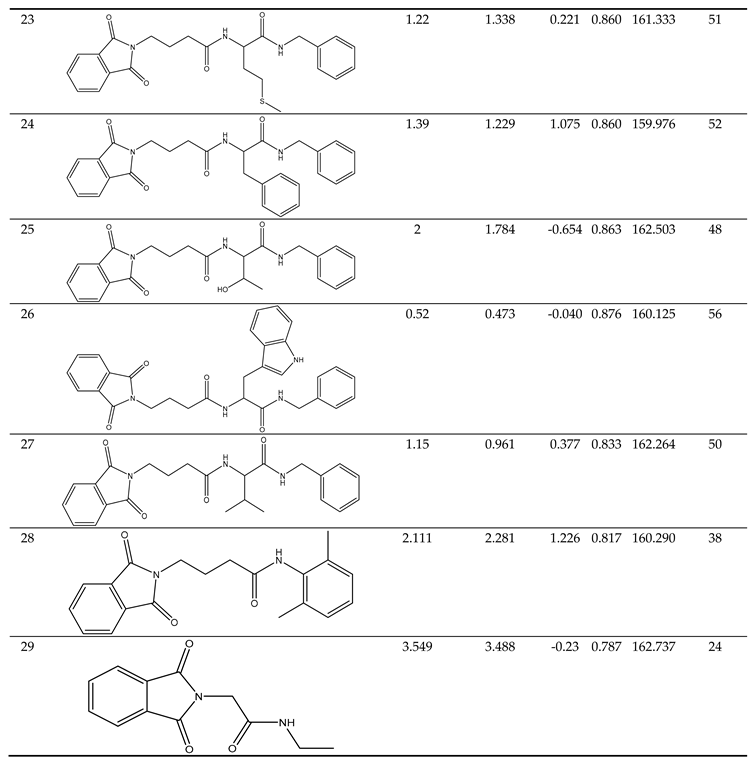
Clarification of Descriptors
The Descriptors included in the models carried specific physical and chemical significance. For instance, one of the descriptors, ALogP, represents the Ghose-Crippen-Viswanadhan octanol-water partition coefficient. This coefficient was incorporated into the model based on an assessment N-octanol-water partition coefficient calculation method. Data from 12,908 chemical compounds were used in the neural network ensemble study that was used to build this approach. The number of hydrogen and other atoms, as well as atom and bond-type E-state indices, were all taken into consideration while calculating ALogP. [
31]
BIC5 was another descriptor in the model. This descriptor exhibited a negative contribution to the anticonvulsant activity, showing that a decrease in its value increases the tested compounds' anticonvulsant efficacy. This particular descriptor quantifies the Bond Information Content index. [
32]
AATS0i was the model's additional descriptor. It contributed negatively to the anticonvulsant activity, This indicates that decreasing its value enhances the anticonvulsant activity of the compounds being investigated.
nBondsS was another descriptor in the model. It made a negative contribution to the anticonvulsant activity, demonstrating that a decrease in its value improves the tested compounds' anticonvulsant effect. This descriptor is associated with the detection of the number of single bonds. [
33]
Prediction of ADMET by Computational Analysis
With the use of the pkCSM server [
34], substances' pharmacokinetic characteristics were identified. In order to ensure that the ADMET parameters fell within their expected ranges, the parameters were computed.
Table 7 displayed the ADMET characteristics of docked molecules.
In the (
Table 7) High Caco-2 permeability was predicted for the chosen compounds 12, 13 and 16. Intestinal absorption patterns for all the compounds were predicted to be excellent. The finding suggests that the compounds have good skin permeability values
Table 7.
All the Compounds has demonstrated good dispersion volume. A substance with log BB > 0.3 has high blood-brain barrier membrane permeability and can easily cross it, whereas a molecule with log BB -1 will find it challenging to do so. Crossing the blood-brain barrier was anticipated for each of the chosen compounds shown in
Table 7. The findings indicate that the ADMET toxicity test did not reveal any toxicity for any of the chosen compounds. The results for the ADMET properties that were anticipated were fairly positive, and more in-depth research will undoubtedly be useful in forecasting the fate of the compounds.
Table 7.
Prediction of ADMET.
Table 7.
Prediction of ADMET.
| Model Name |
Predicted Value Compound 2 |
Predicted Value Compound 11 |
Standard value |
| Distribution |
|
|
|
| Caco2 Permeability |
1.106 |
0.972 |
>0.90 |
| Skin Permeability |
-3.105 |
-3.17 |
< -2.5 |
| Intestinal absorption (human) |
92.774% |
90.873% |
Poor absorption If less 30% |
| Distribution |
|
|
|
| VDss(human) |
-0.123 |
0.148 |
Minimum if Less - 0.15 |
| BBB Permeability |
-0.317 |
-0.26 |
> 0.45 |
| Toxicity |
|
|
|
| Oral Rat Acute Toxicity (LD50) |
2.567 |
2.312 |
Numeric (mol/kg) |
| Oral Rat Chronic Toxicity (LOAEL) |
1.226 |
1.583 |
Numeric (mol/kg bw/day) |
Computational Investigations of Molecular Dynamics
The molecular docking analyses provided insights into the intricate binding modes between the protein and the inhibitor. However, to assess even the slightest discrepancies, molecular dynamics simulations become essential. [
35,
36] Leveraging the most favourable interactions and energetically stable docked conformation obtained from Auto Dock results, we conducted an in-depth exploration of compound 11 within a solvent system. As a reference compound, we selected phenytoin, a GABA-AT inhibitor.
For the molecular dynamics simulation, we identified the best active conformation based on hydrogen bond interactions, root mean square deviation (RMSD), and energy values. A 5 nanosecond (ns) simulation was conducted to evaluate the stability of the protein-inhibitor complex, followed by a comprehensive analysis of the simulation results. [
37]
Analysis of Compound 11_1OHV Complex Simulation.
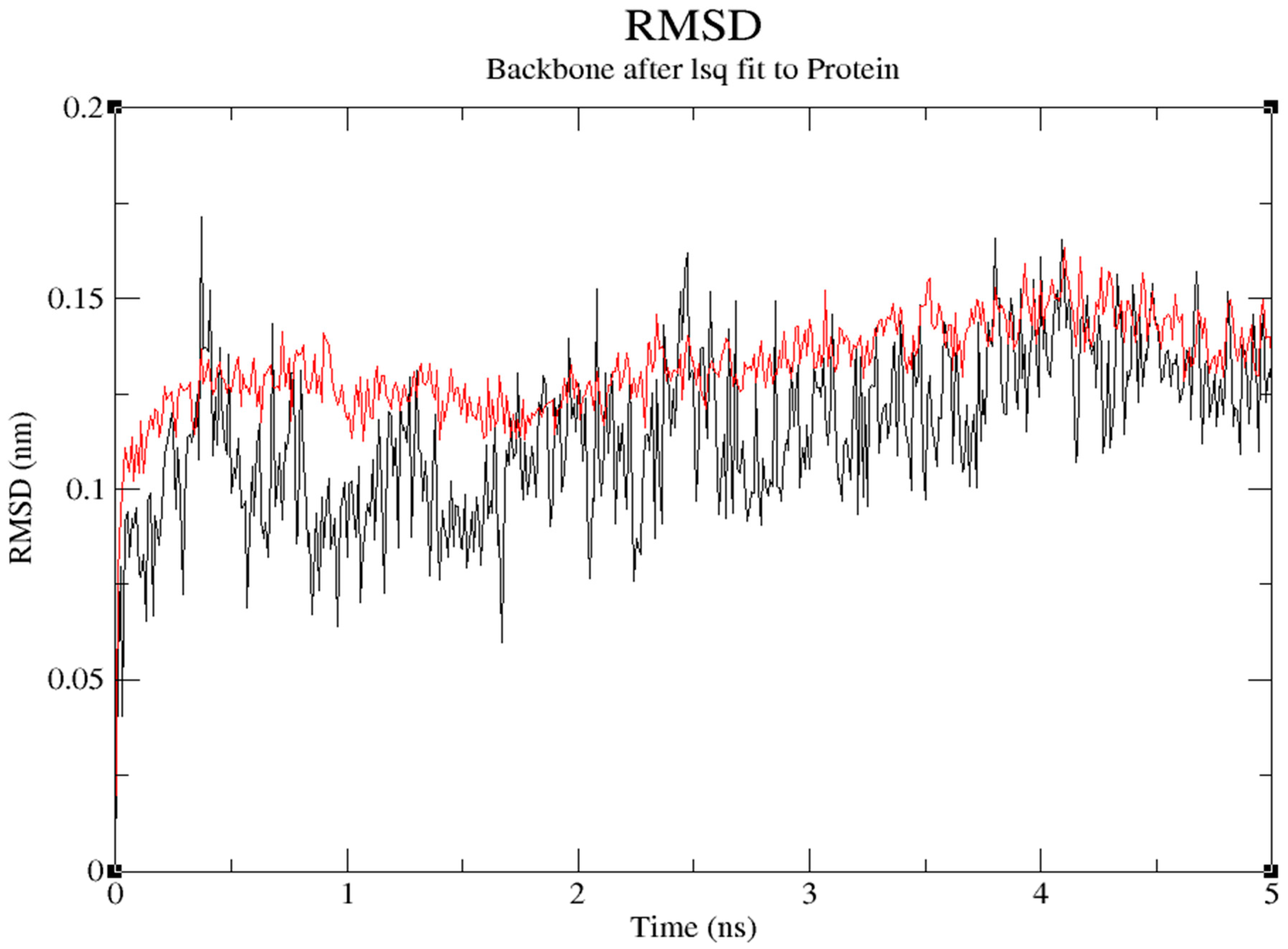
The protein's backbone atom's RMSD value increased in relation to its initial position to 0.136nm for first 0.371ns and decreased to 0.0.114nm at 1.76ns and reached maximum of 0.162nm at 4.102ns in the trajectory. RMSD number for the ligand with protein for its initial position increased to 0.118 nm for first 0.242ns and decreased to 0.059nm at 1.676ns and reached maximum of 0.171nm at 0.371ns in the trajectory. (
Figure 12)
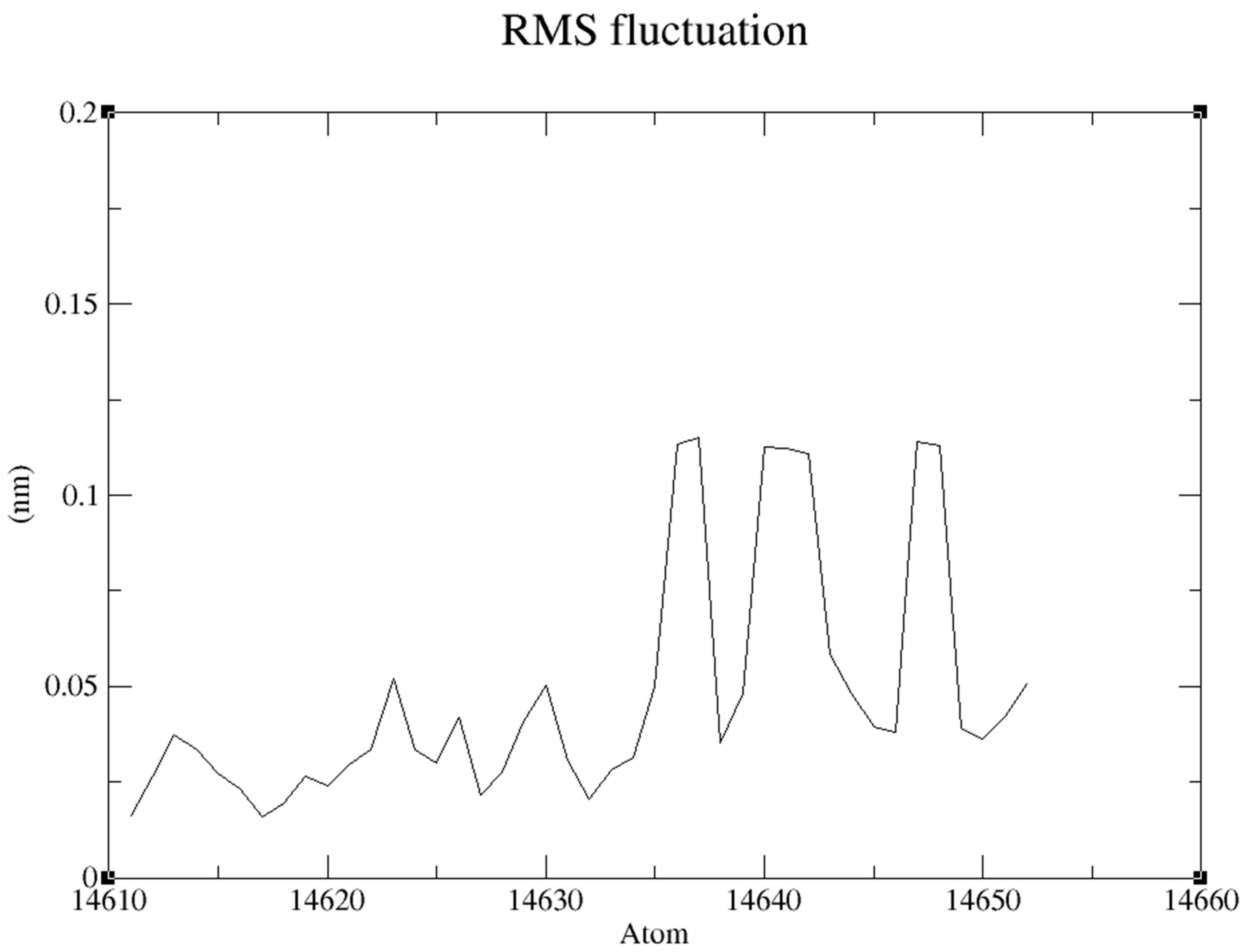
RMSF “root mean square fluctuation” of the Ligand produced an average at about 0.049nm and maximum at 0.114nm an appropriate foundation for further inquiry is established during the simulation. (
Figure 13)
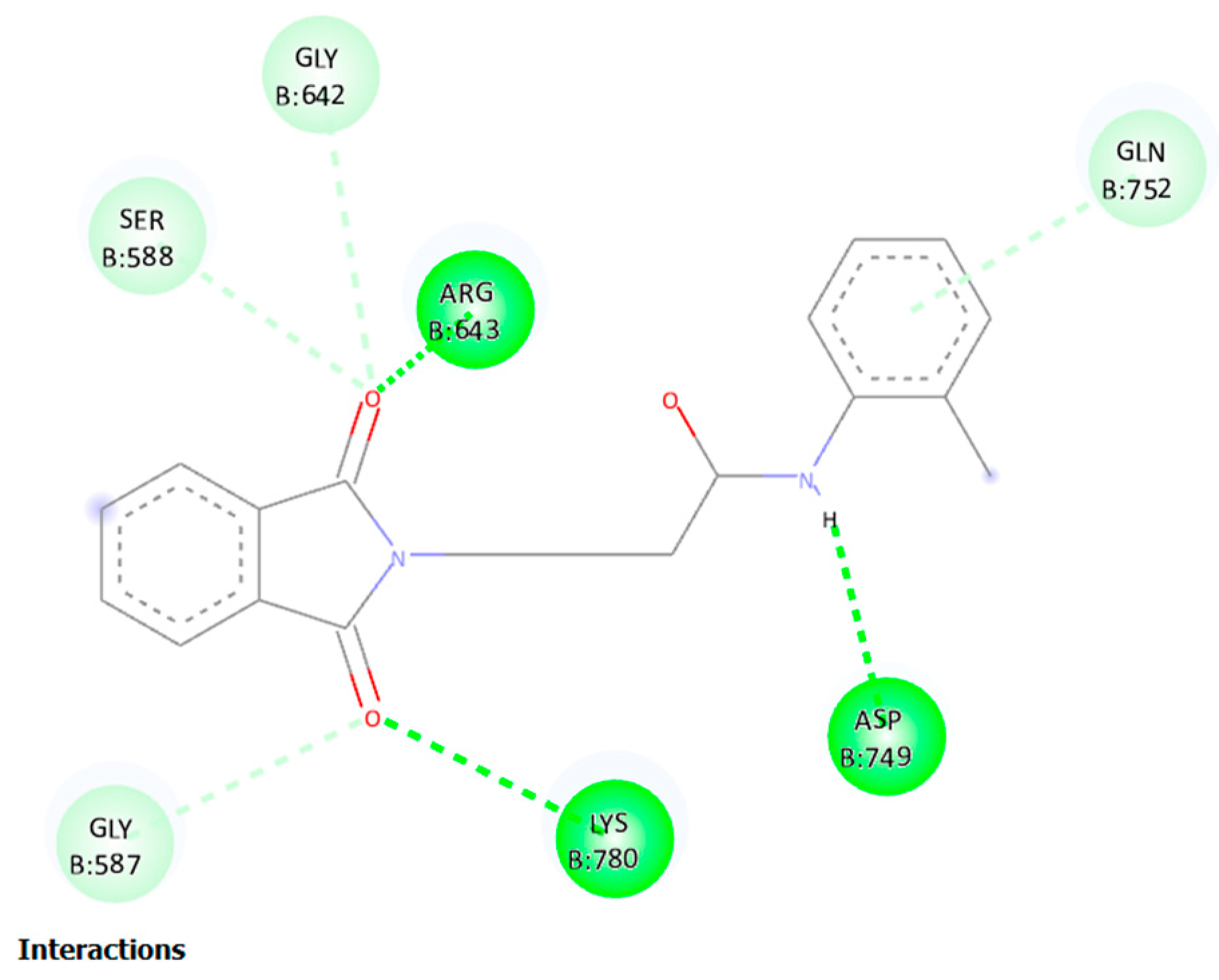
2D structural analysis of MDS Examination of the compound 11 MD presented hydrogen bonds, Carbon and Pi-sigma interactions. within hydrogen bonds the stable regions Ser, Lys, Asp, Gly, Arg, Gln. The compound 11 and the residues Gly, Gln, and Ser interact via carbon and pi-donor hydrogen bonds (
Figure 14 bottom). The amino acid residues Ser, Lys, Asp, Gly, Arg, and Gln exhibited notable interactions within the binding site, and they were deemed crucial for maintaining the stability of compound 11 within the cavity. Compound no. 11 consistently formed hydrogen bonds at a high frequency during the entire simulation. In particular, it was observed that Lys 329 played a significant role in facilitating the adoption of an active conformation by compound no.11.
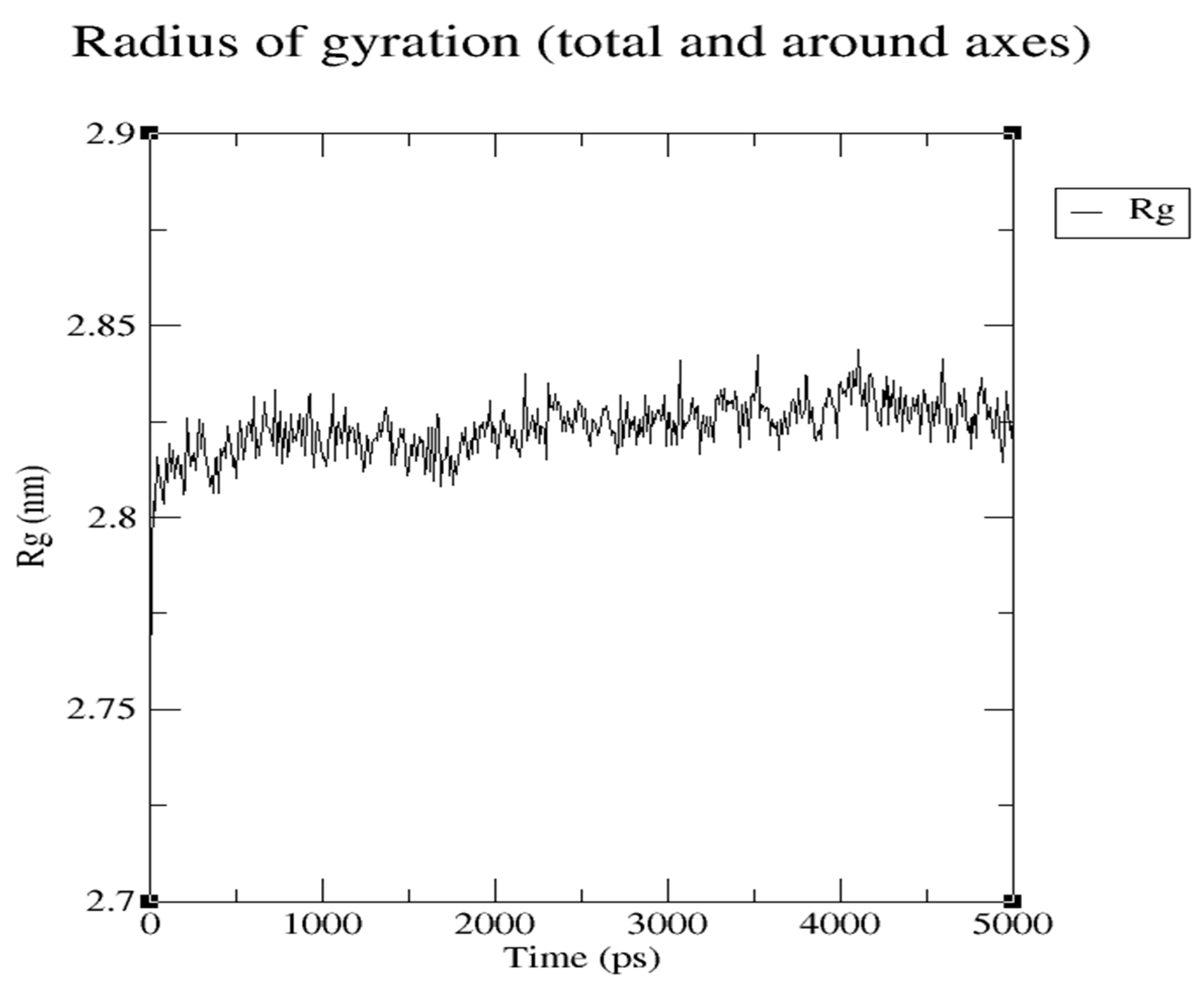
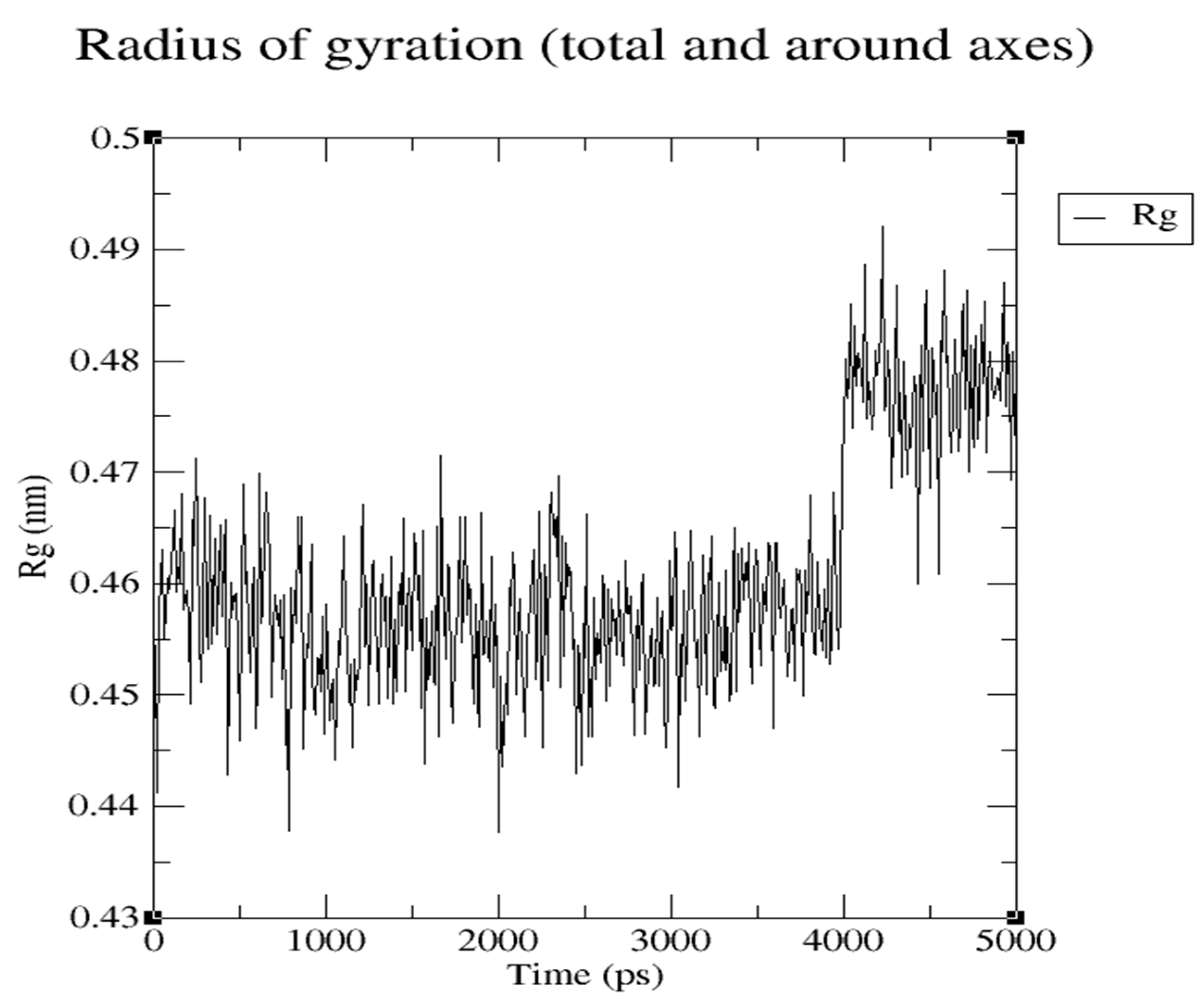
Were radius of gyration represents the hardness of protein and ligand (
Figure 16 and
Figure 17)
Conformational Change Analysis by Post-Docking
In order to examine residue interactions, hydrogen bonds, and binding energy values, molecular docking studies were conducted on the above conformations. The original GABA-AT structure, as found in the protein database (
www.rcsb.org) with the PDB structure 10HV, served as the starting point. For protein preparation, polar hydrogens were added while applying Kollman's united atomic charges. Polar hydrogens were added and Gasteiger charges were assigned in the case of the ligand file. To facilitate the molecular docking process, the Autogrid feature was utilized, enabling the selection of an active site and the customization of the grid size to 60 × 60 × 60 points with a spacing of 0.375 Å. Due to this configuration, the energy map could be calculated using a distance-dependent dielectric constant function.
The active binding site for the enzyme is shown in the designated grid box, providing ample space for ligand rotation and translation. To explore various ligand conformation poses within the active site of GABA-AT, orientations were employed using the Lamarckian genetic algorithm. The docking performance utilized default parameters, with the addition of specific metal parameters as required. The search algorithm analysis was employed to generate ligand poses within the GABA-AT binding site, considering both roto-translational movement and the ligand's internal degrees of freedom. With the help of the Discovery Studio non-bonding interaction visualizer, the best positions were selected based on binding affinity. To assess the accuracy of RMSD from the docking simulation values between the reference and predicted structures were calculated. This analysis helped confirm whether or not the docking simulation was able to accurately identify docking poses.
Simulations of Molecular Dynamics
Simulations of molecular dynamics were conducted using Gromacs 2023 to explore the conformational changes in the protein-ligand complex within a solvent system. For the simulation, the complex was positioned in the centre of an orthorhombic cubic box, and TIP3P water molecules were added to fill the box, with additional buffer space between the box edges and protein atoms to facilitate dynamic simulations.
To account for boundary conditions, the box volume was adjusted based on the type of complex, and To balance the system, counter ions such as Na+ and Cl- were injected at random.
The simulations were carried out under the Berendsen NVT ensemble, maintaining a temperature of 10 K initially to restrain heavy atoms in the solute. Then, under the isothermal isotropic ensemble, At 300 K and 1 atm, simulations of molecular dynamics were run with a thermostat relaxation time of 10 ps (NPT). To control temperature and pressure during MD simulations, By using the Martyna-Tobias-Klein barostat method and the Nose-Hoover thermostat, conditions were kept at 300 K and 1 atm, respectively.
Throughout the MD simulations, trajectory frames were collected and analyzed using simulation interaction diagrams to assess fluctuations and gain insights into the system's behaviour.
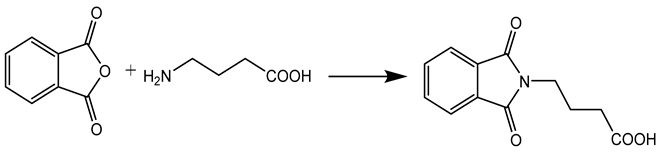
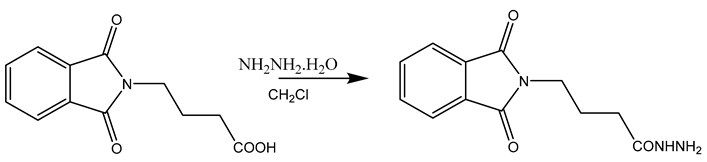
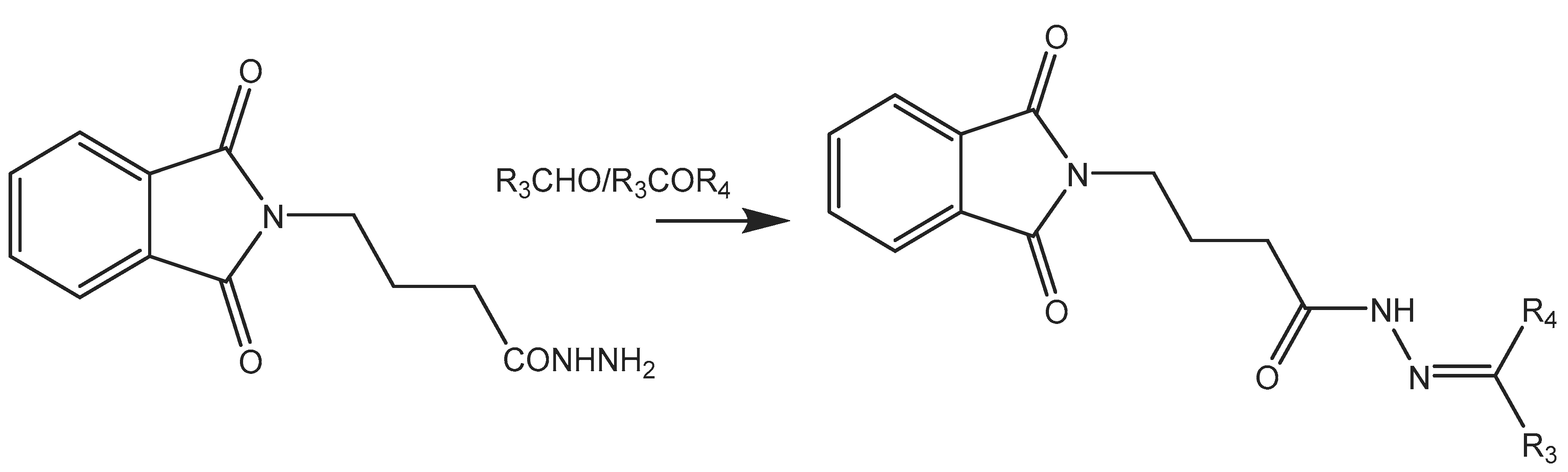
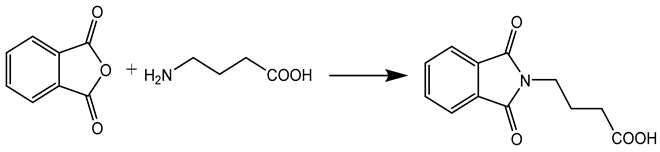
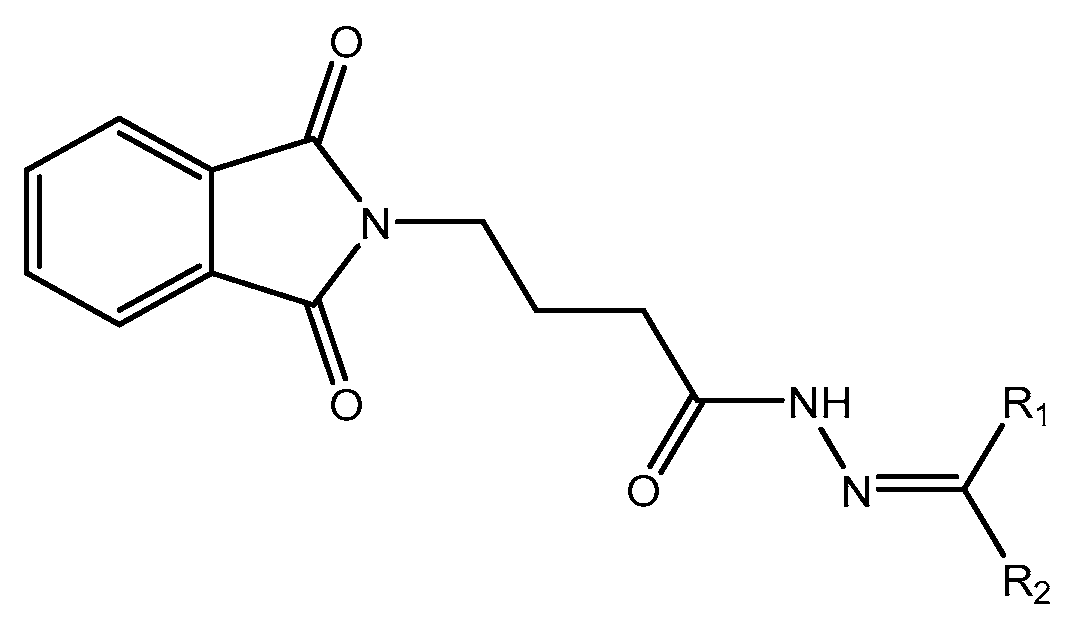
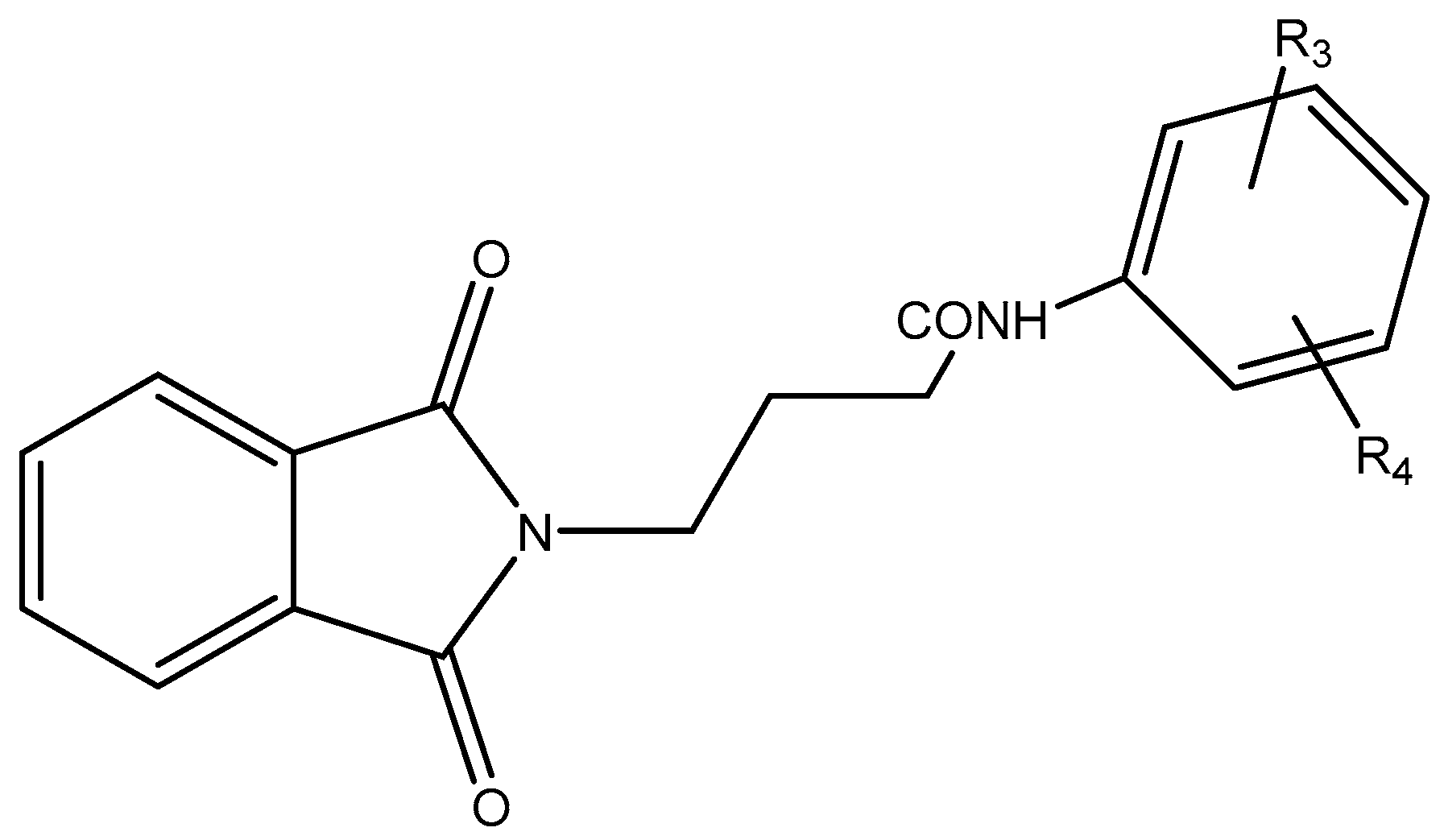
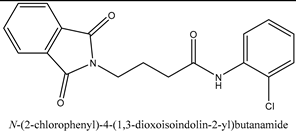
In this work, a series of N,N-phthaloyl GABA was produced and its anticonvulsant effectiveness was assessed. The N,N-phthaloyl GABA was prepared in the manner shown in Scheme. 1-(2-chlorophenyl)-4-(1,3-dioxoisoindolin-2-yl)butanamide that has been substituted. After condensation with aniline and aldehyde in dichloromethane, the 4-(1,3-dioxoisoindolin-2-yl)butanoic acid derivatives produced substituted N,N-phthaloyl GABA. It then further reacted with the appropriate aldehydes, resulting in the production of phthalimide derivatives, namely N'-(2-chlorobenzylidene)-4-(1,3-dioxoisoindolin-2-yl)butanehydrazide. We performed thorough characterizations of every chemical we synthesised using spectral, analytical, and physical analysis. infrared (IR) spectra showed signals from amide bonds between 3300 and 3200 cm-1, a C=N peak between 1620 and 1590 cm-1, and NH stretching vibrations at 3450 cm-1. Further information was also revealed by the 1H-NMR spectra, which showed N-H signals in the form of a singlet at 3324 ppm and 8.35 ppm, indicating the conformation of these molecules (
Table 8)
N-(2-chlorophenyl)-4-(1,3-dioxoisoindolin-2-yl) butanamide, white flaky crystal, m.p:208-210℃, % yield: 70, Rf:0.62, [(CHCl3: MeOH) (9:1), Molecular Formula: C16H18ClN, Mol. wt: 259.77 g/mol, IR: 3269 (NH Stretch), 2925(C-H Stretch Aromatic), 1712 (C=O Stretch), 1530 (C=C Aromatic), 1292 (CN stretch) 717 (C-Cl Stretch), 1H NMR: (CDCL3, 400MHz, δppm); 8.28(s, 1H, CONH), 7.04-7.74 (m, 8H, Aryl-H), 3.85-3.88 (t, 3H, CH2), 3.76-3.79 (t, 1H, CH2), 2.15-2.21 (m, 2H, CH2).
The test compounds were administered intraperitoneally to mice at doses of 30, 100, and 300 mg/kg. The assessment of anticonvulsant activity was conducted 30 minutes and 4 hours after drug administration.
Anti-Convulsant Activity of Active Synthesized Compounds:
Latency in Development of Seizures
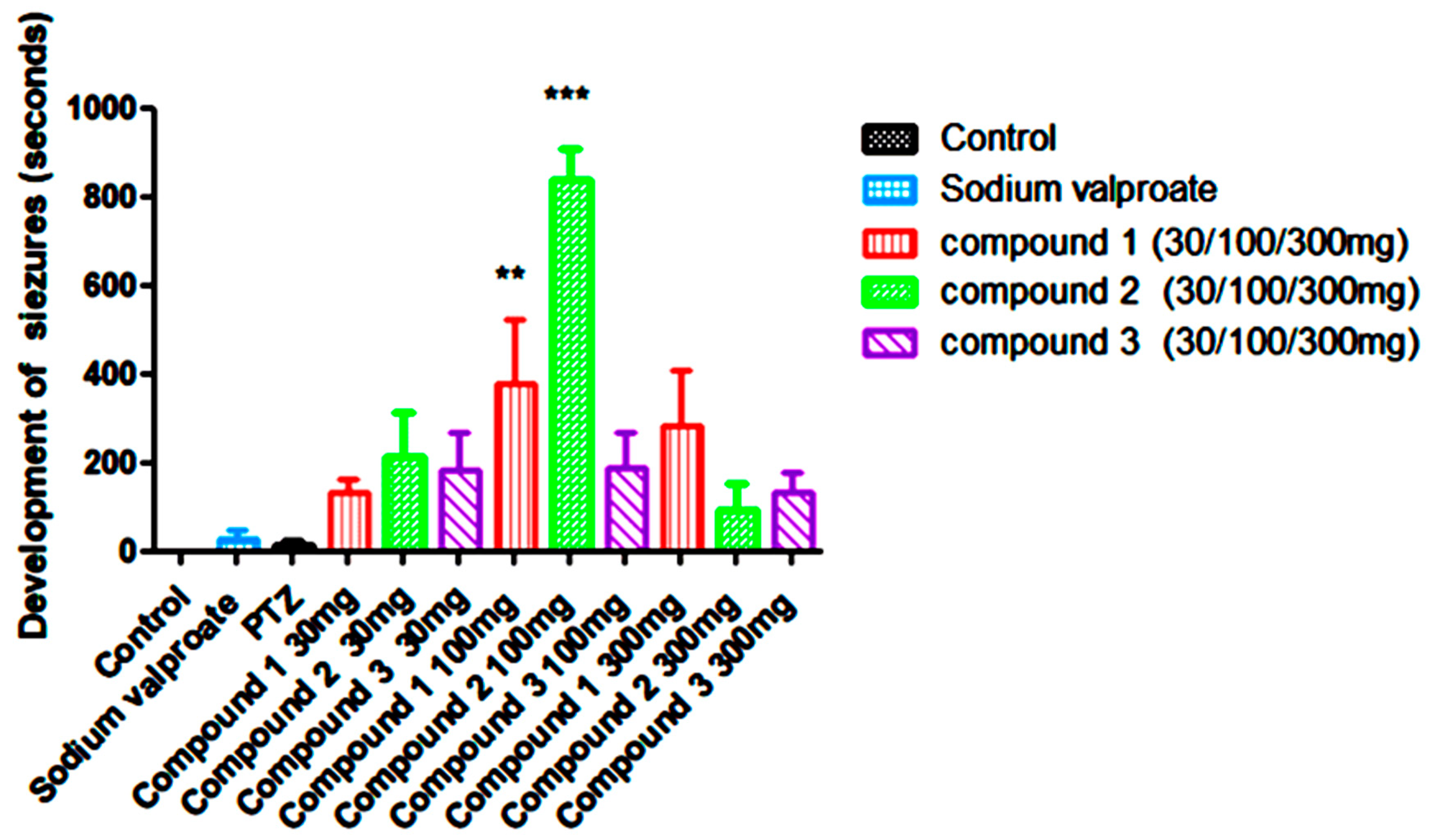
In this study, subcutaneous injection of pentylenetetrazol (ScPTZ, 80mg/kg) was administered to develop convulsions in mice. Results obtained from the study indicated that Compound 1 (Sch1-1) at the doses of 30mg/kg, 100mg/kg, and 300mg/kg increases the latency to develop seizures significantly in dose dependent manner in mice in compared to standard i.e. Sodium valproate and PTZ. Compound 2(Sch-1-3) at the doses of 30mg/kg, 100mg/kg showed significant increase in latency in development of seizures but at 300mg/kg latency was decreased compared to 30 and 100mg/kg yet it is much better compared to standard drug Sodium Valproate and PTZ. Compound 3 (Sch1-6) At 30mg/kg and 100mg/kg exhibited quite similar latency in development of seizures and at the dose of 300mg/kg the latency was reduced compared to initial two doses ie. 30mg/kg, and 100mg/kg but these results were found to be better in comparison to standard group of Sodium Valproate and PTZ (
Figure 19).
Table 9.
Latency in the Development of Seizures (time Second) (Data represented below as mean±SEM).
Table 9.
Latency in the Development of Seizures (time Second) (Data represented below as mean±SEM).
| Control |
Sodium valproate |
PTZ |
Compound 1 (30mg) |
Compound 2 (30mg) |
Compound 3 (30mg) |
Compound 1 (100mg) |
Compound 2 (100mg) |
Compound 3 (100mg) |
Compound 1 (300mg) |
Compound 2 (300mg) |
Compound 3 (300mg) |
| 0 |
30±50.199 |
12.3±21.058 |
132.6±70.47 |
212.16±241.94 |
181.5±208.91 |
454.8±356.59** |
836.1±79.5*** |
186±207.36 |
283.16±311.0 |
93±146.171 |
132±1111.864 |
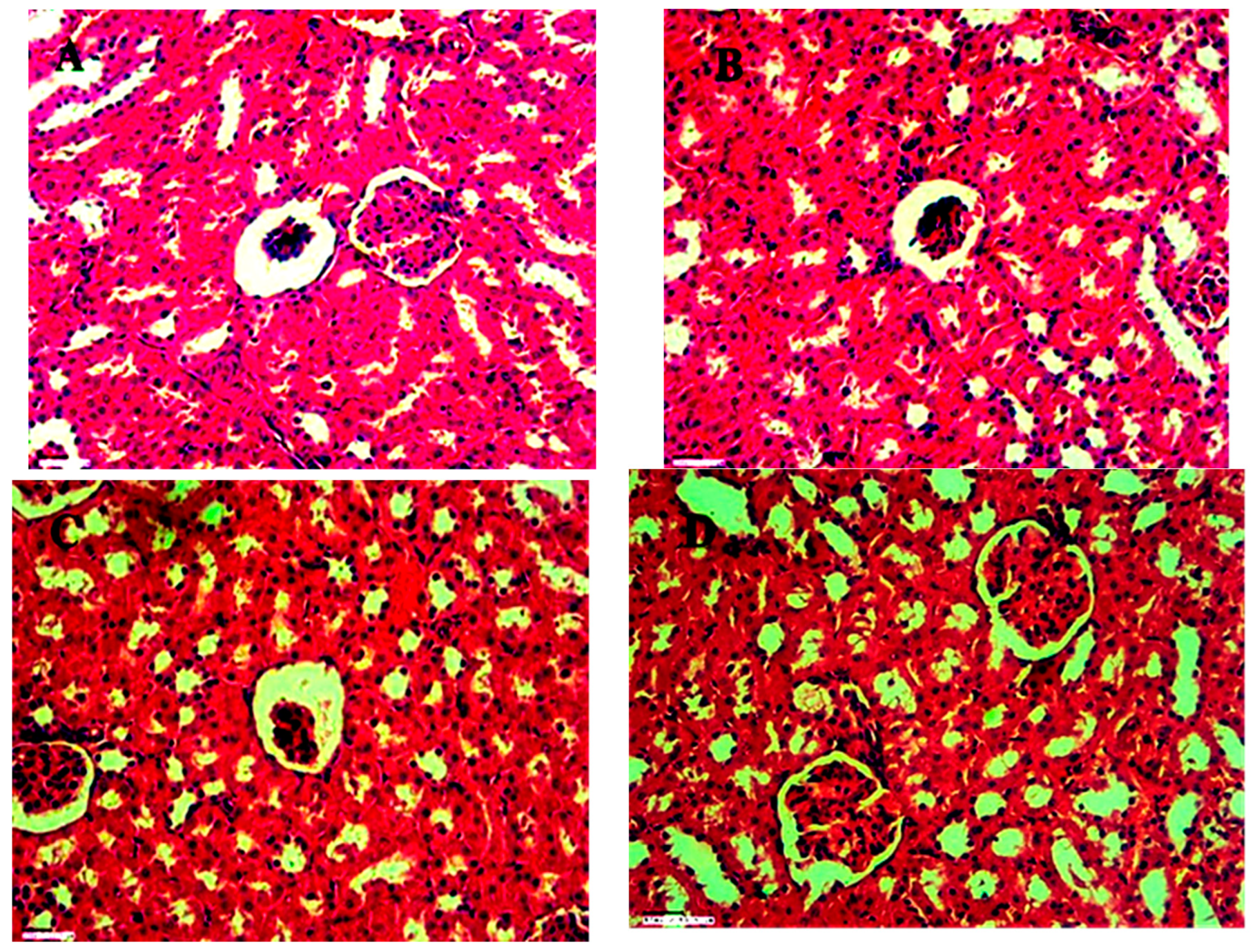
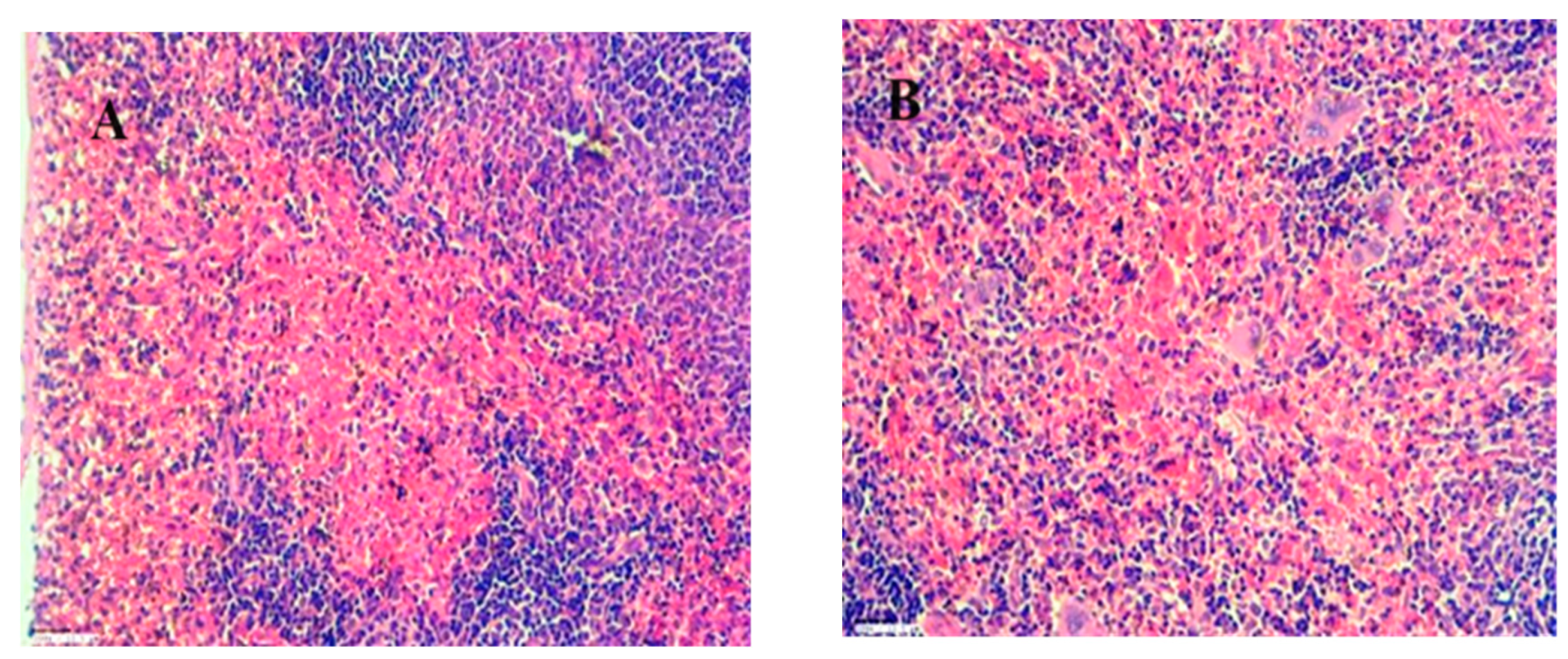
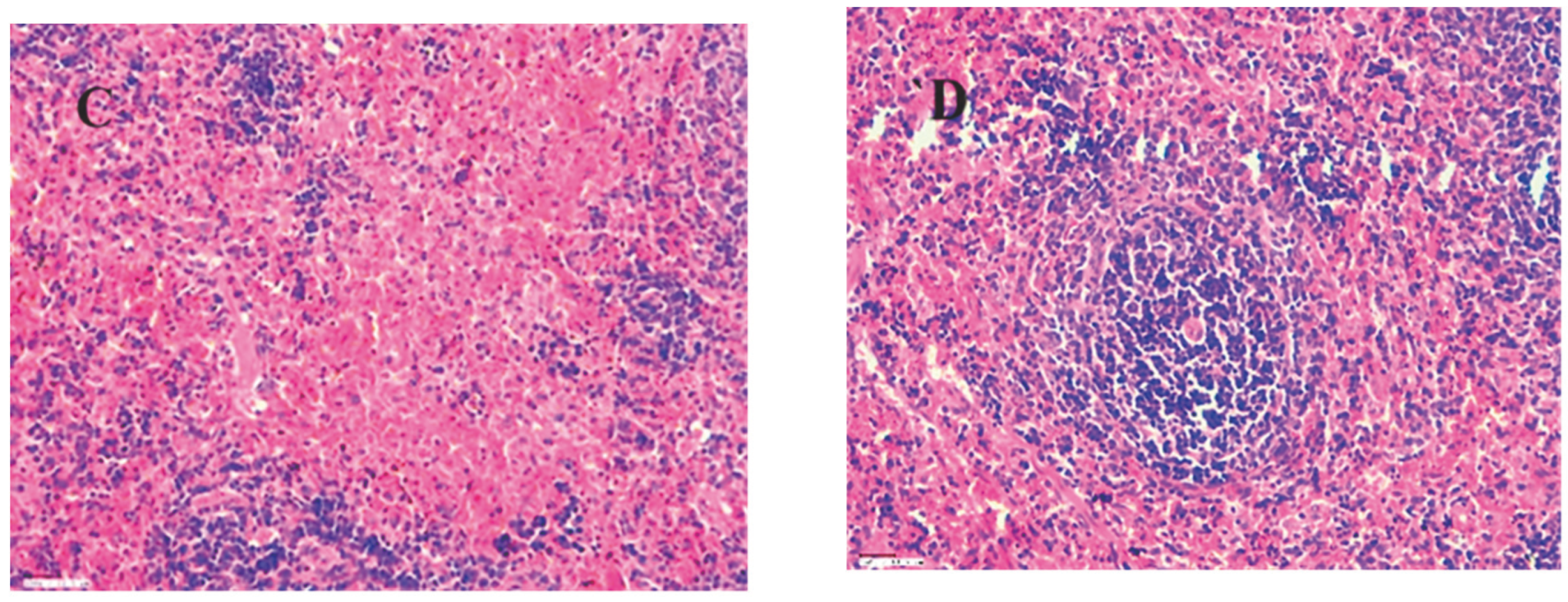
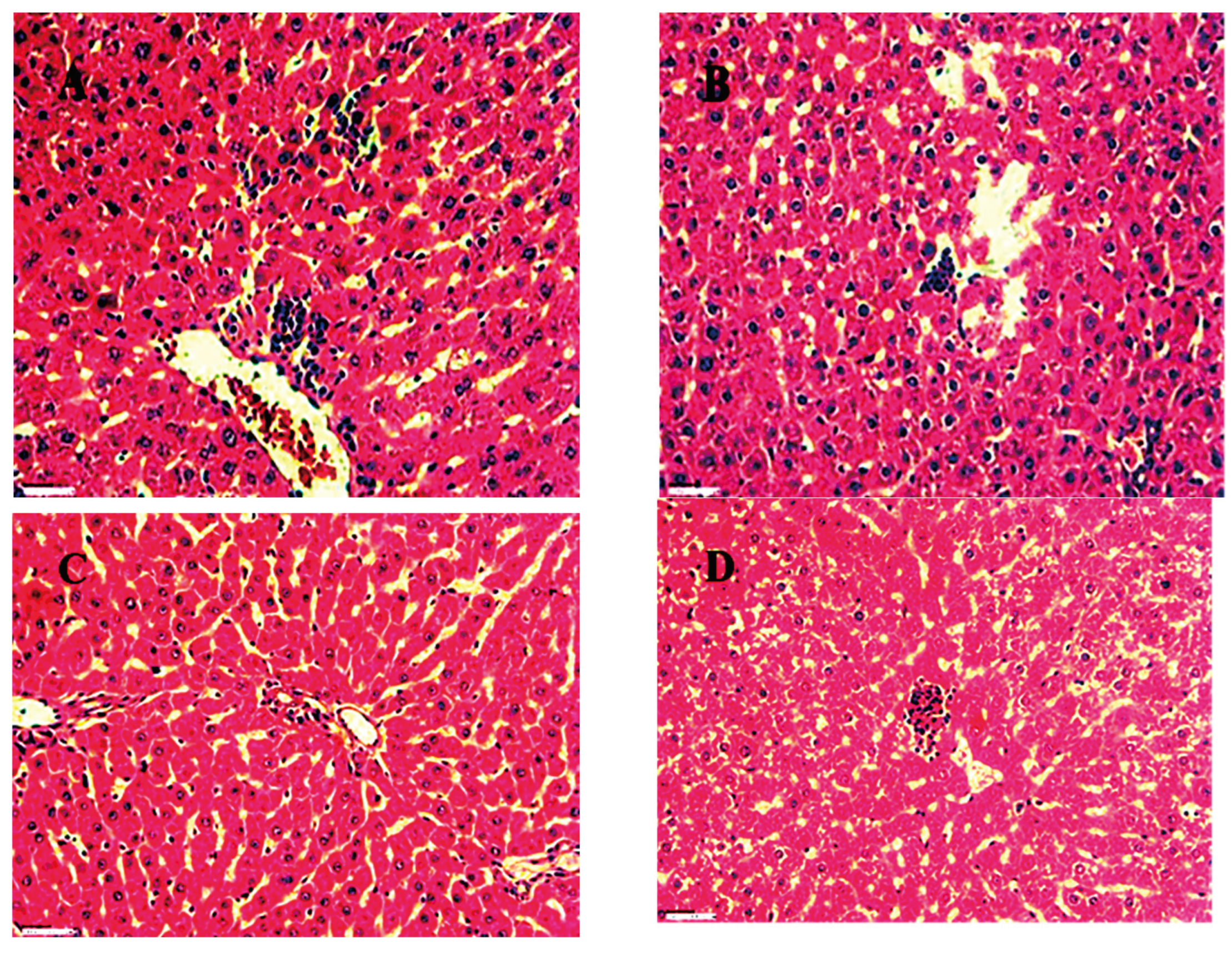
Histopathology result for kidney, spleen and Liver of group 1,2,3 and 4 (control) are shown in
Figure 19,
Figure 20 and
Figure 21 at 45X. for acute oral toxicity.
Acute oral Toxicity Report
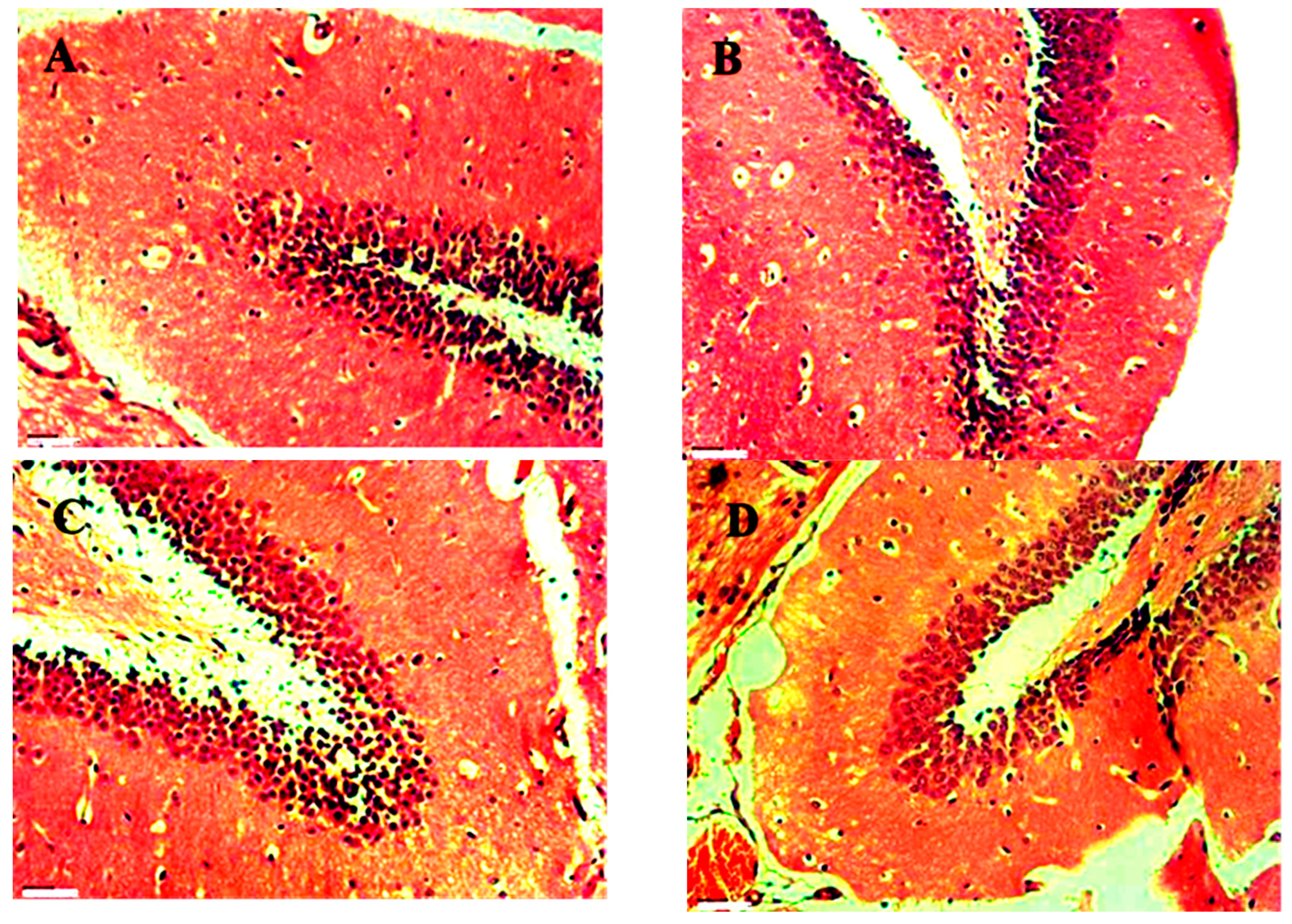
Histopathology Study for Anticonvulsant Activity
Histopathology result for Brain of group standard, PTZ, control and Compound no. 11 are shown in
Figure 22,
Figure 23, Figure 24 and Figure 25 at 45X and 100X. for acute oral toxicity.