Submitted:
24 March 2024
Posted:
25 March 2024
Read the latest preprint version here
Abstract
Keywords:
1. Introduction
2. Results and Discussion
2.1. Model Performance Overview
2.1.1. Training Performance
2.1.2. Testing Performance
2.1.3. Error Metrics
2.2. Model Optimization
2.3. Comparative Analysis: QSAR Modeling
2.4. Model Interpretation
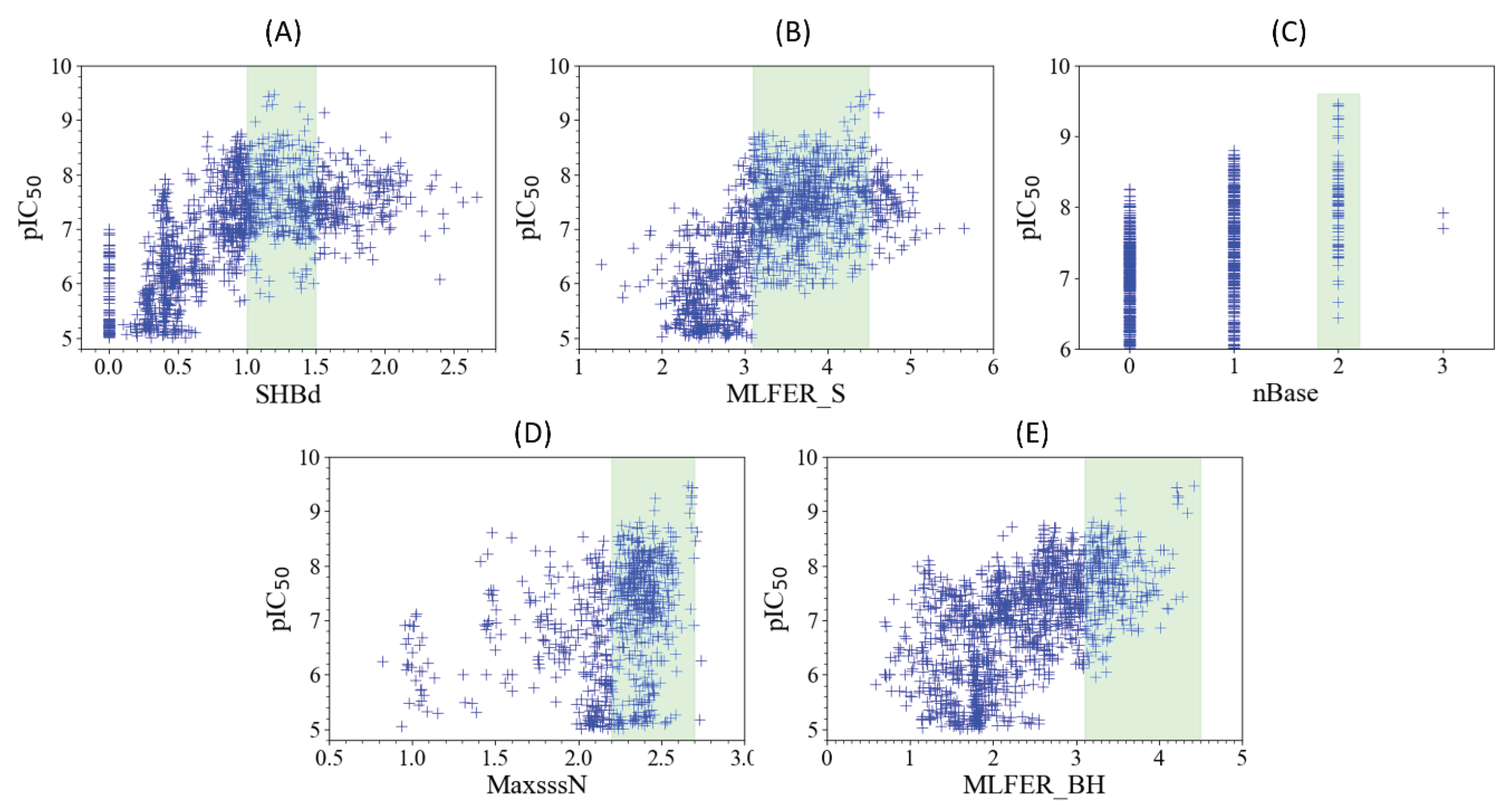
2.4.1. SHBd
2.4.2. MLFER_S
2.4.3. nBase, MaxsssN and MLFER_BH
2.5. Novel FLT3 Inhibitors Identified by Ligand-Based Screening
2.6. Script-Like Tool Description
3. Materials and Methods
3.1. Data Curation
3.2. Molecular Descriptor Calculation
3.3. Benchmarking Machine Learning Methods with External Validation
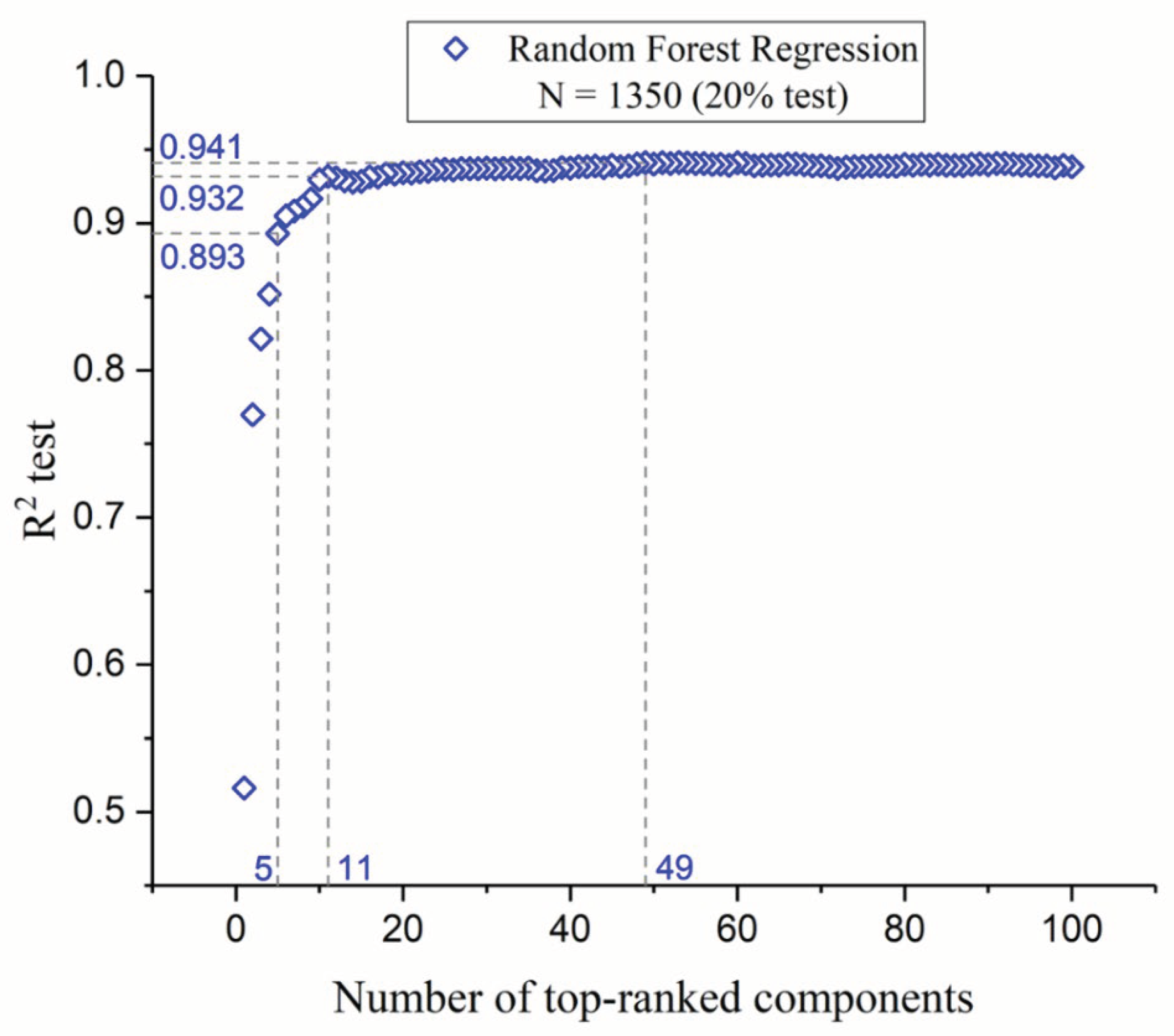
3.4. Component Optimization through Feature Selection
3.4.1. Individual Descriptor Evaluation
3.4.2. Analysis and Feature Selection Process
3.5. Internal Validation
3.6. Ligand-Based Virtual Screening
4. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Birg, F.; Courcoul, M.; Rosnet, O.; Bardin, F.; Pébusque, M.J.; Marchetto, S.; Tabilio, A.; Mannoni, P.; Birnbaum, D. Expression of the FMS/KIT-like gene FLT3 in human acute leukemias of the myeloid and lymphoid lineages. Blood 1992, 80, 2584–2593. [Google Scholar] [CrossRef]
- Small, D. FLT3 mutations: biology and treatment. Hematology / the Education Program of the American Society of Hematology. American Society of Hematology. Education Program 2006, 2006, 178–184. [Google Scholar] [CrossRef] [PubMed]
- Barley, K.; Navada, S.C. Acute myeloid leukemia. Oncology 2019, 373, 308–318. [Google Scholar] [CrossRef]
- Kazi, J.U.; Rönnstrand, L. FMS-like tyrosine kinase 3/FLT3: From basic science to clinical implications. Physiological Reviews 2019, 99, 1433–1466. [Google Scholar] [CrossRef]
- Kantarjian, H.M.; Short, N.J.; Fathi, A.T.; Marcucci, G.; Ravandi, F.; Tallman, M.; Wang, E.S.; Wei, A.H. Acute Myeloid Leukemia: Historical Perspective and Progress in Research and Therapy Over 5 Decades. Clinical Lymphoma, Myeloma and Leukemia 2021, 21, 580–597. [Google Scholar] [CrossRef] [PubMed]
- Wei, A.H.; Tiong, I.S. Midostaurin, enasidenib, CPX-351, gemtuzumab ozogamicin, and venetoclax bring new hope to AML. Blood 2017, 130, 2469–2474. [Google Scholar] [CrossRef]
- Daver, N.; Wei, A.H.; Pollyea, D.A.; Fathi, A.T.; Vyas, P.; DiNardo, C.D. New directions for emerging therapies in acute myeloid leukemia: the next chapter. Blood Cancer Journal 2020, 10, 1–12. [Google Scholar] [CrossRef]
- Kantarjian, H.; Kadia, T.; DiNardo, C.; Daver, N.; Borthakur, G.; Jabbour, E.; Garcia-Manero, G.; Konopleva, M.; Ravandi, F. Acute myeloid leukemia: current progress and future directions. Blood Cancer Journal 2021, 11, 1–25. [Google Scholar] [CrossRef] [PubMed]
- Jaramillo, S.; Schlenk, R.F. Update on current treatments for adult acute myeloid leukemia: To treat acute myeloid leukemia intensively or non-intensively? That is the question. Haematologica 2023, 108, 342–352. [Google Scholar] [CrossRef]
- Kumar Kar, R.; Suryadevara, P.; Roushan, R.; Chandra Sahoo, G.; Ranjan Dikhit, M.; Das, P. Quantifying the Structural Requirements for Designing Newer FLT3 Inhibitors. Medicinal Chemistry 2012, 8, 913–927. [Google Scholar] [CrossRef]
- Shih, K.C.; Lin, C.Y.; Chi, H.C.; Hwang, C.S.; Chen, T.S.; Tang, C.Y.; Hsiao, N.W. Design of novel FLT-3 inhibitors based on dual-layer 3D-QSAR model and fragment-based compounds in silico. Journal of Chemical Information and Modeling 2012, 52, 146–155. [Google Scholar] [CrossRef] [PubMed]
- Abutayeh, R.F.; Taha, M.O. Discovery of novel FLT3 inhibitory chemotypes through extensive ligand-based and new structure- based pharmacophore modelling methods. Journal of Molecular Graphics and Modelling 2019, 88, 128–151. [Google Scholar] [CrossRef] [PubMed]
- Bhujbal, S.P.; Keretsu, S.; Cho, S.J. Design of New Therapeutic Agents Targeting FLT3 Receptor Tyrosine Kinase Using Molecular Docking and 3D-QSAR Approach. Letters in Drug Design& Discovery 2019, 17, 585–596. [Google Scholar] [CrossRef]
- Fernandes, Í.A.; Resende, D.B.; Ramalho, T.C.; Kuca, K.; Da Cunha, E.F.F. Theoretical studies aimed at finding FLT3 inhibitors and a promising compound and molecular pattern with dual aurora B/FLT3 activity. Molecules 2020, 25, 1726. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, S.; Keretsu, S.; Cho, S.J. Molecular modeling studies of n-phenylpyrimidine-4-amine derivatives for inhibiting FMS-like tyrosine kinase-3. International Journal of Molecular Sciences 2021, 22, 12511. [Google Scholar] [CrossRef] [PubMed]
- Sandoval, C.; Torrens, F.; Godoy, K.; Reyes, C.; Farías, J. Application of Quantitative Structure-Activity Relationships in the Prediction of New Compounds with Anti-Leukemic Activity. International Journal of Molecular Sciences 2023, 24, 12258. [Google Scholar] [CrossRef] [PubMed]
- Islam, M.R.; Osman, O.I.; Hassan, W.M. Identifying novel therapeutic inhibitors to target FMS-like tyrosine kinase-3 (FLT3) against acute myeloid leukemia: a molecular docking, molecular dynamics, and DFT study. Journal of Biomolecular Structure and Dynamics 2023. [CrossRef] [PubMed]
- Nasimian, A.; Al Ashiri, L.; Ahmed, M.; Duan, H.; Zhang, X.; Rönnstrand, L.; Kazi, J.U. A Receptor Tyrosine Kinase Inhibitor Sensitivity Prediction Model Identifies AXL Dependency in Leukemia. International Journal of Molecular Sciences 2023, 24, 3830. [Google Scholar] [CrossRef]
- Janssen, A.P.; Grimm, S.H.; Wijdeven, R.H.; Lenselink, E.B.; Neefjes, J.; Van Boeckel, C.A.; Van Westen, G.J.; Van Der Stelt, M. Drug Discovery Maps, a Machine Learning Model That Visualizes and Predicts Kinome-Inhibitor Interaction Landscapes. Journal of Chemical Information and Modeling 2019, 59, 1221–1229. [Google Scholar] [CrossRef]
- Zhao, Y.; Tian, Y.; Pang, X.; Li, G.; Shi, S.; Yan, A. Classification of FLT3 inhibitors and SAR analysis by machine learning methods. Molecular Diversity 2023, 1, 1–17. [Google Scholar] [CrossRef]
- Eckardt, J.N.; Bornhäuser, M.; Wendt, K.; Middeke, J.M. Application of machine learning in the management of acute myeloid leukemia: Current practice and future prospects. Blood Advances 2020, 4, 6077–6085. [Google Scholar] [CrossRef] [PubMed]
- Breiman, L. Random forests. Machine Learning 2001, 45, 5–32. [Google Scholar] [CrossRef]
- Friedman, J.H. Greedy function approximation: A gradient boosting machine. Annals of Statistics 2001, 29, 1189–1232. [Google Scholar] [CrossRef]
- Hearst, M.A.; Dumais, S.T.; Osuna, E.; Platt, J.; Scholkopf, B. Support vector machines. IEEE Intelligent Systems and their applications 1998, 13, 18–28. [Google Scholar] [CrossRef]
- Williams, C.; Rasmussen, C. Gaussian processes for regression. Advances in neural information processing systems 1995, 8. [Google Scholar]
- Altman, N.; Krzywinski, M. Ensemble methods: bagging and random forests. Nature pubchemds 2017, 14, 933–935. [Google Scholar] [CrossRef]
- Chollet, F. Keras, 2015. In: Github Repos. https://github.com/fchollet/keras.
- Paszke, A.; Gross, S.; Massa, F.; Lerer, A.; Bradbury, J.; Chanan, G.; Killeen, T.; Lin, Z.; Gimelshein, N.; Antiga, L.; et al. Pytorch: An imperative style, high-performance deep learning library. Advances in neural information processing systems 2019, 32. [Google Scholar]
- Marino, S.; Zhao, Y.; Zhou, N.; Zhou, Y.; Toga, A.W.; Zhao, L.; Jian, Y.; Yang, Y.; Chen, Y.; Wu, Q.; et al. Compressive Big Data Analytics: An ensemble meta-algorithm for high-dimensional multisource datasets. Plos one 2020, 15, e0228520. [Google Scholar] [CrossRef] [PubMed]
- Hall, L.H.; Kier, L.B. Electrotopological State Indices for Atom Types: A Novel Combination of Electronic, Topological, and Valence State Information. Journal of Chemical Information and Computer Sciences 1995, 35, 1039–1045. [Google Scholar] [CrossRef]
- Euldji, I.; Si-Moussa, C.; Hamadache, M.; Benkortbi, O. QSPR Modelling of the Solubility of Drug and Drug-like Compounds in Supercritical Carbon Dioxide. Molecular Informatics 2022, 41, 2200026. [Google Scholar] [CrossRef]
- Platts, J.A.; Butina, D.; Abraham, M.H.; Hersey, A. Estimation of molecular linear free energy relation descriptors using a group contribution approach. Journal of Chemical Information and Computer Sciences 1999, 39, 835–845. [Google Scholar] [CrossRef]
- Lin, C.; Xiaoxiao, Z. Optimizing Drug Screening with Machine Learning. 2022 19th International Computer Conference on Wavelet Active Media Technology and Information Processing, ICCWAMTIP 2022 2022. [CrossRef]
- Ibrahim, Z.Y.; Uzairu, A.; Shallangwa, G.; Abechi, S. QSAR and molecular docking based design of some indolyl-3-ethanone-α- thioethers derivatives as Plasmodium falciparum dihydroorotate dehydrogenase (PfDHODH) inhibitors. SN Applied Sciences 2020, 2, 1–12. [Google Scholar] [CrossRef]
- Lee, L.Y.; Hernandez, D.; Rajkhowa, T.; Smith, S.C.; Raman, J.R.; Nguyen, B.; Small, D.; Levis, M. Preclinical studies of gilteritinib, a next-generation FLT3 inhibitor. Blood 2017, 129, 257–260. [Google Scholar] [CrossRef] [PubMed]
- Shimada, I.; Kurosawa, K.; Matsuya, T.; Iikubo, K.; Kondoh, Y.; Kamikawa, A.; Tomiyama, H.; Iwai, Y. Patent US8969336, 2015. Available at: https://patents.google.com/patent/US8969336B2.
- PubChem Substructure Fingerprint, 2023. [Accessed December 10, 2023].
- Kim, S.; Chen, J.; Cheng, T.; Gindulyte, A.; He, J.; He, S.; Li, Q.; Shoemaker, B.A.; Thiessen, P.A.; Yu, B.; et al. PubChem 2023 update. Nucleic acids research 2023, 51, D1373–D1380. [Google Scholar] [CrossRef] [PubMed]
- Kenneth Reitz. Requests: HTTP for Humans™— Requests 2.26.0 documentation, 2021. Available at: https://docs.python-requests.org/en/latest/.
- McKinney, W.; Team, P.D. Pandas - Powerful Python Data Analysis Toolkit. https://pandas.pydata.org, 2015.
- Yap, C.W. PaDEL-descriptor: An open source software to calculate molecular descriptors and fingerprints. Journal of Computational Chemistry 2011, 32, 1466–1474. [Google Scholar] [CrossRef] [PubMed]
- Landrum, G. RDKit: Open-source cheminformatics 2022_9_5 (Q3 2022). http://www.rdkit.org, 2023. [CrossRef]
- Fabian, Pedregosa.; Gaël, Varoquaux.; Alexandre, Gramfort.; Vincent, Michel.; Bertrand, Thirion.; Olivier, Grisel.; Mathieu, Blondel.; Peter, Prettenhofer.; Ron, Weiss.; Vincent, Dubourg.; et al. Scikit-learn: Machine Learning in Python. Journal of Machine Learning Research 2011, 12, 2825–2830. [Google Scholar]
- PubChem, 2023. [Accessed November 28, 2023].
- Bajusz, D.; Rácz, A.; Héberger, K. Why is Tanimoto index an appropriate choice for fingerprint-based similarity calculations? Journal of Cheminformatics 2015, 7, 1–13. [Google Scholar] [CrossRef]
| Metrics/ML | RFR | GBR | SVM | KRR | GPR | BRF | ANN-K | ANN-P |
|---|---|---|---|---|---|---|---|---|
| R² training | 0.988 | 0.973 | 0.014 | 0.546 | 0.641 | 0.967 | 0.988 | 0.983 |
| MAE training | 0.082 | 0.126 | 0.756 | 0.489 | 0.469 | 0.136 | 0.070 | 0.082 |
| SD training | 0.102 | 0.154 | 0.933 | 0.638 | 0.526 | 0.172 | 0.101 | 0.121 |
| RMSE training | 0.102 | 0.154 | 0.941 | 0.638 | 0.568 | 0.172 | 0.103 | 0.123 |
| R² test | 0.936 | 0.939 | -0.012 | 0.592 | -0.228 | 0.931 | 0.907 | 0.895 |
| MAE test | 0.197 | 0.195 | 0.786 | 0.484 | 0.876 | 0.207 | 0.235 | 0.248 |
| SD test | 0.246 | 0.237 | 0.975 | 0.619 | 0.932 | 0.255 | 0.296 | 0.313 |
| RMSE test | 0.246 | 0.239 | 0.977 | 0.620 | 1.076 | 0.256 | 0.297 | 0.315 |
| Training set | Test set | |
|---|---|---|
| Size | 1080 | 270 |
| R² | 0.989 | 0.941 |
| MAE | 0.081 | 0.190 |
| SD | 0.101 | 0.235 |
| RMSE | 0.101 | 0.236 |
| Q2LOO | 0.926 | |
| Kara (2012) |
Shiha (2012) |
Abutayeha (2019) |
Bhujbala (2020) |
Fernandesa (2020) |
Ghosha (2021) |
This work |
|
|---|---|---|---|---|---|---|---|
| Dataset size | 67 | 72 | 93 | 63 | 40 | 40 | 1350 |
| Train set size | 51 | 25 | 76 | 45 | 28 | 30 | 1080 |
| Test set size | 16 | 47 | 17 | 18 | 12 | 10 | 270 |
| R² training | 0.956 | 0.98 | 0.86 | 0.956 | 0.80 | 0.983 | 0.989 |
| R² test | 0.891 | 0.76 | 0.57 | 0.707 | 0.80 | 0.698 | 0.941 |
| SD test | 0.435 | 0.66 | – | > 0.895 | 0.31 | 0.452 | 0.235 |
| Q2LOO | 0.747 | 0.58 | 0.65 | 0.57 | 0.60 | 0.802 | 0.926 |
| Priority | Descriptor | Name | Description |
|---|---|---|---|
| 1° | SHBd [30,31] |
Sum of E-States for (strong) hydrogen bond donors | The value is calculated as the sum of each atom capable of donating a hydrogen atom, weighted by its electronic environment and topological position (E-State). |
| 2° | MLFER_S [31,32] |
Molecular Linear-Free Energy Relation_S | Cumulative sum of the free energy contributions of solvatophilic groups in a molecule, calculated using previously established empirical values on their interactions with solvents. |
| 3° | nBase | Number of basic groups | Number of basic groups in the molecule, especially nitrogenous groups. |
| 4° | MaxsssN [30,33] |
Maximum atom-type E-State: >N- | Maximum electrotopological state present in nitrogen atoms with three single bonds. |
| 5° | MLFER_BH [32,34] |
Overall or summation solute hydrogen bond basicity | Total hydrogen bond basicity in a molecule calculated by summing the contributions of all possible hydrogen bond acceptor sites in the molecule. |
| IUPAC name | Structure | pIC50 |
|---|---|---|
| 6-Ethyl-3-[3-methoxy-4-[4-(1-methylpiperidin-4-yl)piperazin-1-yl]anilino]-5-(oxan-4-ylamino)pyrazine-2-carboxamide | 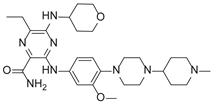 |
9.34 |
| 6-Ethyl-3-[3-methoxy-4-[4-(4-propan-2-ylpiperazin-1-yl) piperidin-1-yl]anilino]-5-(oxan-4-ylamino)pyrazine-2-carboxamide | 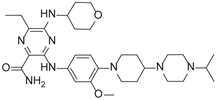 |
9.34 |
| 3-[4-[4-(1-Methylpiperidin-4-yl)piperazin-1-yl]anilino]-5-(oxan-4-ylamino)-6-propan-2-ylpyrazine-2-carboxamide | 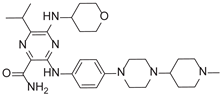 |
9.29 |
| 6-(1-Methyl-3,6-dihydro-2H-pyridin-4-yl)-3-[4-[4-(4-methylpiperazin-1-yl)piperidin-1-yl]anilino]-5-(oxan-4-ylamino)pyrazine-2-carboxamide | 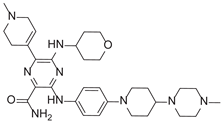 |
9.27 |
| 6-Ethyl-3-[4-[4-(4-methylpiperazin-1-yl)piperidin-1-yl]-3-propan-2-yloxyanilino]-5-(oxan-4-ylamino)pyrazine-2-carboxamide | 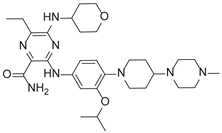 |
9.27 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




