Submitted:
03 February 2024
Posted:
05 February 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Metagenomic Sequencing and Data Analysis
2.2. Complete Genome Sequencing of PoSaV and Genomic Evolutionary Analysis
2.3. Recombination and Selective Pressure Analysis
3. Results
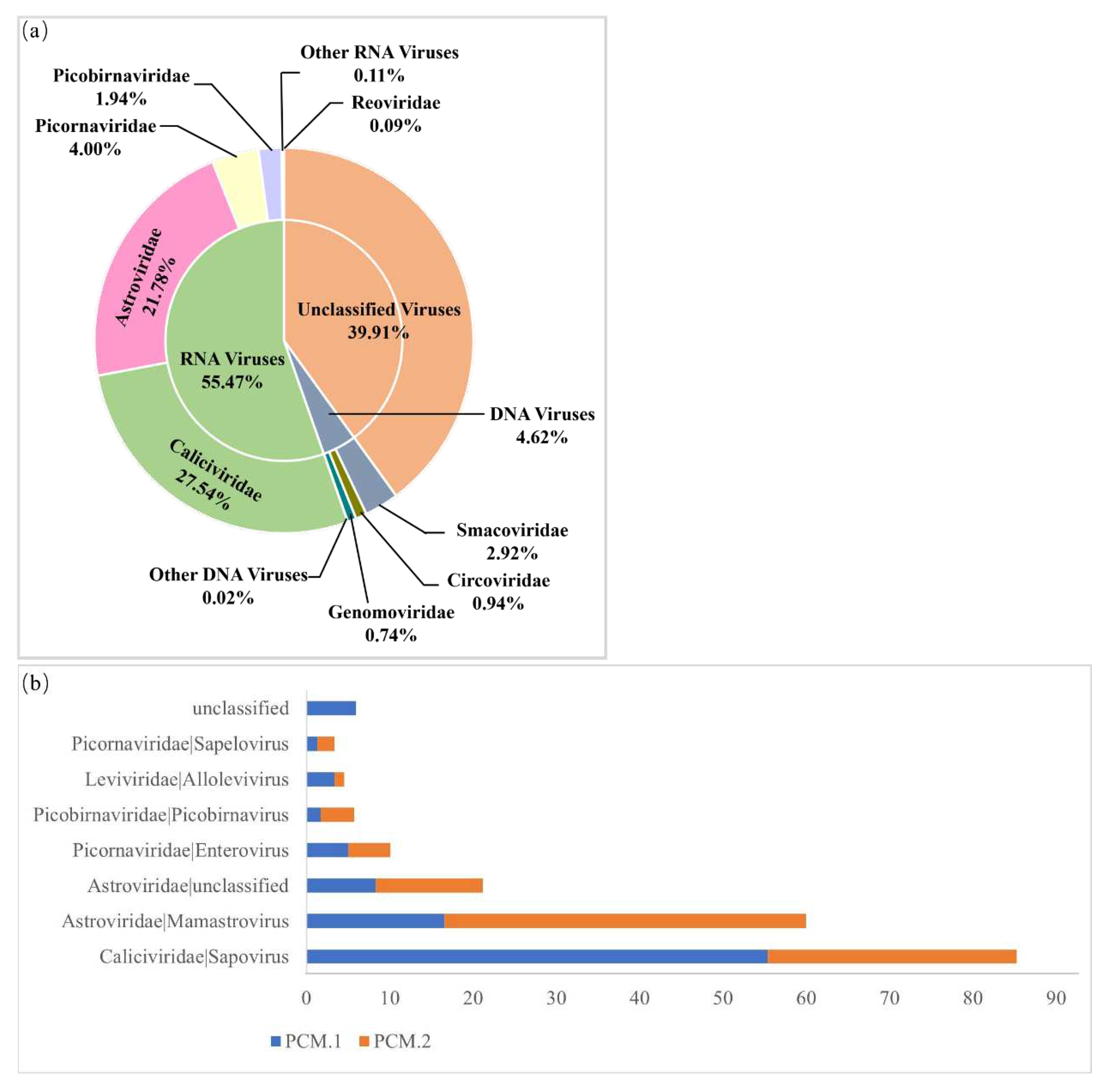
3.1. Sapovirus Is Highly Prevalent in the Piglet Diarrhea Feces
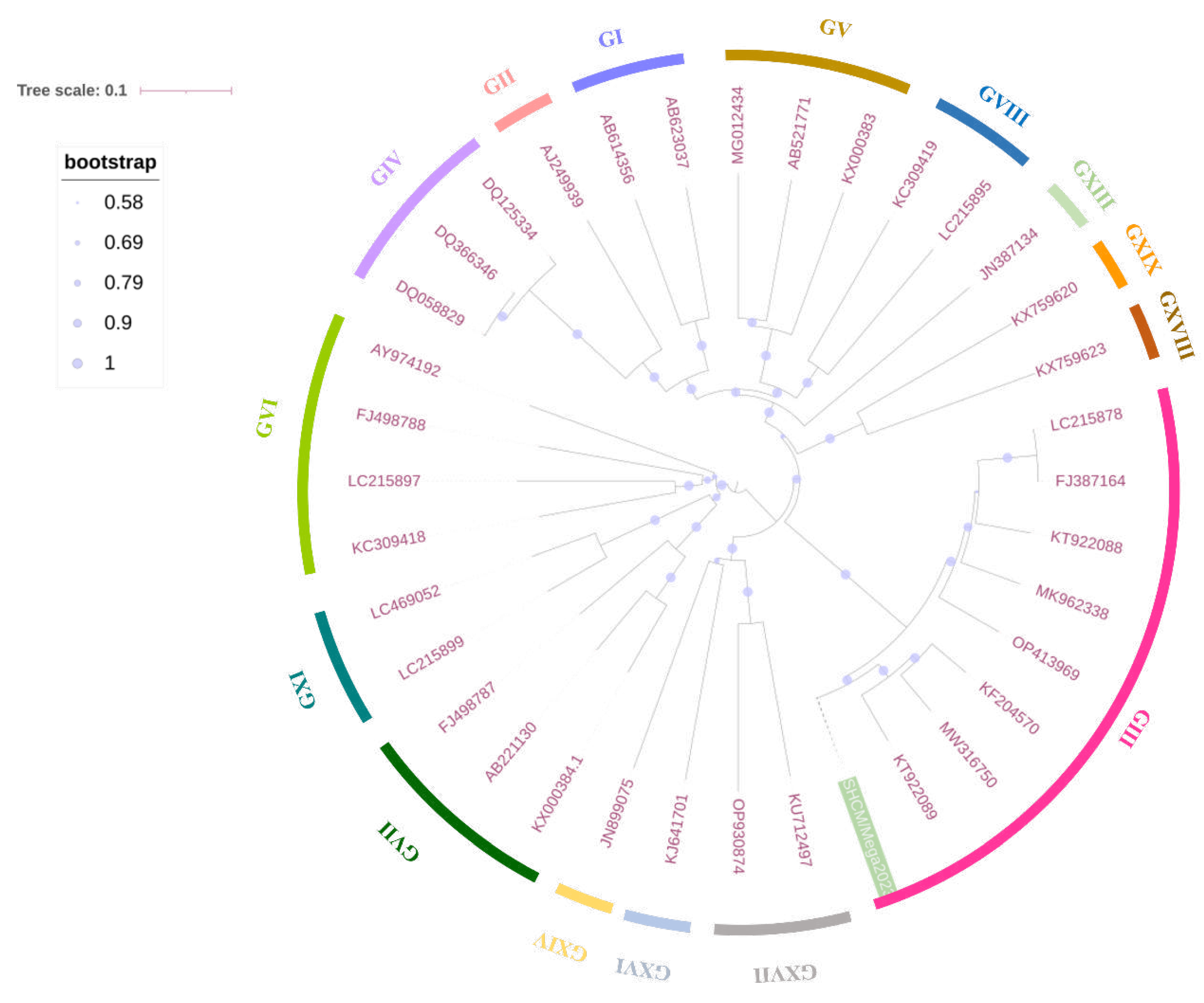
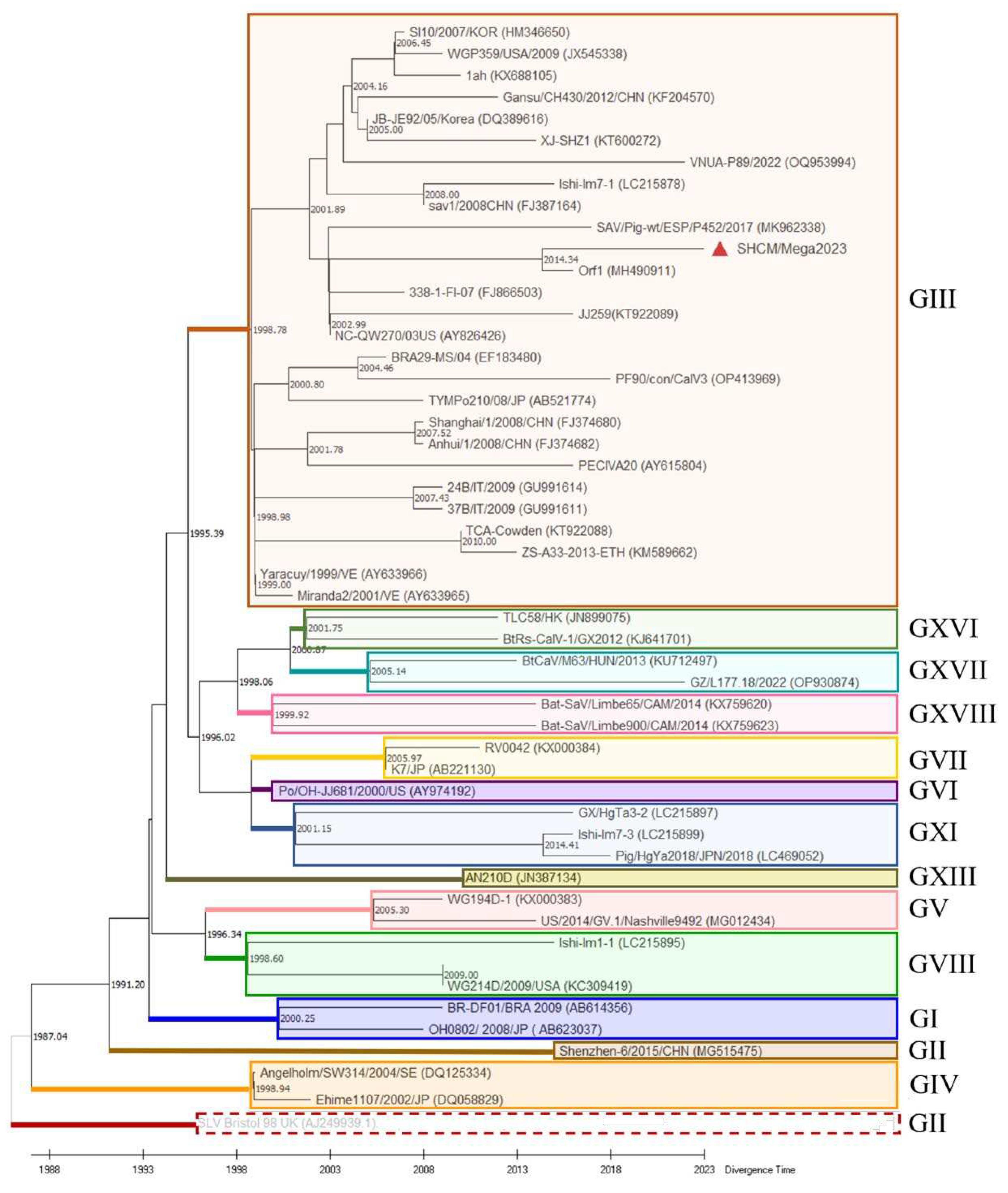
3.2. Phylogenetic and Time Evolutionary Analysis
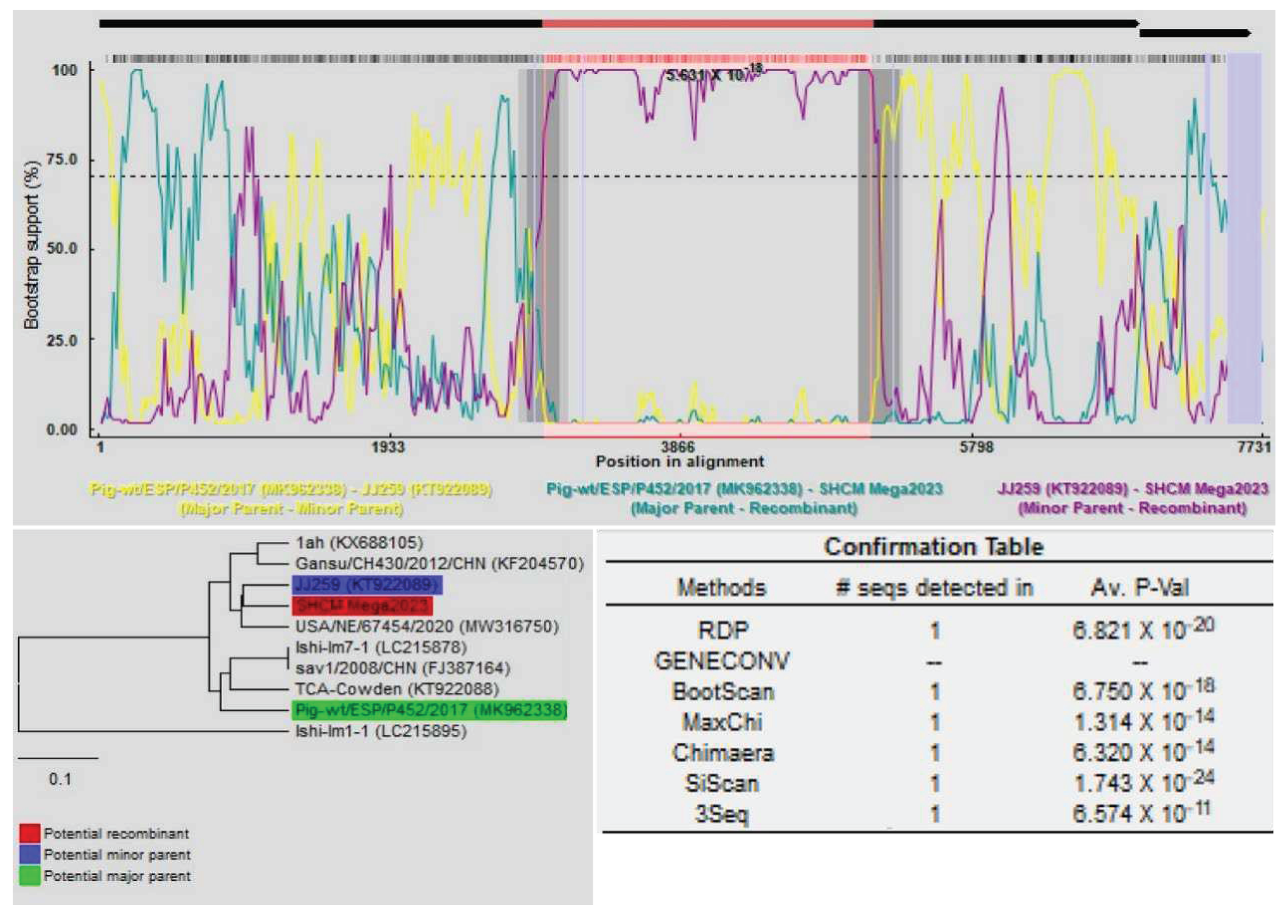
3.3. Recombinant Analysis
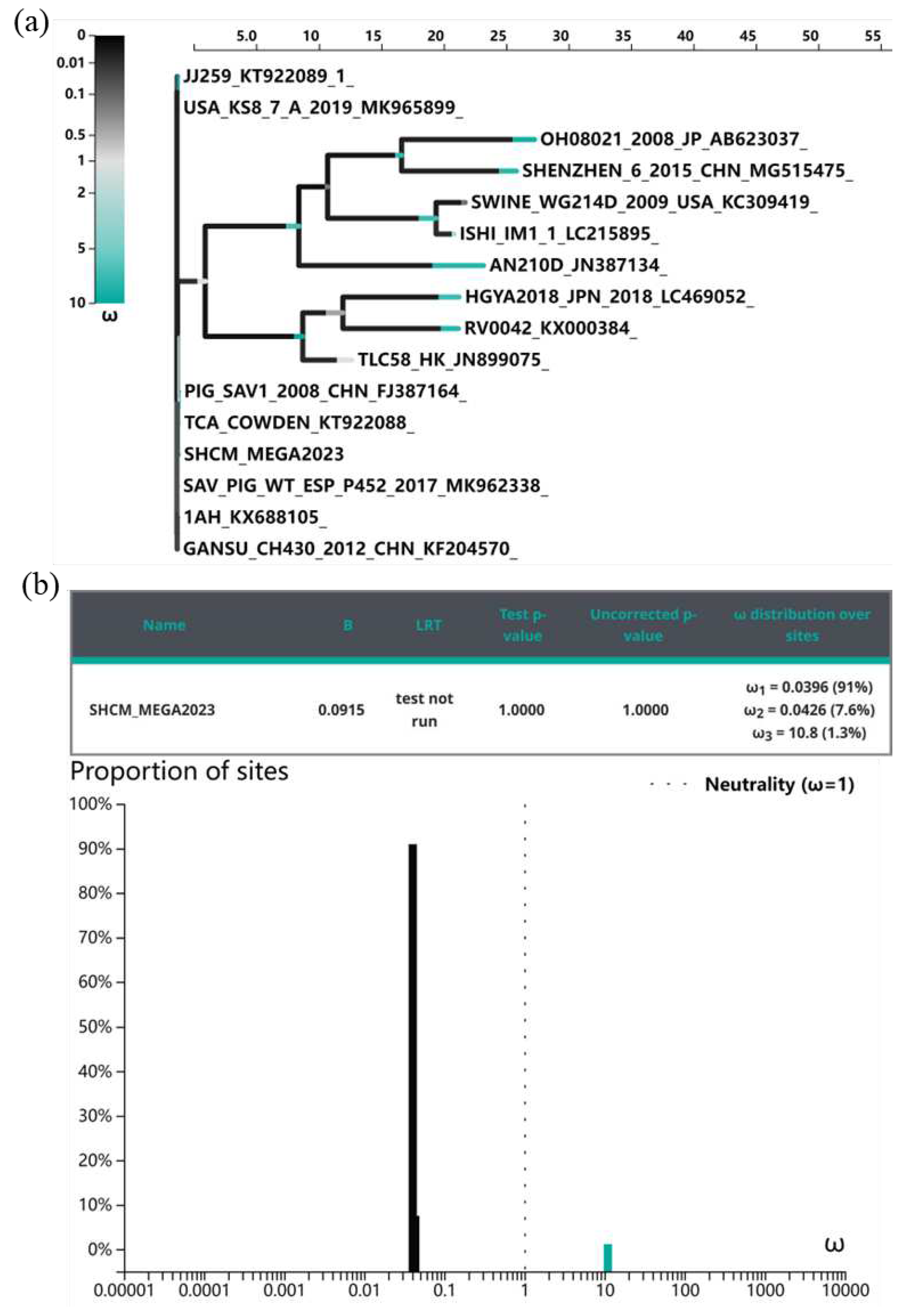
3.4. Evolutionary Selective Pressures on the ORF1 Codons
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Umair, M.; Rehman, Z.; Haider, S.A.; Usman, M.; Rana, M.S.; Ikram, A.; Salman, M. First report of coinfection and whole-genome sequencing of norovirus and sapovirus in an acute gastroenteritis patient from Pakistan. J. Med Virol. 2023, 95, e28458. [Google Scholar] [CrossRef]
- Choi, H.L.; Suh, C.-I.; Park, S.-W.; Jin, J.-Y.; Cho, H.-G.; Paik, S.-Y. Whole-Genome Sequencing Analysis of Sapovirus Detected in South Korea. PLOS ONE 2015, 10, e0132328. [Google Scholar] [CrossRef]
- Yinda, C.K.; Conceição-Neto, N.; Zeller, M.; Heylen, E.; Maes, P.; Ghogomu, S.M.; Van Ranst, M.; Matthijnssens, J. Novel highly divergent sapoviruses detected by metagenomics analysis in straw-colored fruit bats in Cameroon. Emerg. Microbes Infect. 2017, 6, e38. [Google Scholar] [CrossRef] [PubMed]
- Zhuo, R.; Ding, X.; Freedman, S.B.; Lee, B.E.; Ali, S.; Luong, J.; Xie, J.; Chui, L.; Wu, Y.; Pang, X.; Alberta Provincial Pediatric EnTeric Infection, T. Molecular Epidemiology of Human Sapovirus among Children with Acute Gastroenteritis in Western Canada. J. Clin. Microbiol. 2021, 59, JCM0098621. [Google Scholar] [CrossRef] [PubMed]
- Zhou, W.; Ullman, K.; Chowdry, V.; Reining, M.; Benyeda, Z.; Baule, C.; Juremalm, M.; Wallgren, P.; Schwarz, L.; Zhou, E.; et al. Molecular investigations on the prevalence and viral load of enteric viruses in pigs from five European countries. Veter- Microbiol. 2015, 182, 75–81. [Google Scholar] [CrossRef] [PubMed]
- Hoque, S.A.; Pham, N.T.K.; Onda-Shimizu, Y.; Nishimura, S.; Sugita, K.; Kobayashi, M.; Islam, M.T.; Okitsu, S.; Khamrin, P.; Maneekarn, N.; et al. Sapovirus infections in Japan before and after the emergence of the COVID-19 pandemic: An alarming update. J. Med Virol. 2023, 95, e29023. [Google Scholar] [CrossRef] [PubMed]
- Wei, R.; Shang, R.; Cheng, K.; Wang, S.; Wu, J.; Yu, Z. A novel recombinant porcine sapovirus infection in piglets with diarrhea in Shandong Province, China, 2022. Braz. J. Microbiol. 2023, 54, 1309–1314. [Google Scholar] [CrossRef] [PubMed]
- 8. Birch, J.M.; Leijon, M.; Nielsen, S.S.; Struve, T.; Jensen, H.E. Visualization of intestinal infections with astro- and sapovirus in mink (Neovison vison) kits by in situ hybridization. FEMS Microbes 2011, 2, xtab005. [Google Scholar] [CrossRef] [PubMed]
- Mijovski, J.Z.; Poljšak-Prijatelj, M.; Steyer, A.; Barlič-Maganja, D.; Koren, S. Detection and molecular characterisation of noroviruses and sapoviruses in asymptomatic swine and cattle in Slovenian farms. Infect. Genet. Evol. 2010, 10, 413–420. [Google Scholar] [CrossRef]
- Stamelou, E.; Giantsis, I.A.; Papageorgiou, K.V.; Petridou, E.; Davidson, I.; Polizopοulou, Z.S.; Papa, A.; Kritas, S.K. First report of canine Astrovirus and Sapovirus in Greece, hosting both asymptomatic and gastroenteritis symptomatic dogs. Virol. J. 2022, 19, 1–9. [Google Scholar] [CrossRef]
- Tse, H.; Chan, W.M.; Li, K.S.; Lau, S.K.; Woo, P.C.; Yuen, K.Y. Discovery and Genomic Characterization of a Novel Bat Sapovirus with Unusual Genomic Features and Phylogenetic Position. PLOS ONE 2012, 7, e34987. [Google Scholar] [CrossRef]
- dos Anjos, K.; Lima, L.M.; Silva, P.A.; Inoue-Nagata, A.K.; Nagata, T. The possible molecular evolution of sapoviruses by inter- and intra-genogroup recombination. Arch. Virol. 2011, 156, 1953–1959. [Google Scholar] [CrossRef]
- Saif, L.J.; Bohl, E.H.; Theil, K.W.; Cross, R.F.; House, J.A. Rotavirus-Like, Calicivirus-Like, and 23-nm Virus-Like Particles Associated with Diarrhea in Young Pigs. J. Clin. Microbiol. 1980, 12, 105–111. [Google Scholar] [CrossRef]
- Letunic, I.; Bork, P. Interactive tree of life (iTOL) v3: an online tool for the display and annotation of phylogenetic and other trees. Nucleic Acids Res. 2016, 44, W242–W245. [Google Scholar] [CrossRef]
- Tao, J.; Li, B.; Cheng, J.; Shi, Y.; Qiao, C.; Lin, Z.; Liu, H. Genomic Divergence Characterization and Quantitative Proteomics Exploration of Type 4 Porcine Astrovirus. Viruses 2022, 14, 1383. [Google Scholar] [CrossRef] [PubMed]
- Wisotsky, S.R.; Pond, S.L.K.; Shank, S.D.; Muse, S.V. Synonymous Site-to-Site Substitution Rate Variation Dramatically Inflates False Positive Rates of Selection Analyses: Ignore at Your Own Peril. Mol. Biol. Evol. 2020, 37, 2430–2439. [Google Scholar] [CrossRef] [PubMed]
- 17. Valente, C.S.; Alfieri, A.F.; Barry, A.F.; Leme, R.A.; Lorenzetti, E.; Alfieri, A.A. Age distribution of porcine sapovirus asymptomatic infection and molecular evidence of genogroups GIII and GIX? circulation in distinct Brazilian pig production systems. Trop Anim Health Prod 2016, 48, 21–27. [Google Scholar] [CrossRef] [PubMed]
- Rouhani, S.; Yori, P.P.; Olortegui, M.P.; A Lima, A.; Ahmed, T.; Mduma, E.R.; George, A.; Samie, A.; Svensen, E.; Lima, I.; et al. The Epidemiology of Sapovirus in the Etiology, Risk Factors, and Interactions of Enteric Infection and Malnutrition and the Consequences for Child Health and Development Study: Evidence of Protection Following Natural Infection. Clin. Infect. Dis. 2022, 75, 1334–1341. [Google Scholar] [CrossRef] [PubMed]
- Dufkova, L.; Scigalkova, I.; Moutelikova, R.; Malenovska, H.; Prodelalova, J. Genetic diversity of porcine sapoviruses, kobuviruses, and astroviruses in asymptomatic pigs: an emerging new sapovirus GIII genotype. Arch. Virol. 2012, 158, 549–558. [Google Scholar] [CrossRef] [PubMed]
- Nagai, M.; Wang, Q.; Oka, T.; Saif, L.J. Porcine sapoviruses: Pathogenesis, epidemiology, genetic diversity, and diagnosis. Virus Res. 2020, 286, 198025–198025. [Google Scholar] [CrossRef] [PubMed]
- Sunaga, F.; Masuda, T.; Aoki, H.; Ito, M.; Sano, K.; Naoi, Y.; Katayama, Y.; Omatsu, T.; Oba, M.; Furuya, T.; et al. Complete genome sequencing and genetic characterization of porcine sapovirus genogroup (G) X and GXI: GVI, GVII, GX, and GXI sapoviruses share common genomic features and form a unique porcine SaV clade. Infect. Genet. Evol. 2019, 75, 103959. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, K.; Saga, Y.; Iwai, M.; Obara, M.; Horimoto, E.; Hasegawa, S.; Kurata, T.; Okumura, H.; Nagoshi, M.; Takizawa, T. Frequent Detection of Noroviruses and Sapoviruses in Swine and High Genetic Diversity of Porcine Sapovirus in Japan during Fiscal Year 2008. J. Clin. Microbiol. 2010, 48, 1215–1222. [Google Scholar] [CrossRef] [PubMed]
- Mathijs, E.; Muylkens, B.; Mauroy, A.; Ziant, D.; Delwiche, T.; Thiry, E. Experimental evidence of recombination in murine noroviruses. J. Gen. Virol. 2010, 91, 2723–2733. [Google Scholar] [CrossRef] [PubMed]
- Supadej, K.; Khamrin, P.; Kumthip, K.; Malasao, R.; Chaimongkol, N.; Saito, M.; Oshitani, H.; Ushijima, H.; Maneekarn, N. Distribution of norovirus and sapovirus genotypes with emergence of NoV GII.P16/GII.2 recombinant strains in Chiang Mai, Thailand. J. Med Virol. 2018, 91, 215–224. [Google Scholar] [CrossRef]
- Dey, S.K.; Mizuguchi, M.; Okitsu, S.; Ushijima, H. Novel recombinant sapovirus in Bangladesh. Clin Lab 2011, 57, 91–94. [Google Scholar] [PubMed]
- Li, J.; Shen, Q.; Zhang, W.; Zhao, T.; Li, Y.; Jiang, J.; Yu, X.; Guo, Z.; Cui, L.; Hua, X. Genomic organization and recombination analysis of a porcine sapovirus identified from a piglet with diarrhea in China. Virol. J. 2017, 14, 1–8. [Google Scholar] [CrossRef]
- Oka, T.; Wang, Q.; Katayama, K.; Saif, L.J. Comprehensive Review of Human Sapoviruses. Clin. Microbiol. Rev. 2015, 28, 32–53. [Google Scholar] [CrossRef]
- Suzuki, Y. Natural Selection on the Influenza Virus Genome. Mol. Biol. Evol. 2006, 23, 1902–1911. [Google Scholar] [CrossRef]
- Correia, V.; Abecasis, A.B.; Rebelo-De-Andrade, H. Molecular footprints of selective pressure in the neuraminidase gene of currently circulating human influenza subtypes and lineages. Virology 2018, 522, 122–130. [Google Scholar] [CrossRef]
- Air, G.M.; Laver, W.G.; Luo, M.; Stray, S.J.; Legrone, G.; Webster, R.G. Antigenic, sequence, and crystal variation in influenza B neuraminidase. Virology 1990, 177, 578–587. [Google Scholar] [CrossRef]
- Abondio, P.; Cilli, E.; Luiselli, D. Inferring Signatures of Positive Selection in Whole-Genome Sequencing Data: An Overview of Haplotype-Based Methods. Genes 2022, 13, 926. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



