Submitted:
10 January 2024
Posted:
11 January 2024
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
2.1. Computational Results
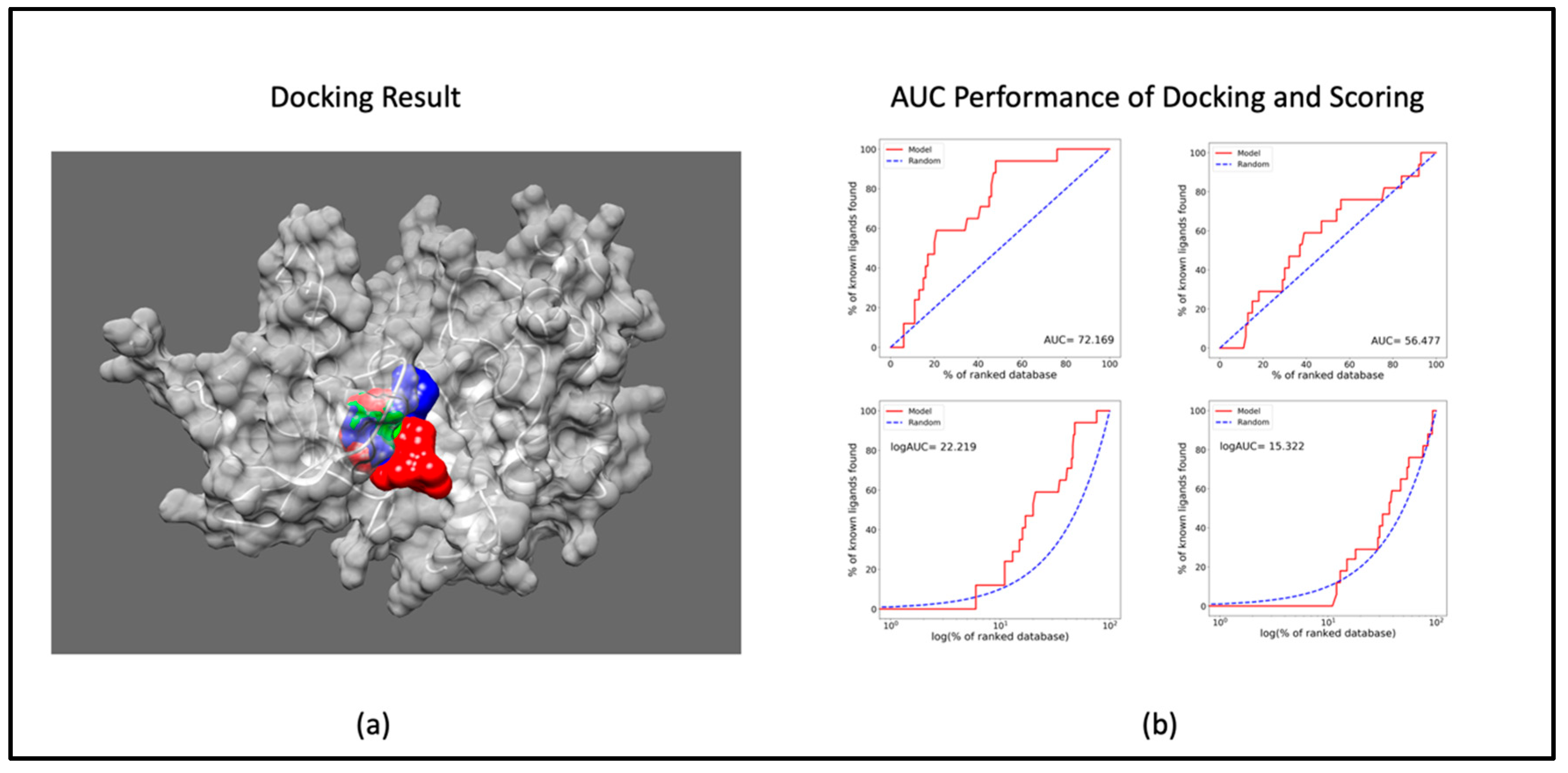
2.1.1. Docking Study
2.1.2. ADMET Analysis
2.1.3. MD Simulations
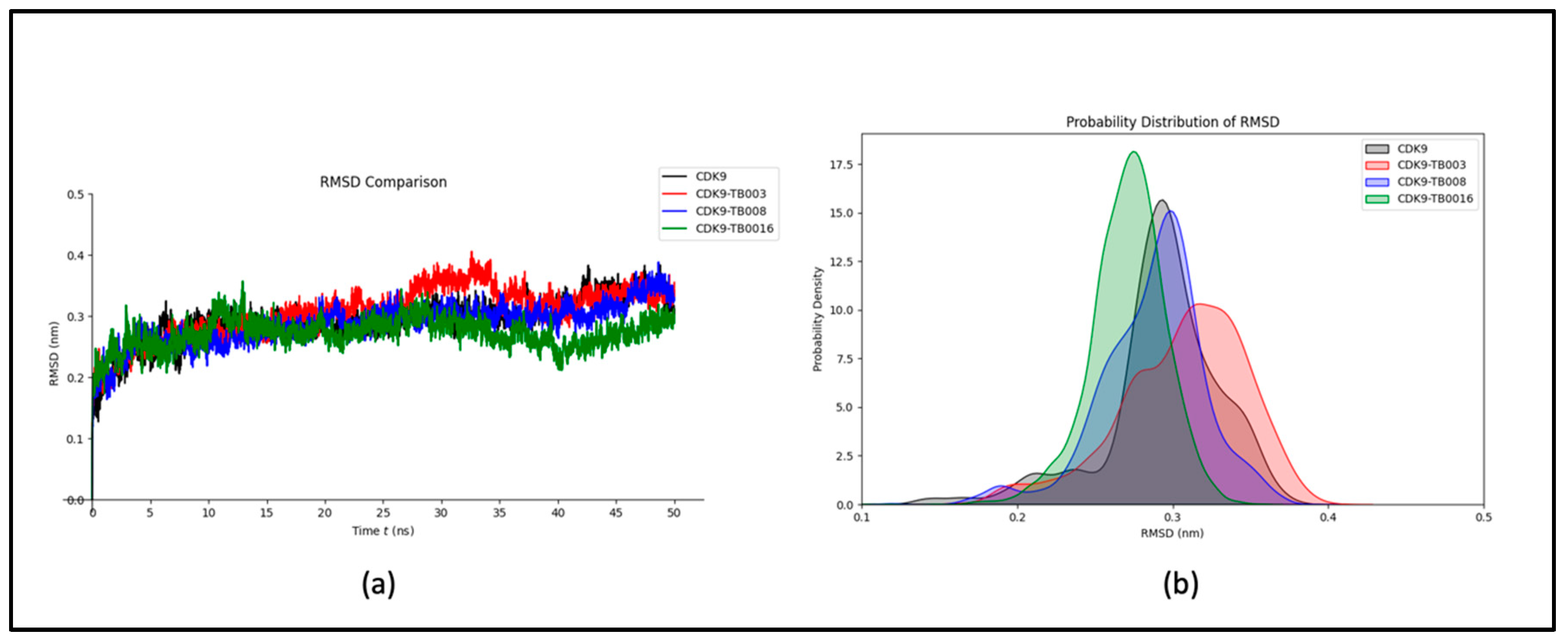
a) RMSD Analysis
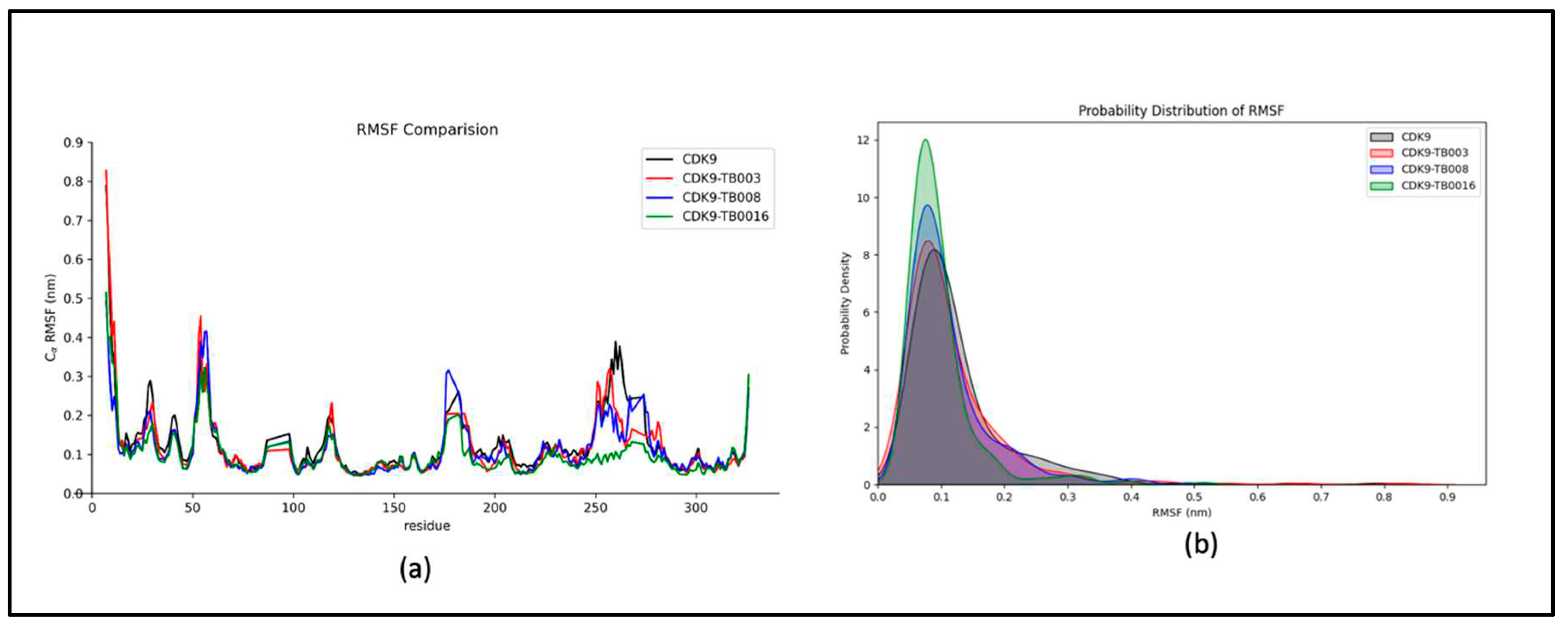
a) RMSF Analysis
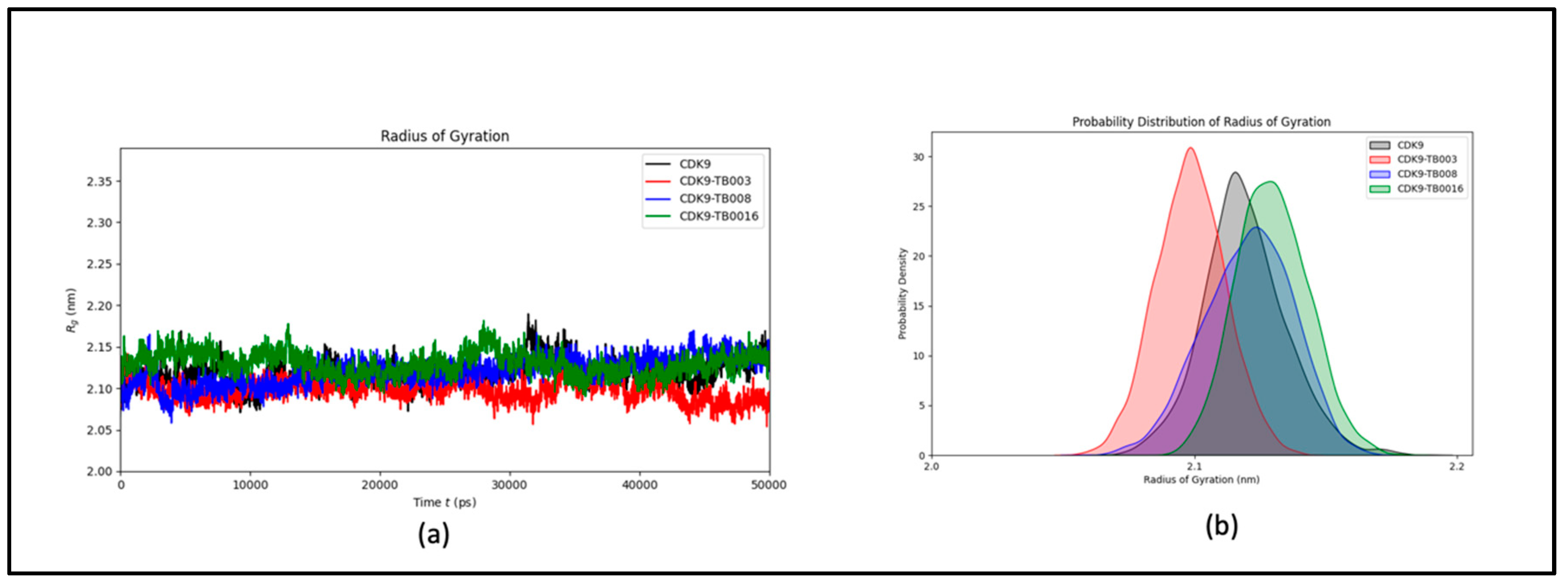
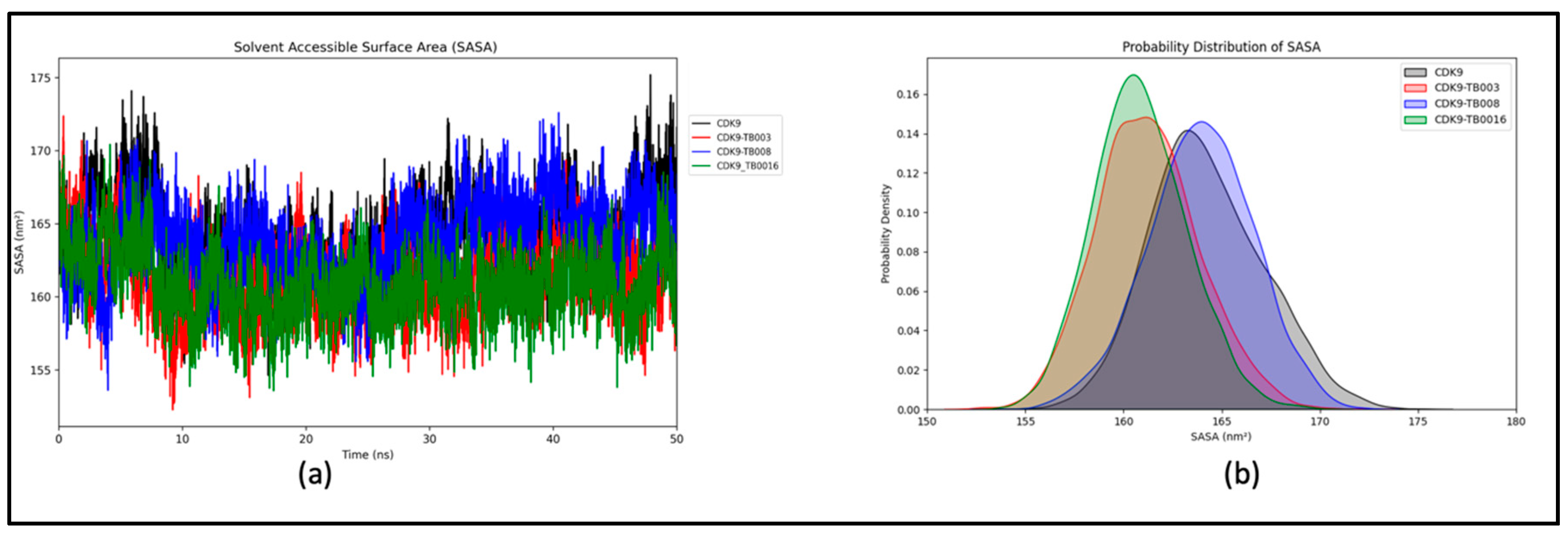
a) Radius of Gyration Analysis
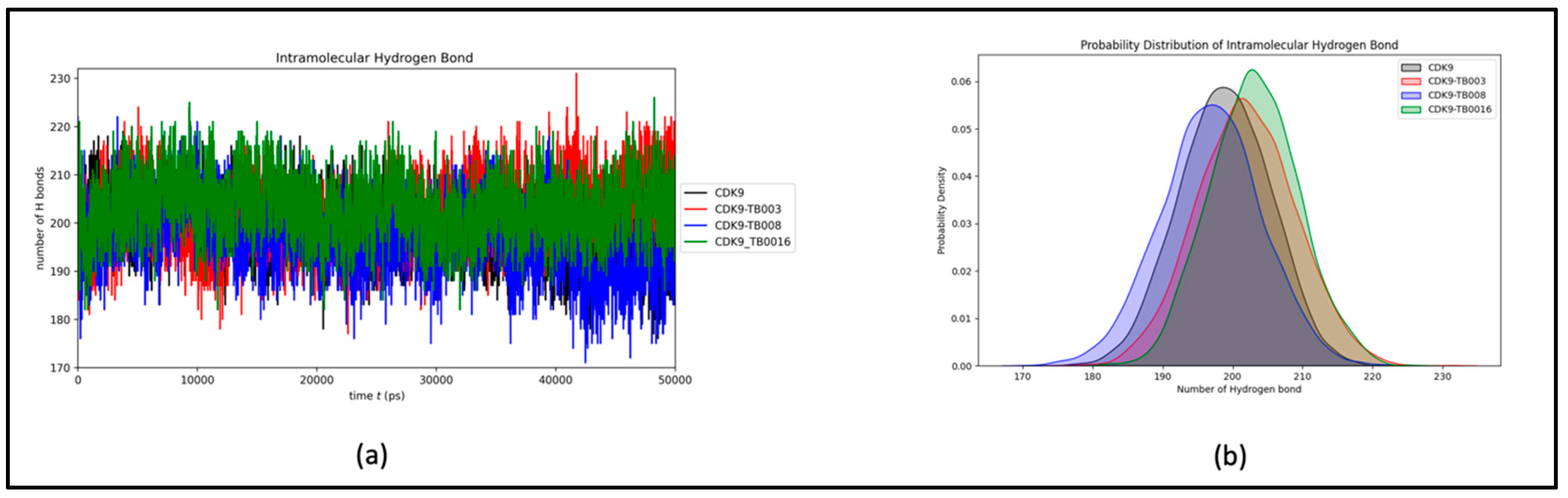
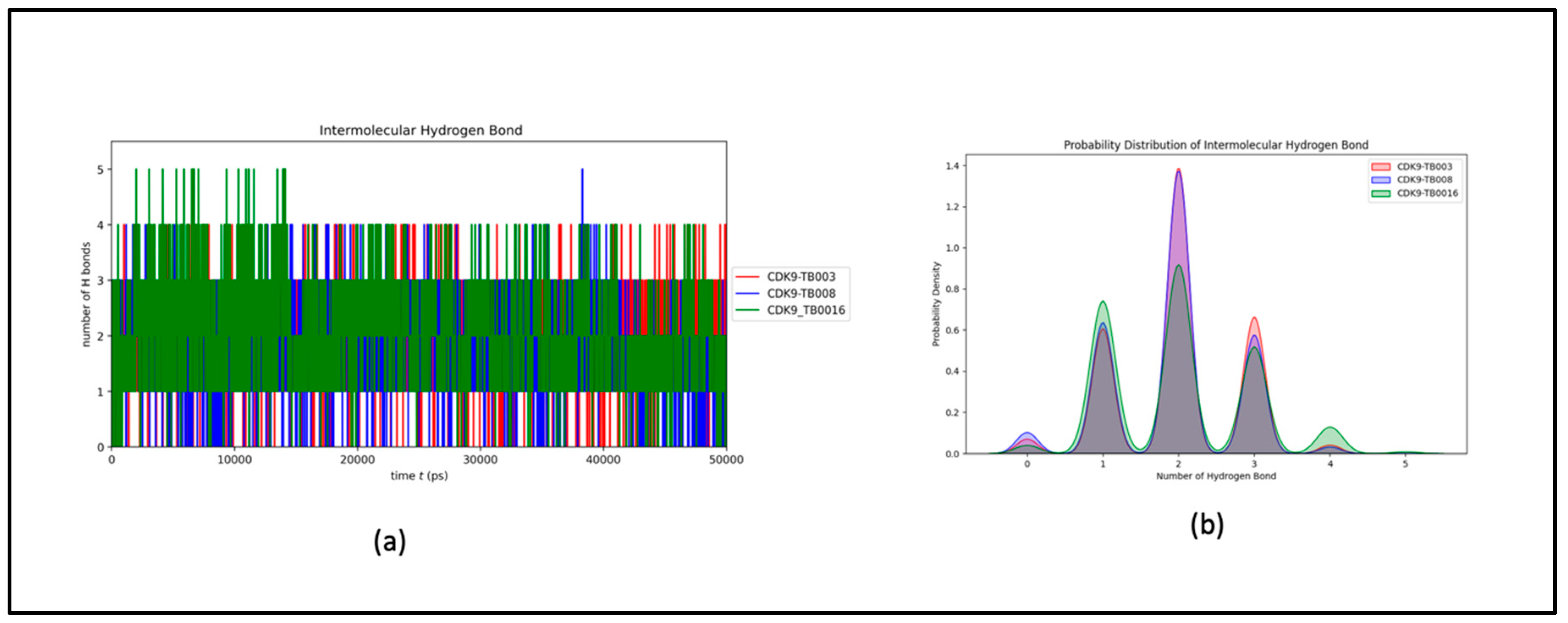
a) Hydrogen Bonds Dynamics
2.2. Experimental Results
2.2.1. Assay
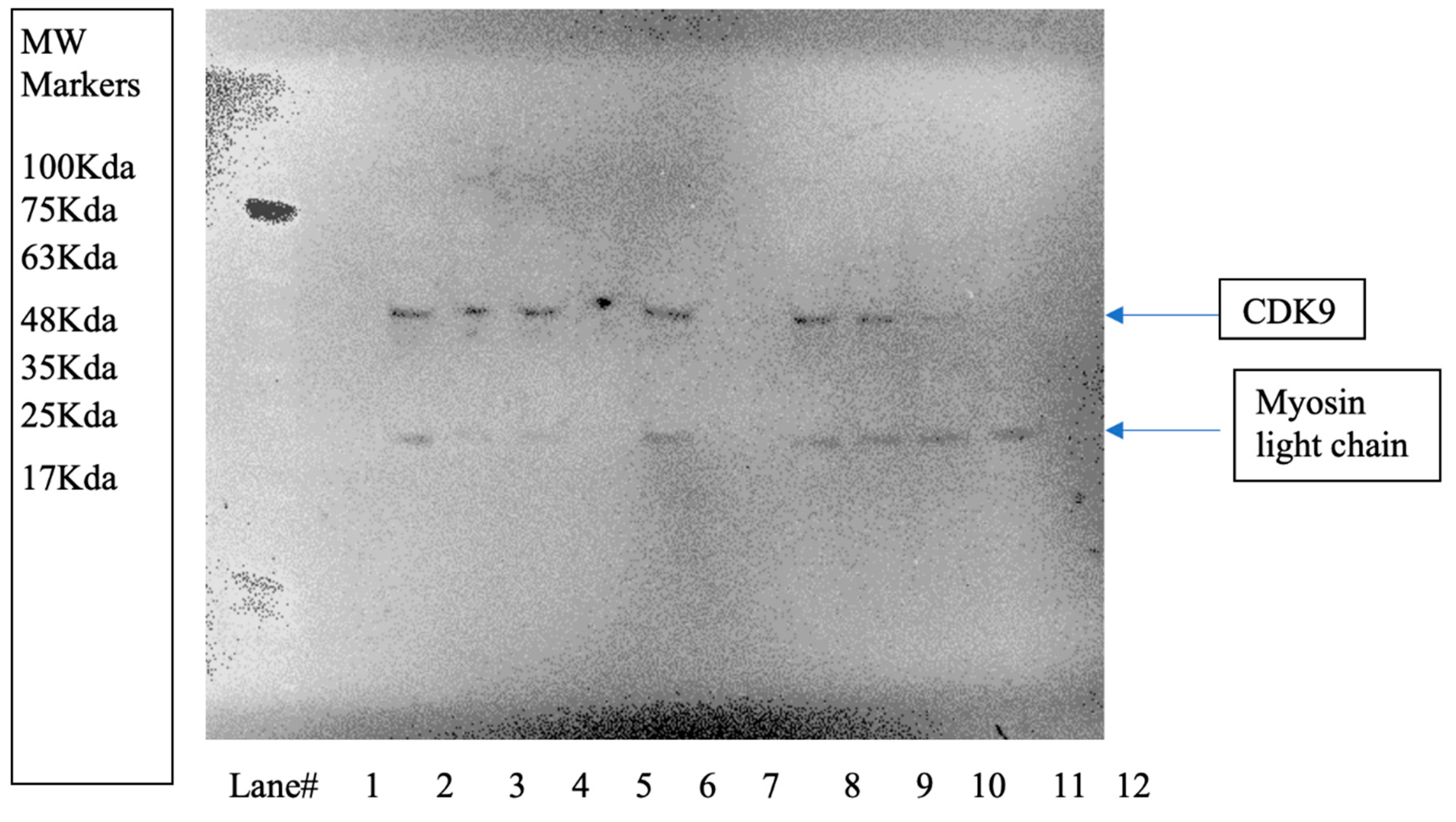
2.2.2. CDK9 Degradation by Western Blot
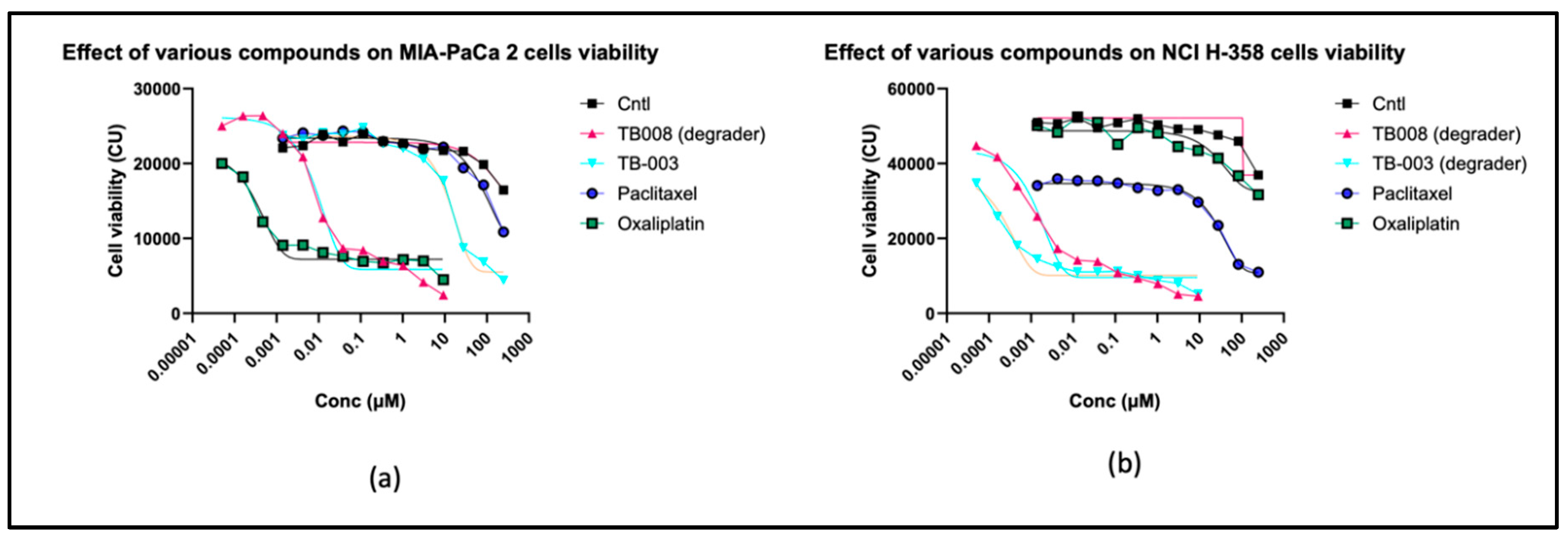
2.2.3. Cell viability
3. Discussions
4. Materials and Methods
4.1. Computational Methods
4.1.1. Compound Selection and Analysis
4.1.2. Target Preparation
4.1.3. Molecular Docking
4.1.3. ADMET Properties
4.1.4. Molecular Dynamics Calculations
4.2. Experimental Methods
4.2.1. Chemical Compounds
4.2.2. Biochemical Assays
a) Adapta Assay
a) Z-Lyte Assay
4.2.3. CDK9 Degradation with TB003
4.2.4. Effect of Degrader on Cell Viability
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Tai, W.; Mahato, R.; Cheng, K. The role of HER2 in cancer therapy and targeted drug delivery. J. Control. Release 2010, 146, 264–275. [Google Scholar] [CrossRef] [PubMed]
- Bondeson, D.P.; Mares, A.; Smith, I.E.D.; Ko, E.; Campos, S.; Miah, A.H.; Mulholland, K.E.; Routly, N.; Buckley, D.L.; Gustafson, J.L.; et al. Catalytic in vivo protein knockdown by small-molecule PROTACs. Nat. Chem. Biol. 2015, 11, 611–617. [Google Scholar] [CrossRef]
- Garriga, J.; Graña, X. CDK9 inhibition strategy defines distinct sets of target genes. BMC Res. Notes 2014, 7, 301–301. [Google Scholar] [CrossRef] [PubMed]
- Sajjadi-Dokht, M.; Mohamad, T.A.M.; Rahman, H.S.; Maashi, M.S.; Danshina, S.; Shomali, N.; Solali, S.; Marofi, F.; Zeinalzadeh, E.; Akbari, M.; et al. MicroRNAs and JAK/STAT3 signaling: A new promising therapeutic axis in blood cancers. Genes Dis. 2021, 9, 849–867. [Google Scholar] [CrossRef]
- Zhou, Q.; Li, T.; Price, D.H. RNA Polymerase II Elongation Control. Annu. Rev. Biochem. 2012, 81, 119–143. [Google Scholar] [CrossRef]
- Ott, M.; Geyer, M.; Zhou, Q. The Control of HIV Transcription: Keeping RNA Polymerase II on Track. Cell Host Microbe 2011, 10, 426–435. [Google Scholar] [CrossRef] [PubMed]
- Blachly, J.S.; Byrd, J.C.; Grever, M. Cyclin-dependent kinase inhibitors for the treatment of chronic lymphocytic leukemia. Semin. Oncol. 2016, 43, 265–273. [Google Scholar] [CrossRef]
- Blake, D.R.; Vaseva, A.V.; Hodge, R.G.; Kline, M.P.; Gilbert, T.S.K.; Tyagi, V.; Huang, D.; Whiten, G.C.; Larson, J.E.; Wang, X.; et al. Application of a MYC degradation screen identifies sensitivity to CDK9 inhibitors in KRAS-mutant pancreatic cancer. Sci. Signal. 2019, 12. [Google Scholar] [CrossRef]
- Lyu, J.; Yang, E.J.; Zhang, B.; Wu, C.; Pardeshi, L.; Shi, C.; Mou, P.K.; Liu, Y.; Tan, K.; Shim, J.S. Synthetic lethality of RB1 and aurora A is driven by stathmin-mediated disruption of microtubule dynamics. Nat. Commun. 2020, 11, 1–16. [Google Scholar] [CrossRef]
- Knight, Z.A.; Shokat, K.M. Chemical Genetics: Where Genetics and Pharmacology Meet. Cell 2007, 128, 425–430. [Google Scholar] [CrossRef]
- Lu, H.; et al. Compensatory induction of MYC expression by sustained CDK9 inhibition via a BRD4-dependent mechanism. Elife 2015, 4, e06535. [Google Scholar] [CrossRef] [PubMed]
- Morales, F.; Giordano, A. Overview of CDK9 as a target in cancer research. Cell Cycle 2016, 15, 519–527. [Google Scholar] [CrossRef] [PubMed]
- Rahaman, M.H.; Kumarasiri, M.; Mekonnen, L.B.; Yu, M.; Diab, S.; Albrecht, H.; Milne, R.W.; Wang, S. Targeting CDK9: A promising therapeutic opportunity in prostate cancer. Endocrine-Related Cancer 2016, 23, T211–T226. [Google Scholar] [CrossRef] [PubMed]
- Burslem, G.M.; Crews, C.M. Proteolysis-Targeting Chimeras as Therapeutics and Tools for Biological Discovery. Cell 2020, 181, 102–114. [Google Scholar] [CrossRef] [PubMed]
- Wu, T.; Qin, Z.; Tian, Y.; Wang, J.; Xu, C.; Li, Z.; Bian, J. Recent Developments in the Biology and Medicinal Chemistry of CDK9 Inhibitors: An Update. J. Med. Chem. 2020, 63, 13228–13257. [Google Scholar] [CrossRef] [PubMed]
- Olson, C.M.; Jiang, B.; Erb, M.A.; Liang, Y.; Doctor, Z.M.; Zhang, Z.; Zhang, T.; Kwiatkowski, N.; Boukhali, M.; Green, J.L.; et al. Pharmacological perturbation of CDK9 using selective CDK9 inhibition or degradation. Nat. Chem. Biol. 2018, 14, 163–170. [Google Scholar] [CrossRef]
- O’Boyle, N.M.; Banck, M.; James, C.A.; Morley, C.; Vandermeersch, T.; Hutchison, G.R. Open babel: An open chemical toolbox. J. Cheminform. 2011, 3, 33. [Google Scholar] [CrossRef] [PubMed]
- Mukherjee, S.; Yang, J.W.; Hoffmann, S.; List, B. Asymmetric Enamine Catalysis. Chem. Rev. 2007, 107, 5471–5569. [Google Scholar] [CrossRef] [PubMed]
- Schwede, T.; Kopp, J.; Guex, N.; Peitsch, M.C. SWISS-MODEL: An automated protein homology-modeling server. Nucleic Acids Res. 2003, 31, 3381–3385. [Google Scholar] [CrossRef]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Couch, G.S.; Greenblatt, D.M.; Meng, E.C.; Ferrin, T.E. UCSF Chimera? A visualization system for exploratory research and analysis. J. Comput. Chem. 2004, 25, 1605–1612. [Google Scholar] [CrossRef]
- DeLano, W.L. Pymol: An open-source molecular graphics tool. CCP4 Newsl. Protein Crystallogr 2002, 40, 82–92. [Google Scholar]
- Consortium, U. UniProt: A hub for protein information. Nucleic acids research 2015, 43, D204–D212. [Google Scholar] [CrossRef] [PubMed]
- Marchand, J.-R.; Pirard, B.; Ertl, P.; Sirockin, F. CAVIAR: A method for automatic cavity detection, description and decomposition into subcavities. J. Comput. Mol. Des. 2021, 35, 737–750. [Google Scholar] [CrossRef] [PubMed]
- Huey, R.; Morris, G.M.; Forli, S. Using AutoDock 4 and AutoDock vina with AutoDockTools: A tutorial. The Scripps Research Institute Molecular Graphics Laboratory 2012, 10550, 1000. [Google Scholar]
- Quiroga, R.; Villarreal, M.A. Vinardo: A Scoring Function Based on Autodock Vina Improves Scoring, Docking, and Virtual Screening. PLoS ONE 2016, 11, e0155183. [Google Scholar] [CrossRef] [PubMed]
- Masters, L.; Eagon, S.; Heying, M. Evaluation of consensus scoring methods for AutoDock Vina, smina and idock. J. Mol. Graph. Model. 2020, 96, 107532. [Google Scholar] [CrossRef] [PubMed]
- Mysinger, M.M.; Carchia, M.; Irwin, J.J.; Shoichet, B.K. Directory of Useful Decoys, Enhanced (DUD-E): Better Ligands and Decoys for Better Benchmarking. J. Med. Chem. 2012, 55, 6582–6594. [Google Scholar] [CrossRef] [PubMed]
- Daina, A.; Michielin, O.; Zoete, V. SwissADME: A free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci. Rep. 2017, 7, 42717. [Google Scholar] [CrossRef] [PubMed]
- Berendsen, H.J.C.; Van Der Spoel, D.; Van Drunen, R. GROMACS: A message-passing parallel molecular dynamics implementation. Comput. Phys. Commun. 1995, 91, 43–56. [Google Scholar] [CrossRef]
- Shukla, R.; Tripathi, T. Molecular dynamics simulation of protein and protein–ligand complexes. Computer-aided drug design 2020, 133–161. [Google Scholar]
- Huang, J.; Mackerell, A.D., Jr. CHARMM36 all-atom additive protein force field: Validation based on comparison to NMR data. J. Comput. Chem. 2013, 34, 2135–2145. [Google Scholar] [CrossRef] [PubMed]
- Vanommeslaeghe, K.; MacKerell, A.D. Automation of the CHARMM General Force Field (CGenFF) I: Bond Perception and Atom Typing. J. Chem. Inf. Model. 2012, 52, 3144–3154. [Google Scholar] [CrossRef] [PubMed]
- Bjørkøy, G.; et al. Monitoring autophagic degradation of p62/SQSTM1. Methods in enzymology 2009, 452, 181–197. [Google Scholar] [PubMed]
- Jang, J.; To, C.; De Clercq, D.J.H.; Park, E.; Ponthier, C.M.; Shin, B.H.; Mushajiang, M.; Nowak, R.P.; Fischer, E.S.; Eck, M.J.; et al. Mutant-Selective Allosteric EGFR Degraders are Effective Against a Broad Range of Drug-Resistant Mutations. Angew. Chem. 2020, 132, 14589–14597. [Google Scholar] [CrossRef]
| Ligand Name | Binding affinity (kcal/mol) | Docking Method |
|---|---|---|
| TB003 | -7.8 | Vina |
| TB003 | -8.4 | Smina |
| TB008 | -9.6 | Vina |
| TB008 | -9.8 | Smina |
| TB0016 | -9.7 | Vina |
| TB0016 | -8.8 | Smina |
| Compound | Lipophilicity (LogP) | Absorption | Solubility (LogS) |
|---|---|---|---|
| TB003 | 5.09 (High) | Low | -6.29 (Poorly Soluble) |
| TB008 | 2.75 (Moderate) | High | -3.83 (Soluble) |
| TB0016 | 2.55 (Moderate) | High | -4.28 (Moderately Soluble) |
| TB003 PROTAC | 4.28 (High) | Low | -6.65 (Poorly Soluble) |
| TB008 PROTAC | 3.19 (High) | Low | -5.59 (Moderately Soluble) |
| Compound | TB003 | TB0016 | TB008 | |
|---|---|---|---|---|
| Modality | CDK9 degrader | CDK9 ligand | CDK9 degrader | |
| Potency (IC50) | 5nM | > 1µM | 3.5nM | |
| Fold selectivity CDK9 vs other CDK family members | CDK7 | >200 | > 1µM | >500 |
| CDK5 | NT | > 1µM | >500 | |
| CDK4 | NT | > 1µM | >500 | |
| CDK2 | >200 | > 1µM | >500 | |
| CDK1 | >200 | > 1µM | >500 | |
| Route of administration | IP/Oral | IP/Oral | IP/Oral | |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




