Submitted:
26 November 2023
Posted:
27 November 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Material and Methods
- -
- Histone deacetylase 8 ( PDB Code 3mz6) : Binding site center (ångström) x( 0,1808), y( 42,7338) z( 2,1861)
- -
- Histone deacetylase 6 (PDB Code 3gv4):Binding site center(ångström) x( -7,3659), y( 4,9697) z( 0,9734)
- -
- Histone deacetylase-like amidohydrolase ( PDB Code1zz1) :Binding site center(ångström) x( -28,5197), y( 16,2263) z( -16,5053)
- -
- Histone deacetylase 4( PDB Code 2vqq) Binding site center (ångström) x( -17,5922), y( 7,3406 z( -11,4742)
- -
- Histone deacetylase 2( PDB Code 3max) Binding site center (ångström) x( 48,9732), y( 15,6496z( -43,4309)
3. Results and Discussion
4. Conclusions
Authors Contributions
Funding
Conflict of Interest
References
- Marks, P. A., Miller, T., & Richon, V. M. (2003). Histone deacetylases. Current opinion in pharmacology, 3(4), 344-351.
- Gray, S. G., & Ekström, T. J. (2001). The human histone deacetylase family. Experimental cell research, 262(2), 75-83. [CrossRef]
- Barneda-Zahonero, B., & Parra, M. (2012). Histone deacetylases and cancer. Molecular oncology, 6(6), 579-589.
- Cress, W. D., & Seto, E. (2000). Histone deacetylases, transcriptional control, and cancer. Journal of cellular physiology, 184(1), 1-16.
- Marks, P. A., Richon, V. M., Miller, T., & Kelly, W. K. (2004). Histone deacetylase inhibitors. Advances in cancer research, 91, 137-168.
- Glozak, M. A., & Seto, E. (2007). Histone deacetylases and cancer. Oncogene, 26(37), 5420-5432.
- Dokmanovic, M., Clarke, C., & Marks, P. A. (2007). Histone deacetylase inhibitors: overview and perspectives. Molecular cancer research, 5(10), 981-989. [CrossRef]
- Aggarwal, B. B., Kumar, A., & Bharti, A. C. (2003). Anticancer potential of curcumin: preclinical and clinical studies. Anticancer research, 23(1/A), 363-398.
- Vallianou, N. G., Evangelopoulos, A., Schizas, N., & Kazazis, C. (2015). Potential anticancer properties and mechanisms of action of curcumin. Anticancer research, 35(2), 645-651.
- Meng, X. Y., Zhang, H. X., Mezei, M., & Cui, M. (2011). Molecular docking: a powerful approach for structure-based drug discovery. Current computer-aided drug design, 7(2), 146-157. [CrossRef]
- Odhar, H. A., Rayshan, A. M., Ahjel, S. W., Hashim, A. A., & Albeer, A. A. M. A. (2019). Molecular docking enabled updated screening of the matrix protein VP40 from Ebola virus with millions of compounds in the MCULE database for potential inhibitors. Bioinformation, 15(9), 627.
- Wang, A. H., Kruhlak, M. J., Wu, J., Bertos, N. R., Vezmar, M., Posner, B. I., ... & Yang, X. J. (2000). Regulation of histone deacetylase 4 by binding of 14-3-3 proteins. Molecular and cellular biology, 20(18), 6904-6912. [CrossRef]
- Vega, R. B., Matsuda, K., Oh, J., Barbosa, A. C., Yang, X., Meadows, E., ... & Olson, E. N. (2004). Histone deacetylase 4 controls chondrocyte hypertrophy during skeletogenesis. Cell, 119(4), 555-566. [CrossRef]
- Grozinger, C. M., & Schreiber, S. L. (2000). Regulation of histone deacetylase 4 and 5 and transcriptional activity by 14-3-3-dependent cellular localization. Proceedings of the National Academy of Sciences, 97(14), 7835-7840. [CrossRef]
| Histone deacetylases | Binding Energies ( kcal/mol) |
|---|---|
| Histone deacetylase 8 | -6.4 |
| Histone deacetylase 6 | -6.5 |
| Histone deacetylase 2 | -8.4 |
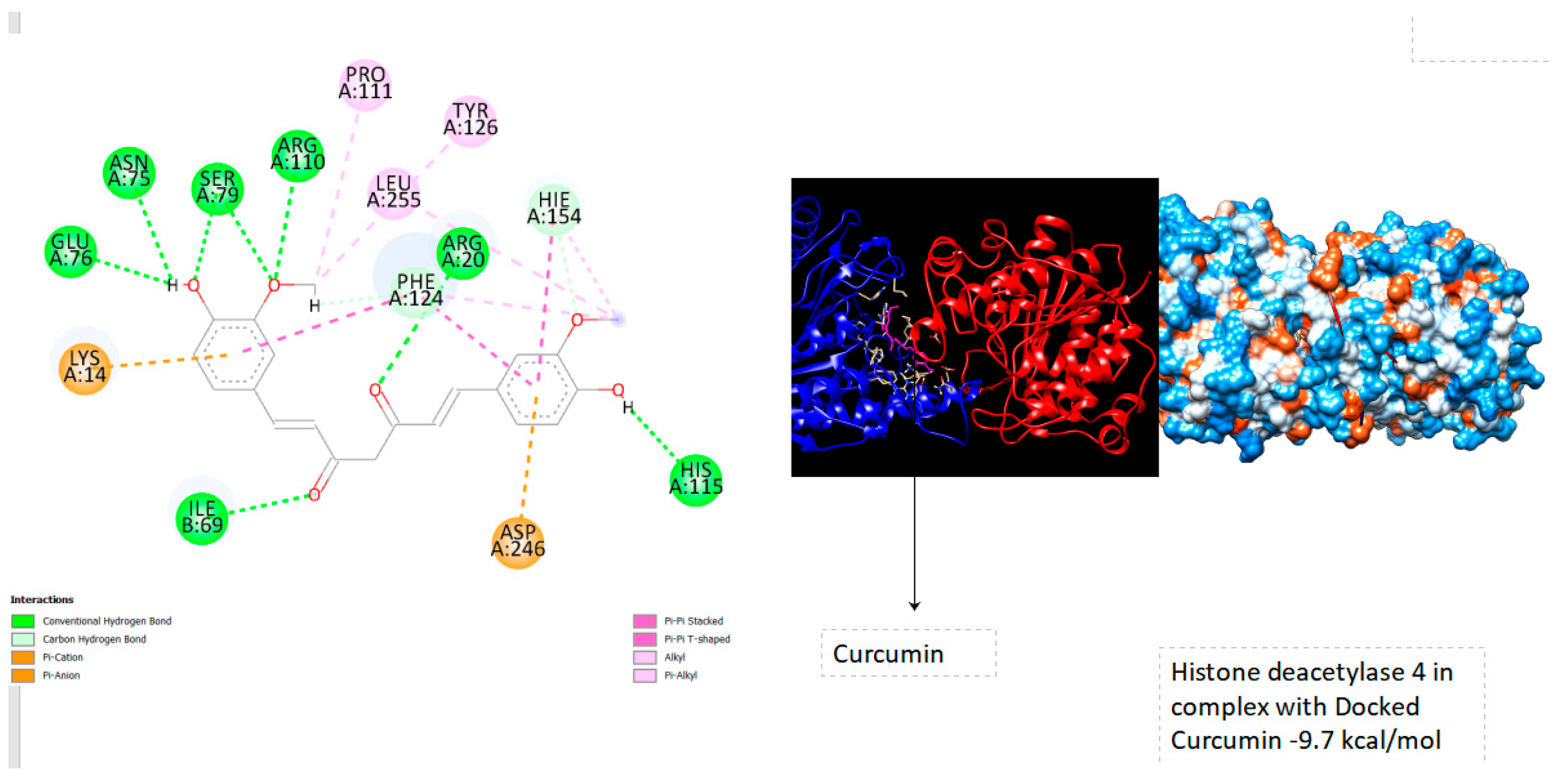
| Histone deacetylase 4 | -9.7 |
| Histone deacetylase-like amidohydrolas | -5.8 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




