2. Background
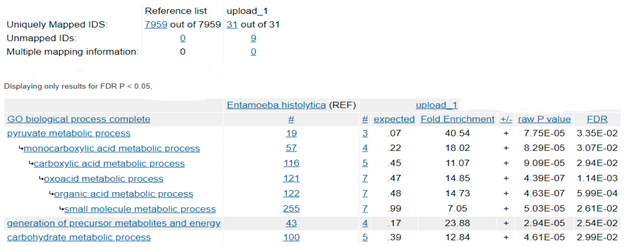
Based on the presence or absence of mitochondria, protozoa can be generally classified as either mitochondriate or amitochondriate organisms. The oxygen-sensitive enzyme PFOR, which is responsible for the anaerobic fermentative decarboxylation of pyruvate, has varying localizations in amitochondriate protozoa (Chabrière et al. 1999, Charon et al. 1999). Based on the localization of PFOR, amitochondriate protozoa are classified further into type 1 and type 2 protists (Hrdý et al. 1995, Muller 1998). In type 1 protists such as G. duodenalis, E. histolytica and C. parvum, PFOR is localized in the cytosol (Embley et al. 2003, Ctrnacta et al. 2006). The PFOR in type 2 protists is localized in the hydrogenosome, as observed in the trichomonads (Müller 1993, Muller 1998, Kulda 1999). The hydrogenosome is a double-membrane-bounded organelle (Benchimol et al. 1996, Benchimol 2009), which is postulated to have resulted either from an ancestral mitochondrial transformation with subsequent gene loss or from endosymbiosis of an anaerobic bacterium (Roger et al. 1996, Hrdy et al. 2004, Makiuchi et al. 2014, Lewis et al. 2020). The hydrogenosome is the powerhouse for the fermentative decarboxylation of pyruvate to produce hydrogen, carbon dioxide, and ATP molecules in the absence of molecular oxygen (Lindmark et al. 1973, Hackstein et al. 1999, Boxma et al. 2005). Unlike mitochondria, hydrogenosomes have no genetic material; they contain proteins that are encoded by nuclear genes, which are post-translationally translocated into the organelle (Johnson et al. 1993, Rada et al. 2011). Interestingly, the free-living, mitochondriate, photosynthetic, protist-like algae C. reinhardtii is able to switch from aerobic to anaerobic metabolism by upregulating the synthesis of PFOR, which localizes and functions in the chloroplastic stroma (Noth et al. 2013, van Lis et al. 2013). However, the N-terminus of the PFOR of the facultative anaerobic, photosynthetic protist E. gracilis localizes in the mitochondrion (Rotte et al. 2001, Ctrnacta et al. 2006). Some prokaryotes possess orthologs of these enzymes that function in a similar manner as observed in protozoa. Due to the bizarrely restricted occurrence, localization, and distribution of these enzymes in protozoa, potential bacterial origins have been proposed in previous studies (Hrdý et al. 1995, Horner et al. 1999, Rotte et al. 2001). One plausible mechanism to explain this phenomenon is by HGT. This study seeks to test the hypothesis that some ancestors of protozoa initially acquired these redox enzyme-encoding genes by multiple independent horizontal gene transfer events from bacteria.
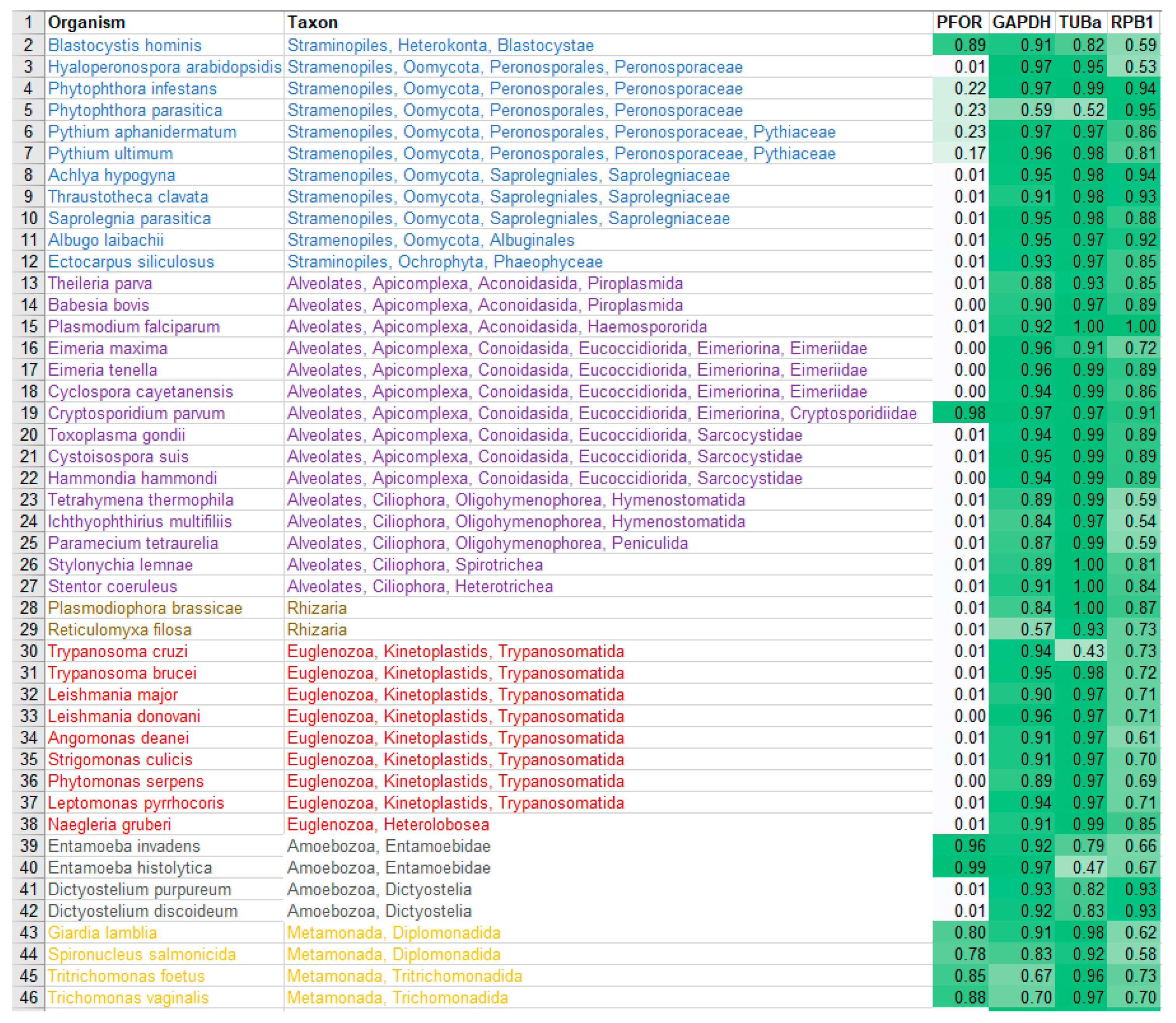
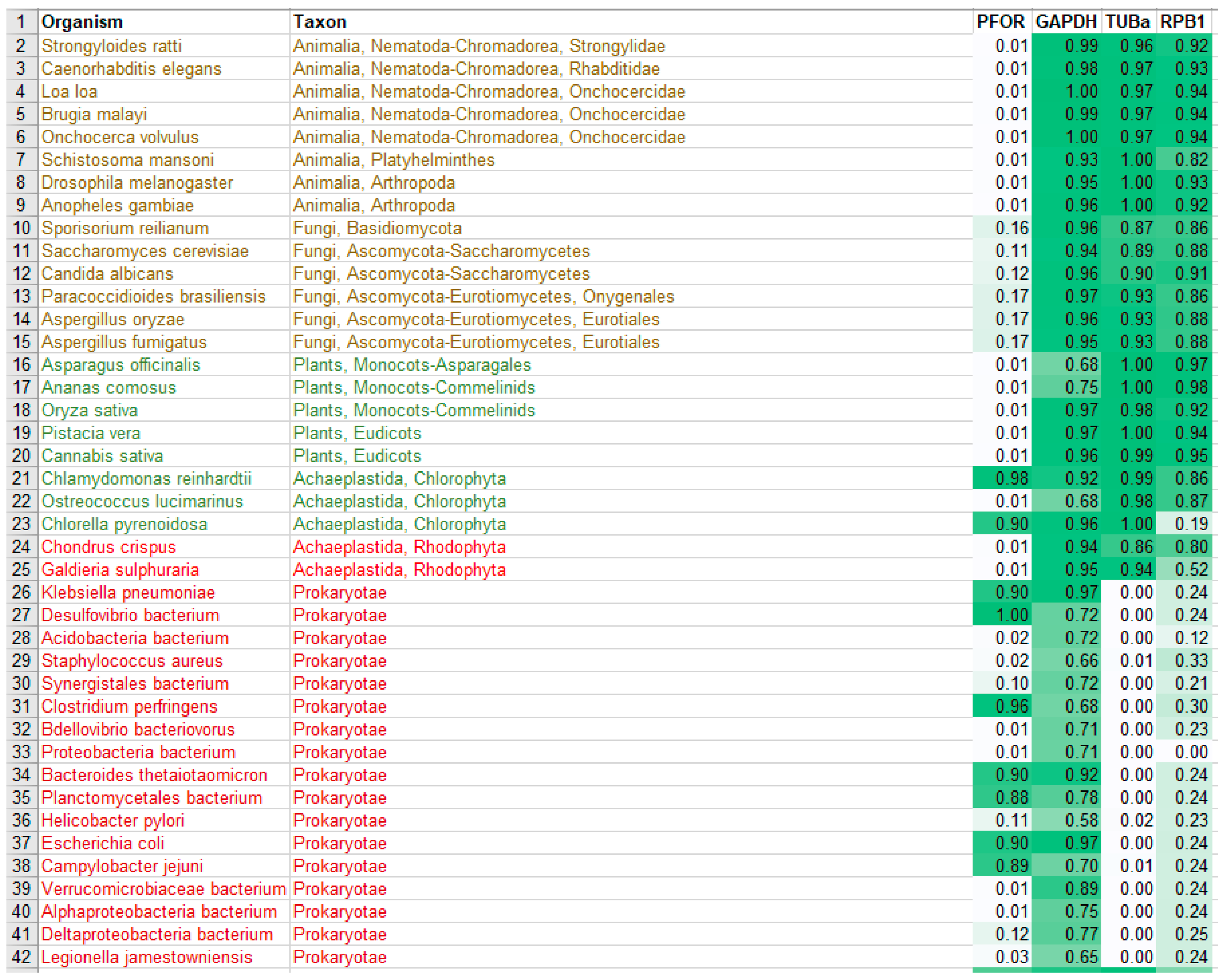
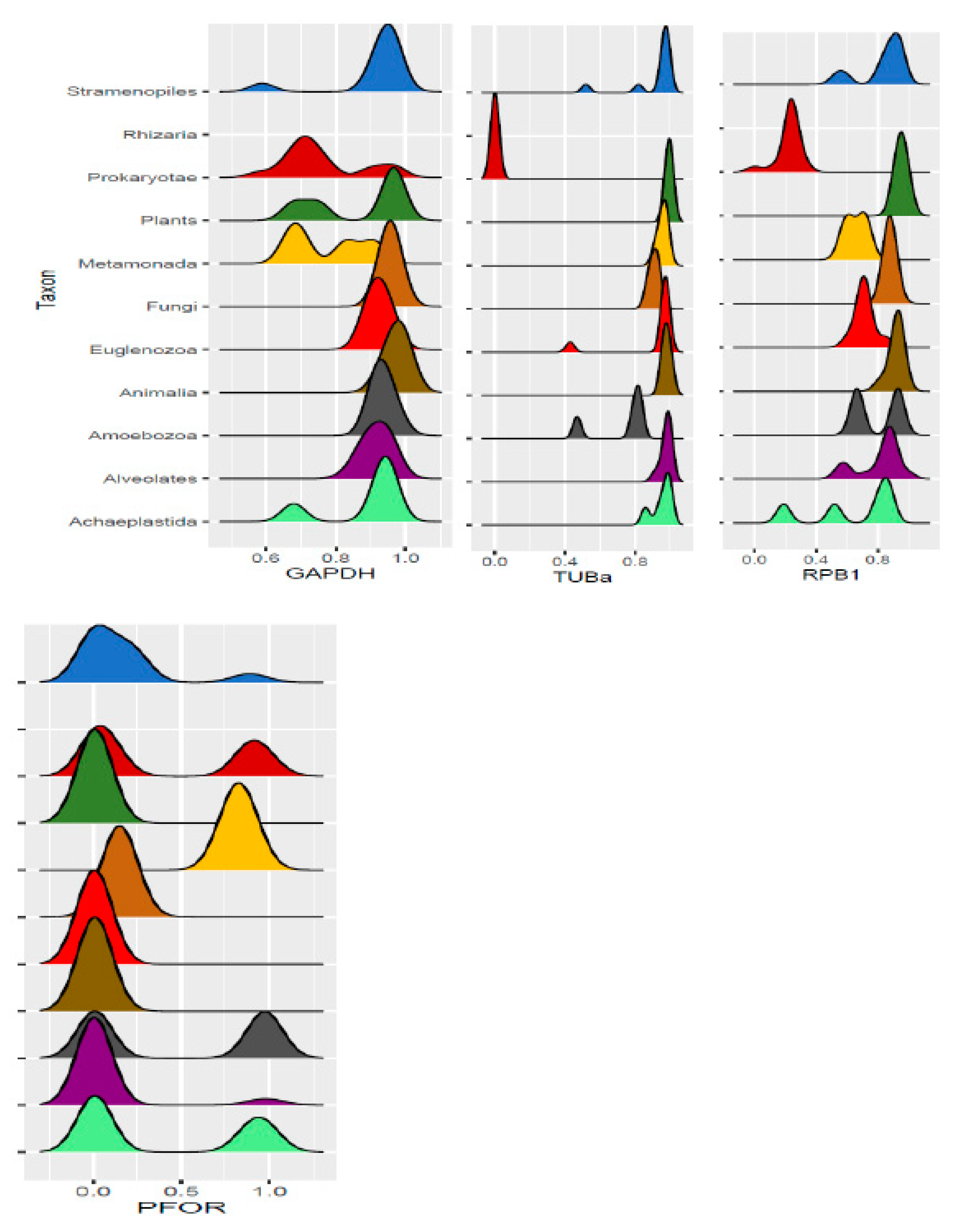
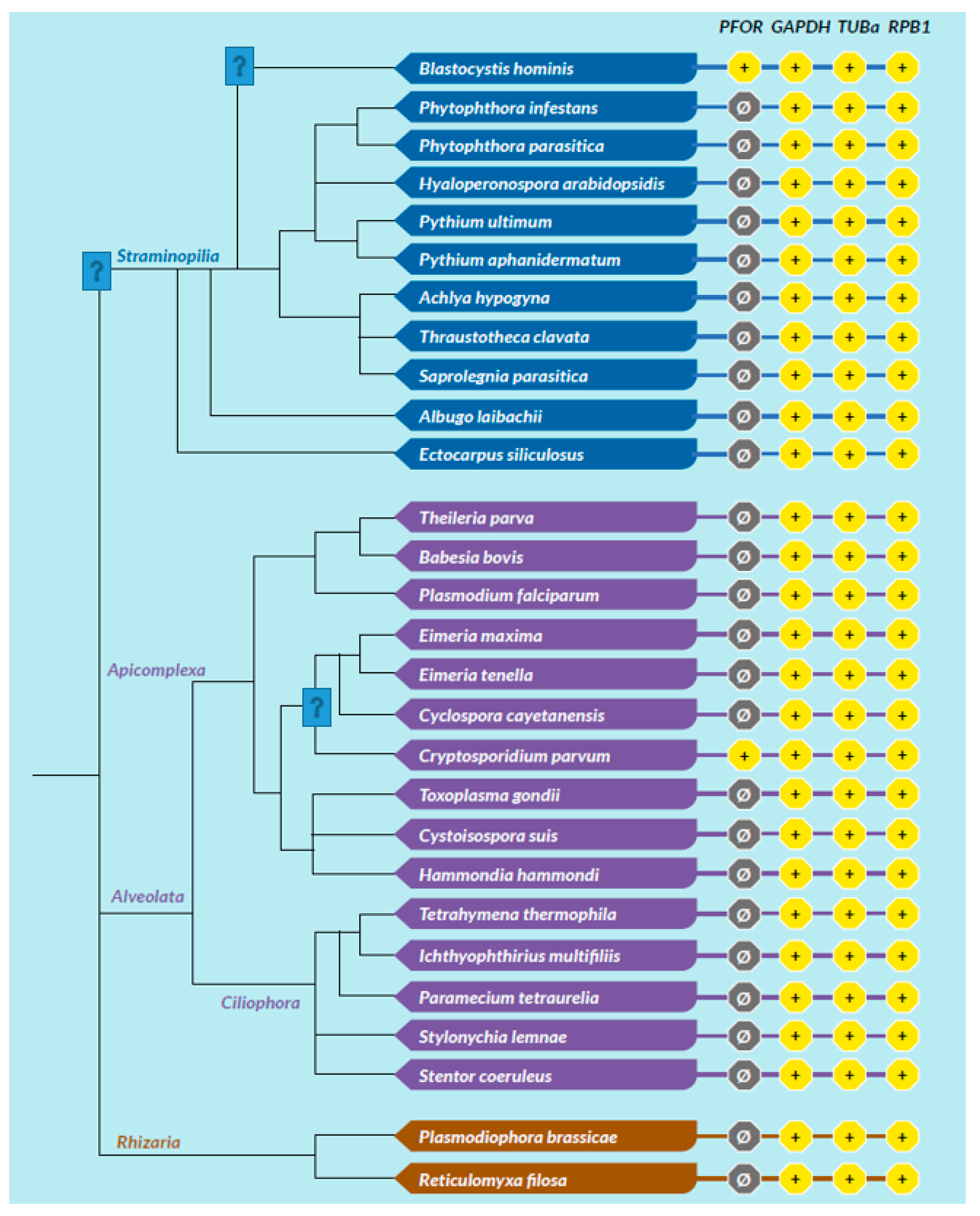
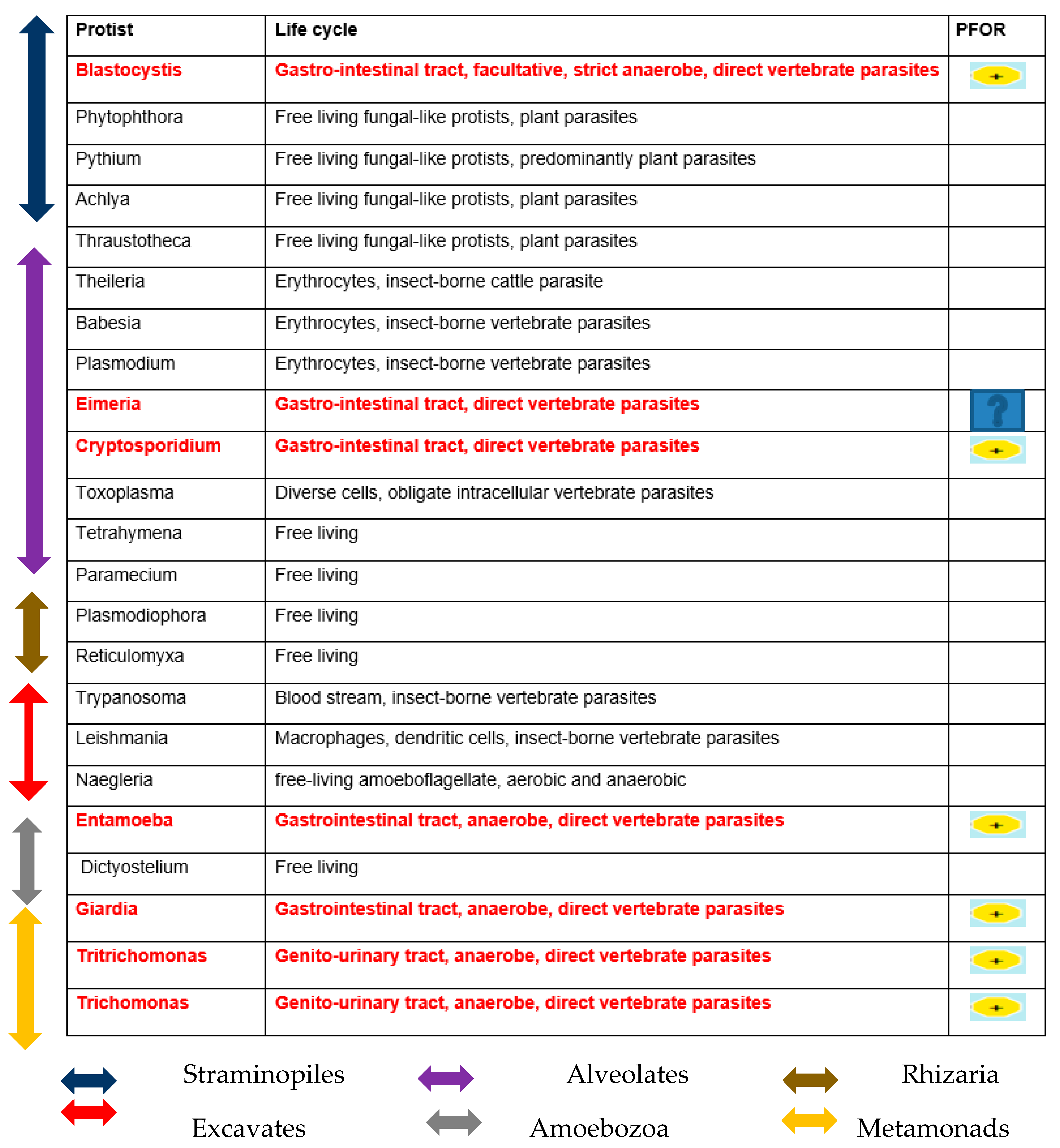
Table 1.
Diversity of the presence and cellular location of PFOR in protists.
Table 1.
Diversity of the presence and cellular location of PFOR in protists.
| Protist |
Localization of PFOR |
Source |
| Entamoeba histolytica |
Cytosol |
(Rodríguez et al. 1998) |
| Giardia duodenalis |
Cytosol |
(Emelyanov et al. 2011) |
| Cryptosporidium parvum |
Cytosol |
(Ctrnacta et al. 2006) |
| Blastocystis hominis |
Hydrogenosome |
(Wawrzyniak et al. 2008) |
| Trichomonas vaginalis |
Hydrogenosome |
(Hrdý et al. 1995) |
| Tritrichomonas foetus |
Hydrogenosome |
(Docampo et al. 1987) |
| Euglena gracilis |
Mitochondrion |
(Rotte et al. 2001) |
| Chlamydomonas reinhardtii |
Chloroplast |
(van Lis et al. 2013) |
Based on the cellular localization of PFOR E. histolytica, G. duodenalis and C. parvum are classified as type-1 protists whereas T. vaginalis, T. foetus and B. hominis are classified as type 2 protists.
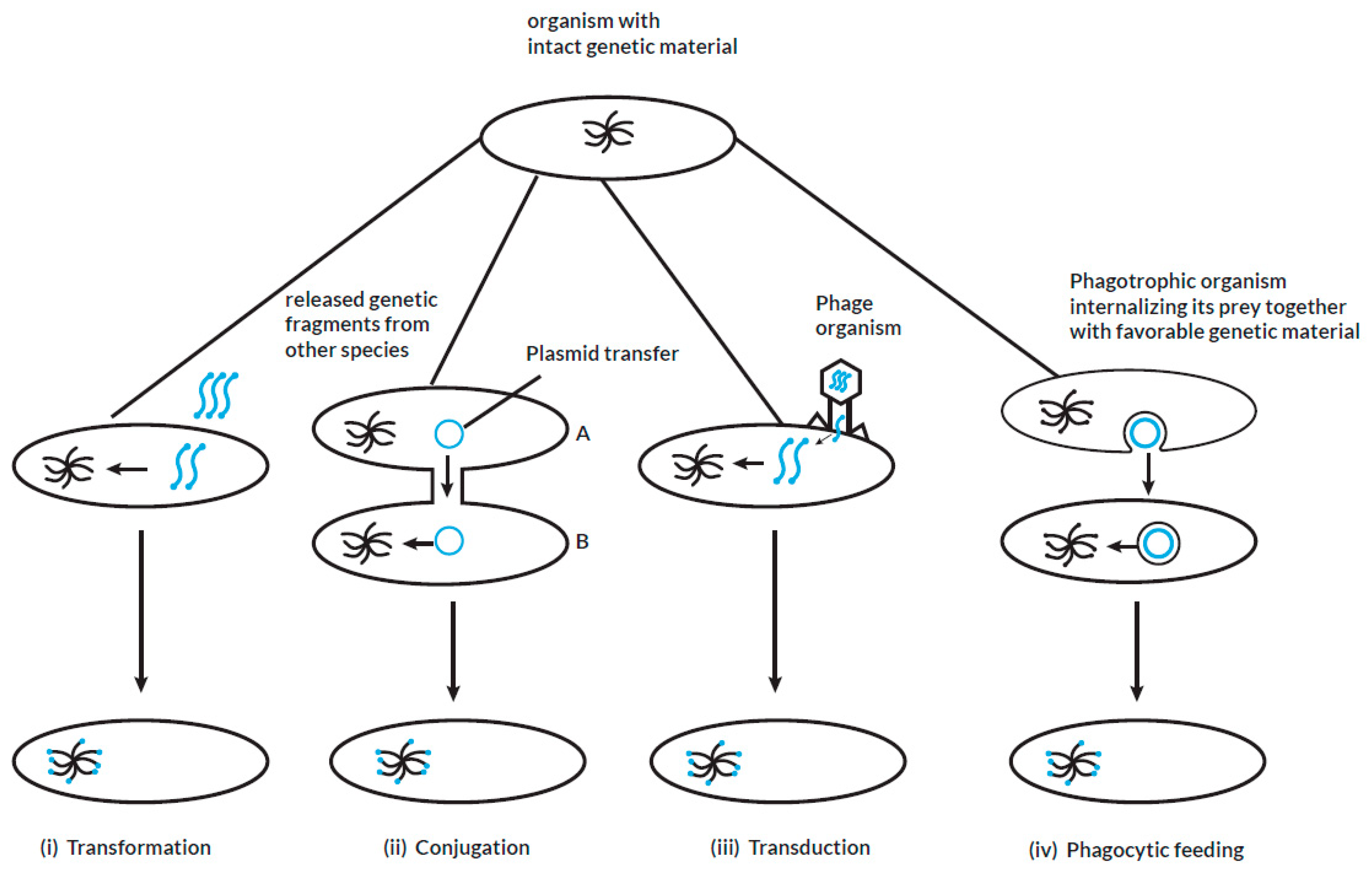
HGT, alternatively referred to as lateral gene transfer (LGT), is a term used in evolutionary genetics to explain the transfer of genetic material or genomic fragments between genetically related or unrelated organisms that do not share an inherent parent-to-offspring reproductive relationship (Soucy et al. 2015, Daubin et al. 2016). HGT is likely to occur through naturally occurring processes such as transformation or transfection, genetic conjugation and transduction: a detailed illustration of the different mechanisms of HGT is shown in
Figure 1. It may result in a substantial change in the genetic composition of the recipient organism or may be subsequently lost, leading to no overall significant genetic change. However, some HGT events may lead to the maintenance of pre-existing functions in the recipient organism that had been lost over the course of time in an evolutionary process termed as maintenance transfer (Husnik et al. 2018). The role of HGT in the acquisition of virulence and multiple drug resistance genes within prokaryotes is well established (Juhas 2015, Deng et al. 2019, Wyres et al. 2019). However, its involvement in the transfer of genetic material from prokaryotes to eukaryotes or within eukaryotes remains controversial (Keeling et al. 2008, Lacroix et al. 2016, Lacroix et al. 2018). Studies postulate that, HGT is an essential evolutionary mechanism by which unicellular eukaryotes acquire functional genetic material from bacteria that reside in close proximity such as the host gastrointestinal or genito-urinary tract (Ricard et al. 2006, Eme et al. 2017). It is also proposed that, some protozoan parasites residing in the same host environment with bacteria can receive foreign DNA fragments through phagocytic feeding on bacteria (Doolittle 1998). The acquisition of exogenous genetic material facilitates the adaptation, pathogenicity, and survival of protozoa in the face of strong selective pressures (Field et al. 2000, Hedges et al. 2001, Ricard et al. 2006, Keeling 2009, Schönknecht et al. 2013, Husnik et al. 2018, Sibbald, Eme et al. 2020, Van Etten et al. 2020). These protozoa may have acquired genes encoding bacterial redox enzymes via HGT to facilitate their survival in deteriorating environments (Nixon et al. 2002). Since protozoa have evolved to survive in oxidative and xenobiotic environments together with bacteria, these enzymes have remained central to their overall survival and were not lost after HGT (Gomaa et al. 2021). The proportion of bacteria-like genes proposed as potential candidates of HGT as apparent from whole genome sequencing of protozoa is summarized in table 2. A previous study that utilized conservative phylogenetic methods of the whole genome sequences of
E. histolytica and
T. vaginalis inferred 68 and 153 cases of HGT, respectively. Most of the transferred genes were found to encode enzymes involved in intermediary metabolism in prokaryotes, and the study concluded that prokaryotes that reside in close proximity to
E. histolytica and
T. vaginalis were potential gene donors (Alsmark, Sicheritz-Ponten et al. 2009).
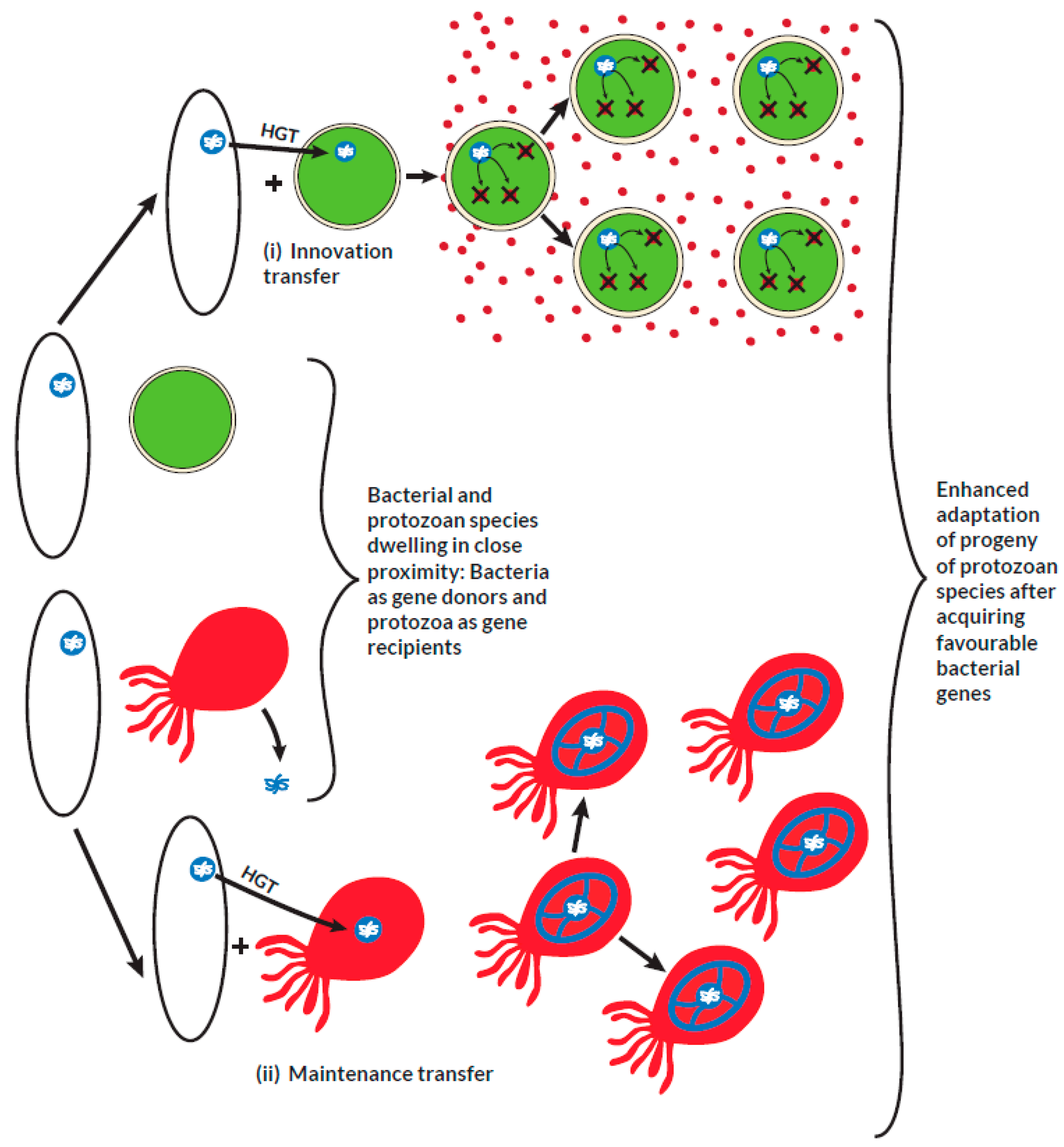
During the process of transformation (i), free genetic fragments from other organisms that reside in close proximity with the recipient organisms are horizontally transferred. The foreign genetic material is incorporated into the genome of the recipient organisms. In conjugation (ii), the genetic material in the form of plasmids is transferred from a specific donor organism after close contact is established with the recipient organisms. The facilitator of a transduction process (iii) is a phage organism that directly transfers its genetic material by infecting the recipient organism. In all cases, the horizontally acquired genetic material is passed on vertically to the respective progeny. If the acquired genetic material enhances adaptation and fitness in the corresponding species, it is maintained over several generations in the lineage. In phagocytic feeding (IV), a phagotrophic organism obtains foreign genetic material by engulfing other species that reside in close proximity. The engulfed species are trapped in food vacuoles which then undergo digestion with subsequent internalization of the favorable foreign genetic material.
Prokaryotes dwelling in close proximity with some protists are potential sources of genes that encode proteins involved in important metabolic and adaptive functions required for growth and survival. In an innovation transfer, a protist that originally lacks such genes acquires them from a bacterial donor. From the example illustrated above in (i), a metabolically less endowed protist incorporates a bacterial gene responsible for expressing proteins involved in the detoxification of toxins. This serves as an adaptive advantage for the survival of the protist in an otherwise toxin-enriched environment that usually favors bacterial species. During the course of evolution, some vital genes that are lost may result in an adaptive disadvantage to the protist. In a maintenance transfer, as illustrated in (ii), a lost gene may be regained subsequently from bacterial sources to maintain the adaptation of the protist.
By targeting the pathogen-specific redox function of PFOR, some nitro-based antimicrobial prodrugs are bio-activated through electron transfer reactions intracellularly in invading pathogens. Substances containing one or more nitro groups, such as the nitro-heterocyclic and nitro-aromatic compounds, are used as prodrugs against these pathogens because they undergo bio-activation to form potent nitro radicals (Fleck et al. 2014). The anionic potential of the radical intermediates results in intracellular trapping and subsequent accumulation of more active metabolites. As a result, these metabolic radicals exert their cytotoxic effects selectively in invading pathogens with no significant effect on the host cells (Chung et al. 2011, Nepali et al. 2019).
Figure 3.
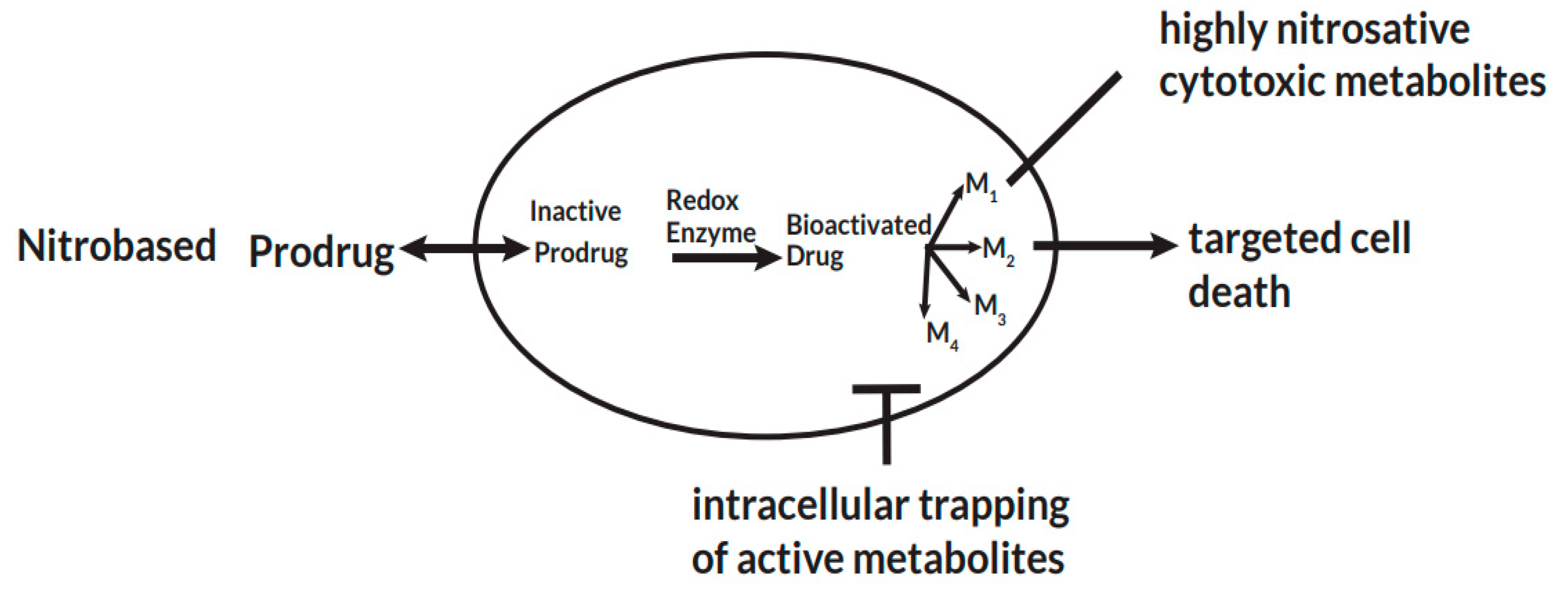
Pathogen specific bio-activation of nitro-based prodrugs.
Figure 3.
Pathogen specific bio-activation of nitro-based prodrugs.
After a prodrug is taken up intracellular by a pathogen, pathogen-specific redox enzymes catalyze electron transfer reactions that lead to the bio-activation of the prodrug. The activated drug undergoes a series of transformation reactions to radical cytotoxic metabolites that remain trapped intracellular and exert their cytotoxic effects. This leads to selective cell death by disrupting nucleic acid and protein synthesis of the pathogen.
Table 2.
Proportion of bacterial-like genes estimated from whole genome sequencing of some protozoan parasites.
Table 2.
Proportion of bacterial-like genes estimated from whole genome sequencing of some protozoan parasites.
| Protozoan parasite |
Total number of genes |
Percentage of bacterial-like genes |
Source |
| Plasmodium falciparum |
5,268 (212) |
4.02% |
(Gardner et al. 2002, Huang et al. 2004) |
| Blastocystis hominis |
6,020 (133) |
2.21% |
(Denoeud et al. 2011) |
| Entamoeba histolytica |
9,938 (96) |
0.97% |
(Loftus et al. 2005) |
| Toxoplasma gondii |
8,300 (72) |
0.87% |
(Huang et al. 2004, Bontell et al. 2009) |
| Trypanosoma brucei |
9,068 (47) |
0.52% |
(Berriman et al. 2005) |
| Cryptosporidium parvum |
5,519 (24) |
0.43% |
(Huang et al. 2004) |
| Dictyostelium discoideum |
13,522 (49) |
0.36% |
(Andersson 2011) |
| Naegleria gruberi |
15,727 (45) |
0.29% |
(Fritz-Laylin et al. 2010) |
| Tetrahymena thermophilia |
26,460 (74) |
0.28% |
(Xiong et al. 2015) |
| Trichomonas vaginalis |
59,681 (152) |
0.25% |
(Carlton et al. 2007) |
This table shows the varying proportion of bacterial-like genes in the whole genome sequencing of some protozoan species proposed as potential candidates for HGT from bacteria. Of all the protozoa of economic importance to humans that have been fully sequenced, P. falciparum seems to be the parasite with the highest proportion of bacterial-genes. The numbers in parenthesis are the absolute number of bacterial-like genes.
2.1. Pyruvate-ferredoxin oxidoreductase
The ability to metabolize pyruvate through the process of oxidative decarboxylation remains a fundamental biochemical reaction in all living organisms (Inui et al. 1999). The pyruvate dehydrogenase multi-enzyme complex (PDC), an enzyme present in the mitochondria of almost all aerobic organisms, catalyzes this reaction (Hucho 1975). However, in most anaerobic and microaerophilic organisms, the enzyme PFOR, which is absent in higher eukaryotes, is responsible for catalyzing this reaction (Chabrière et al. 1999, Charon et al. 1999). PFOR is an ancient, oxygen sensitive, iron-sulfur protein that belongs to the 2-oxo acid ferredoxin oxidoreductase (OFOR) superfamily. It is found in diverse cellular locations in some anaerobic organisms (Emelyanov et al. 2011). Archaea, most anaerobic bacteria, and some anaerobic eukaryotes possess this enzyme (Blamey et al. 1993). The fermentative decarboxylation of pyruvate to acetyl-CoA and carbon dioxide (CO2) by PFOR generates electrons that are transferred to either ferredoxin or flavodoxin (Marczak et al. 1983, Moreno et al. 1985). Due to the reversibility of this reaction, whereby pyruvate will be regenerated, PFOR is also called pyruvate synthase.
According to previous studies, the innate function of PFOR is to protect anaerobic organisms against oxidative stress by catalyzing the transfer of electrons that eventually reduce and inactivate free oxygen radicals during unfavorable oxidative conditions (Raj, et al. 2014). In an alternative pathway, the reduced ferredoxin derived from the decarboxylation of pyruvate can be re-oxidized by ferredoxin: NAD oxidoreductase (FNOR). This involves the transfer of electrons from ferredoxin to NAD(P) (Ctrnacta et al. 2006). The resulting NAD(P)H then transfers the electrons to oxygen radicals by another reaction catalyzed by NAD(P)H oxidase to form water molecules (Mastronicola et al. 2016). In this way, PFOR contributes its redox function together with other enzymes as an antioxidant detoxification system.
Figure 4.
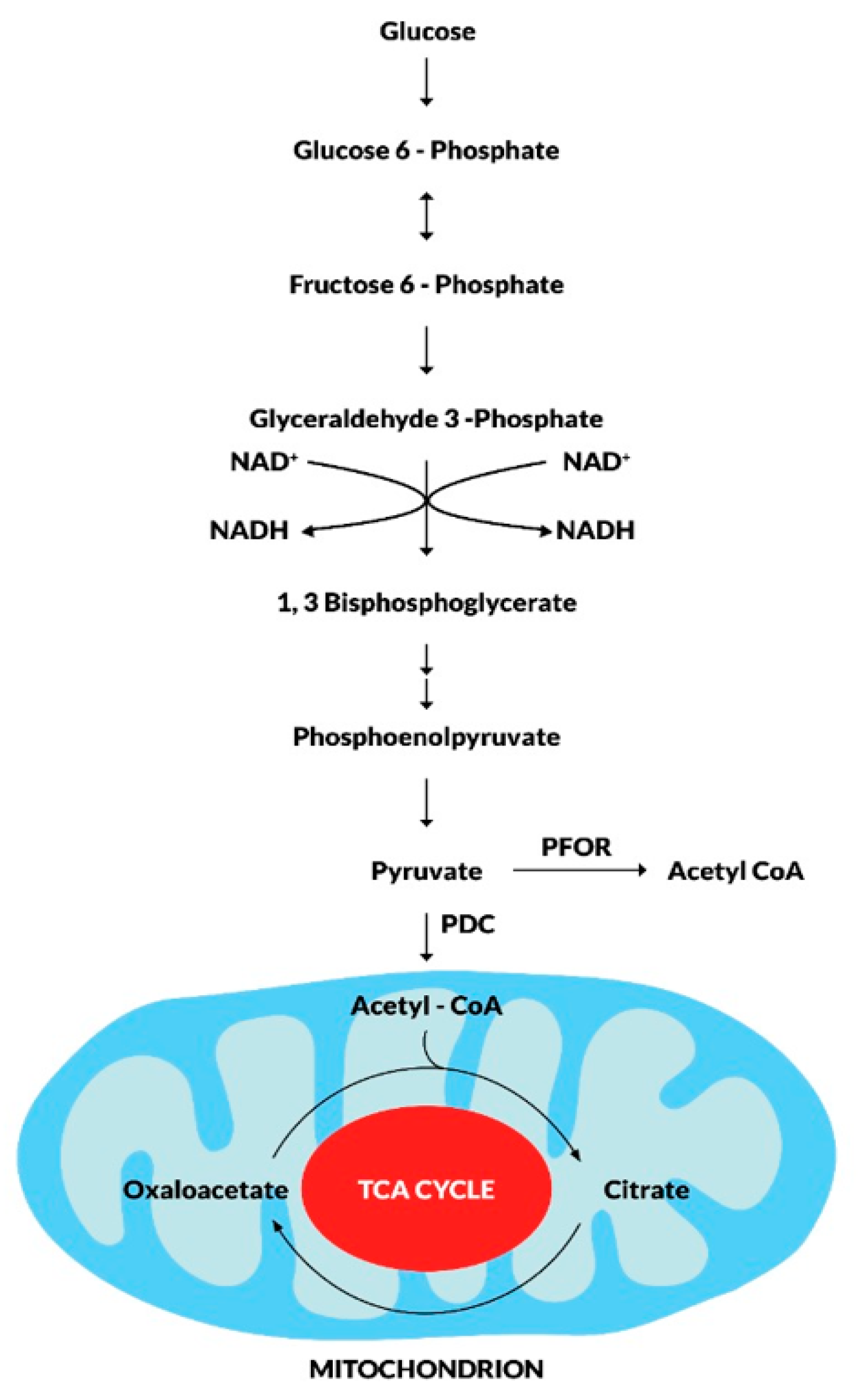
Metabolic fate of pyruvate in mitochondriate and some amitochondriate organisms.
Figure 4.
Metabolic fate of pyruvate in mitochondriate and some amitochondriate organisms.
During the fundamental metabolic pathway of glycolysis, glucose is converted to pyruvate along a series of phosphorylation and dehydrogenation reactions. The fate of pyruvate is mainly dependent on the presence of mitochondrion. In mitochondriate organisms, pyruvate is further dehydrogenated to acetyl CoA by the pyruvate dehydrogenase enzyme complex (PDC). Acetyl CoA enters the tricarboxylic acid cycle as the primary substrate for oxidative reactions that result in the production of CO2 and ATP molecules for intermediary metabolism. In some amitochondriate organisms, however, pyruvate undergoes fermentative decarboxylation to produce Acetyl CoA in a reversible redox reaction catalyzed by the pyruvate-ferredoxin oxidoreductase (PFOR).
Figure 5.
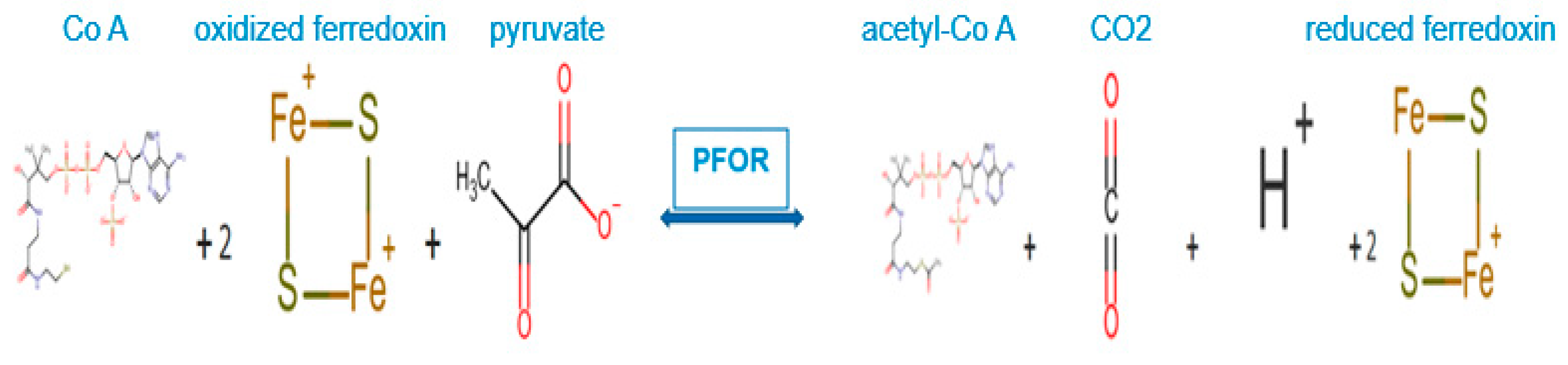
PFOR-mediated anaerobic metabolism of pyruvate.
Figure 5.
PFOR-mediated anaerobic metabolism of pyruvate.
In the fermentative decarboxylation of pyruvate, PFOR catalyzes the reduction of two molecules of ferredoxin using Co-enzyme A as a co-factor to produce acetyl-CoA, carbon dioxide and hydrogen ions. The reduced ferredoxin can be utilized as an electron donor for further redox reactions.
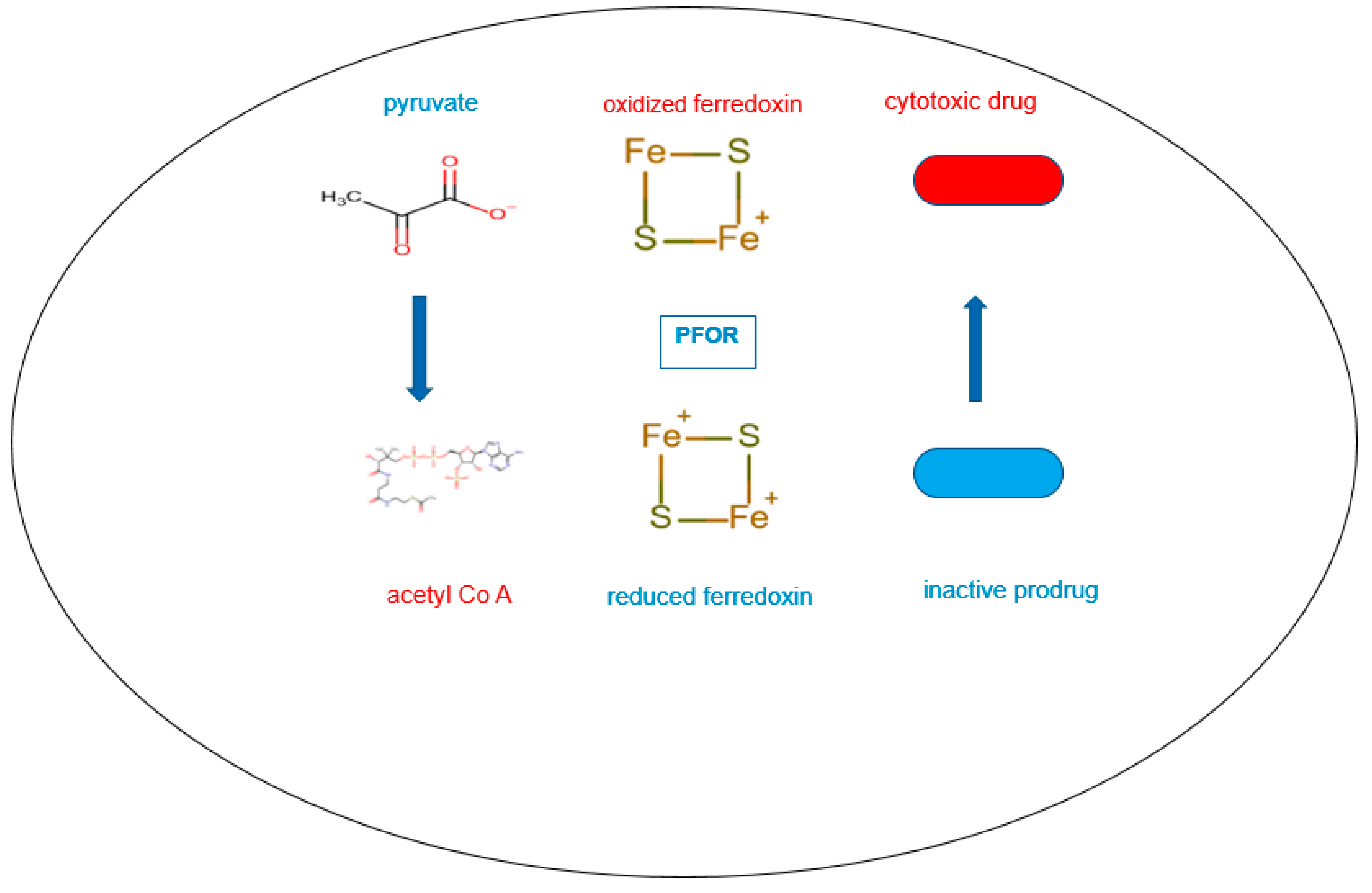
In the presence of nitro-based prodrugs, this same redox machinery is used as a drug target to make pathogens vulnerable to nitro-radical-induced cytotoxic damage. In this instance, the resulting reduced ferredoxin serves as an electron donor by transferring electrons to nitro-prodrugs. These then become pharmacologically activated and subsequently transformed into nitroso and hydroxylamine derivatives that cause cytotoxic damage (Horner et al. 1999, Ragsdale 2003).
Figure 6.
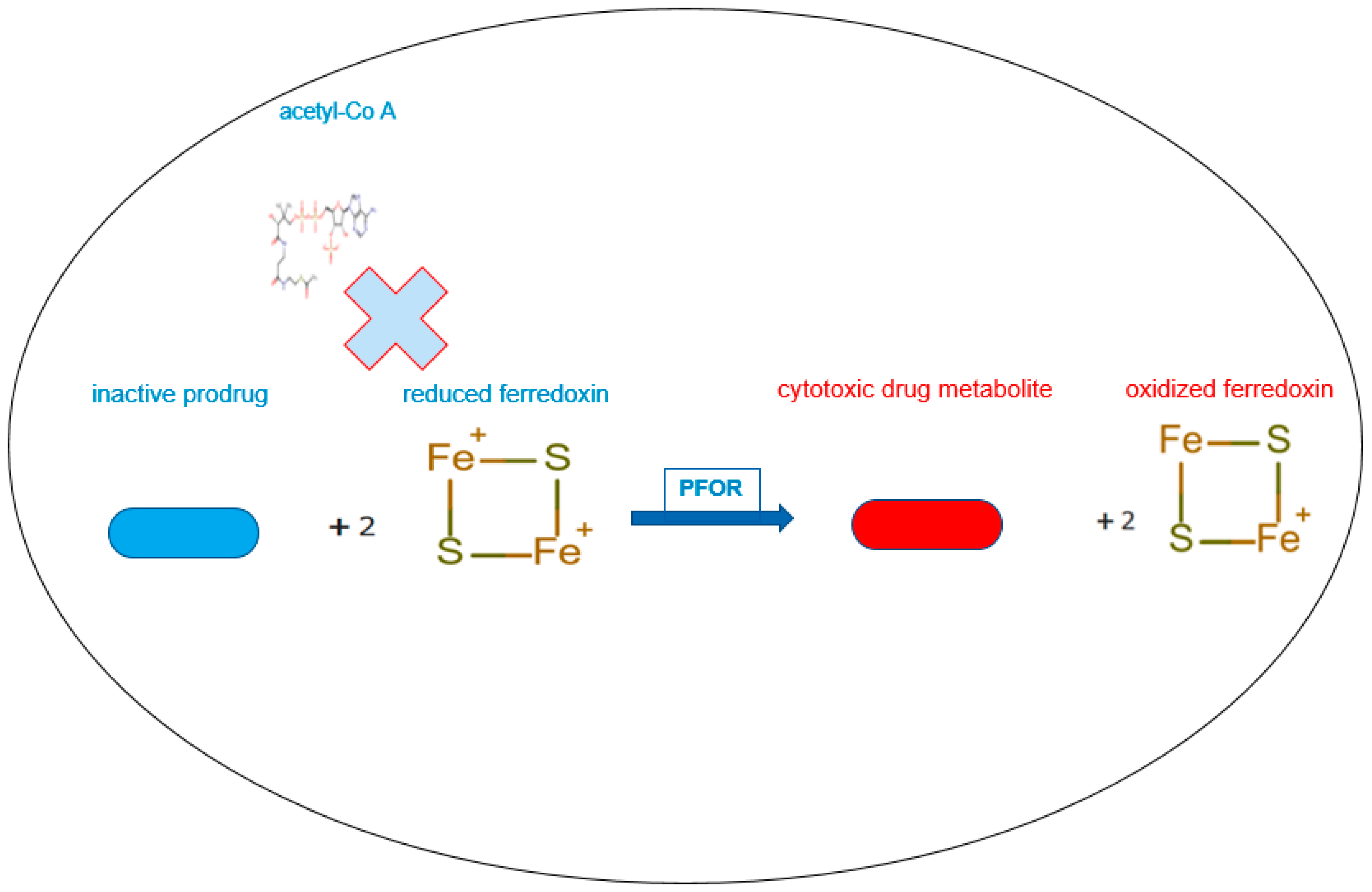
PFOR-mediated re-oxidation of ferredoxin and bio-activation of nitro-based prodrugs.
Figure 6.
PFOR-mediated re-oxidation of ferredoxin and bio-activation of nitro-based prodrugs.
In the presence of sufficient concentrations of the inactive prodrug, reduced ferredoxin donates electrons to the prodrug instead of acetyl-Co A by the catalytic activity of PFOR. In this reverse redox reaction, ferredoxin is re-oxidized and the prodrug is activated to its cytotoxic radical intermediate.
Figure 7.
A summary of the PFOR-mediated fermentative decarboxylation of pyruvate and reductive activation of nitro-based prodrugs.
Figure 7.
A summary of the PFOR-mediated fermentative decarboxylation of pyruvate and reductive activation of nitro-based prodrugs.
This figure depicts the reversible redox reaction catalyzed by PFOR in the presence of very high concentrations of the inactive prodrug. In the forward reaction, PFOR catalyzes the decarboxylation of pyruvate to acetyl-Co A using oxidized ferredoxin as electron acceptor. The resulting reduced ferredoxin then donates electrons to the inactive prodrug to reductively bio-activate it. This cycle continues until the concentration of the radical intermediates of the prodrug reaches therapeutic levels within the pathogen to disrupt nucleic acid and protein synthesis.
2.1.1. Activation of metronidazole by the PFOR of G. duodenalis and E. histolytica
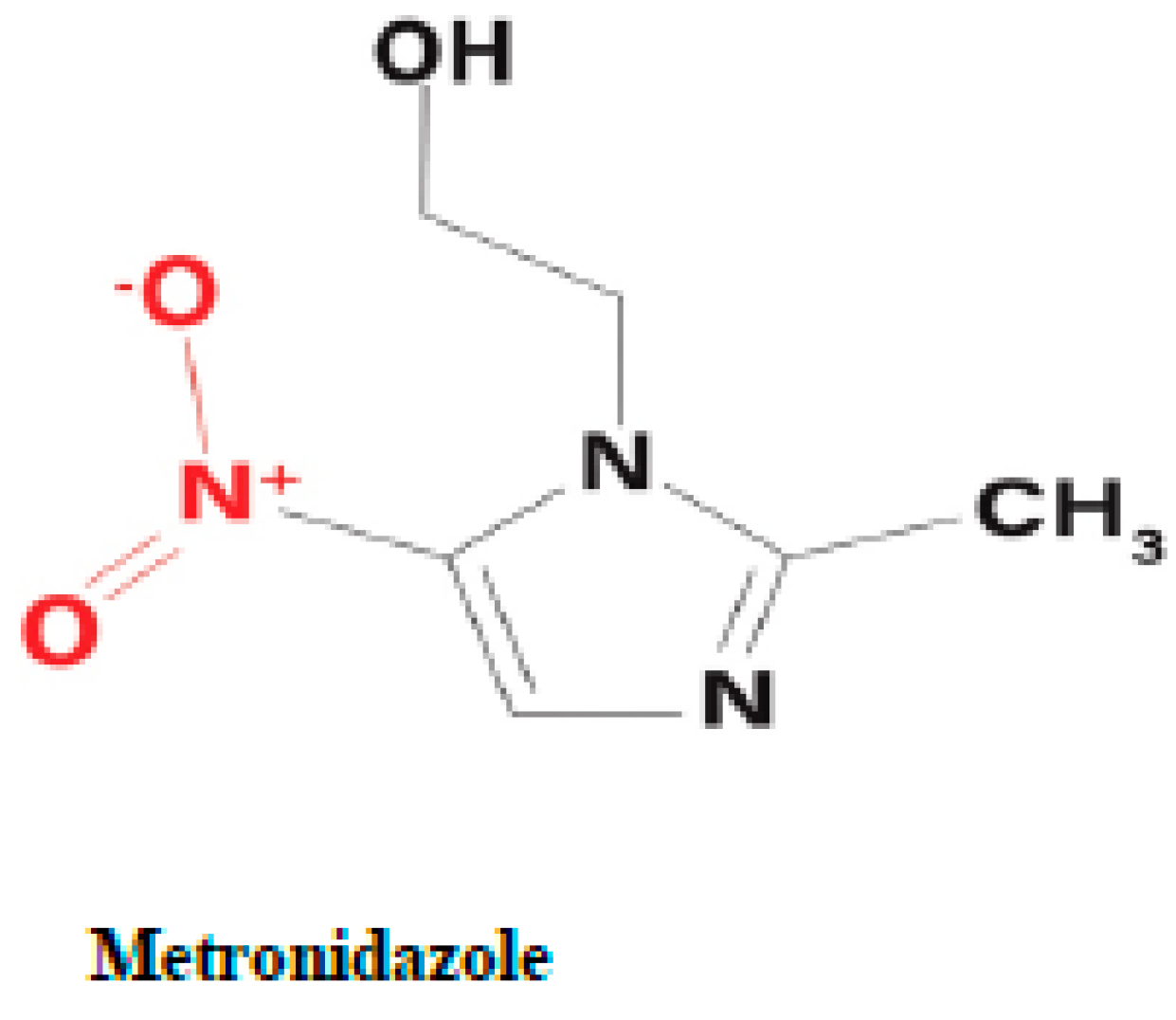
Metronidazole is a synthetic nitro-imidazole derivative effective against most obligate anaerobic bacteria and protozoan parasites such as
G. duodenalis,
E. histolytica,
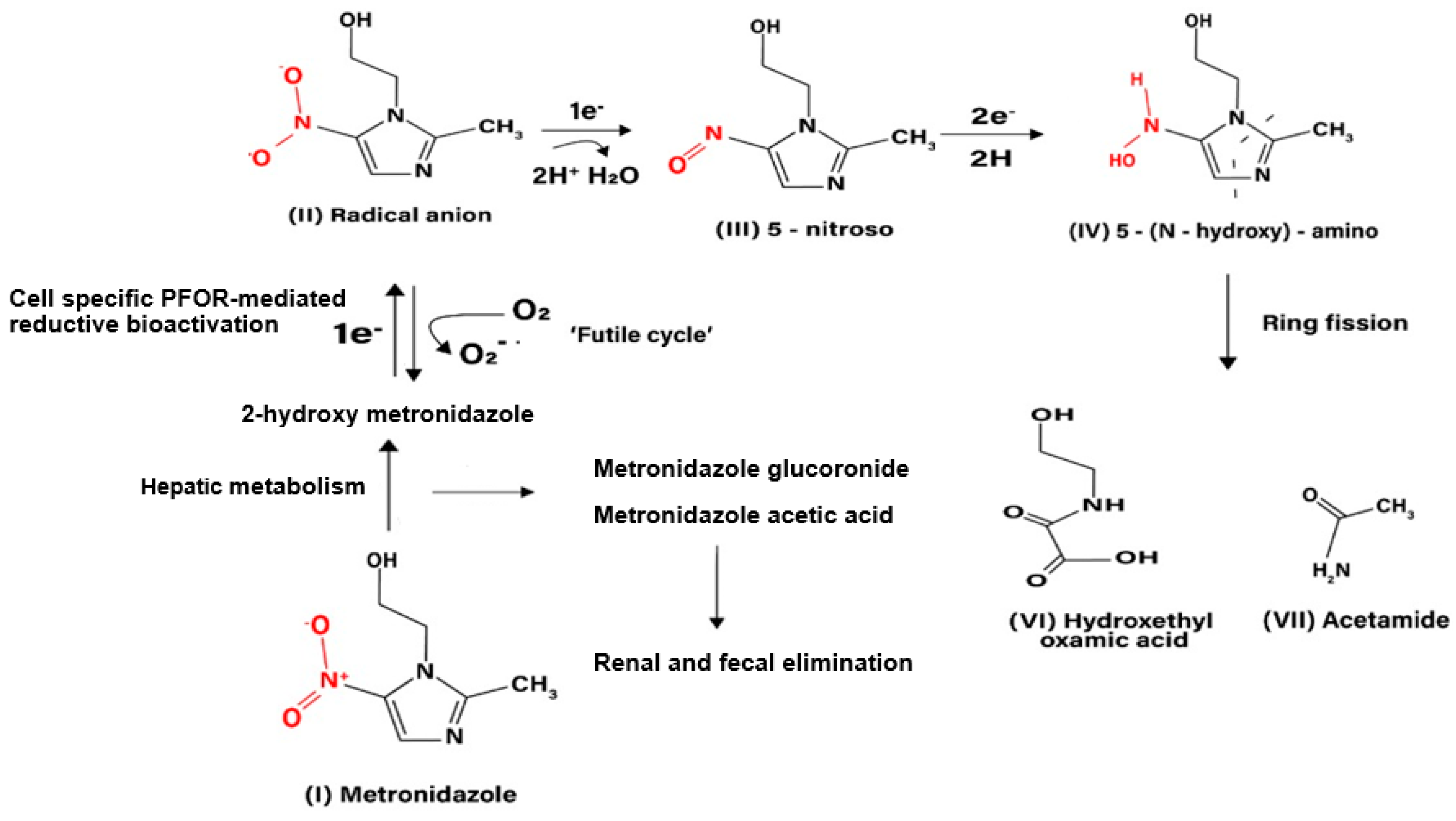
T. vaginalis (Upcroft et al. 2001, Pal et al. 2009). It exerts its antimicrobial effect by interfering with nucleic acid synthesis resulting in DNA damage in the target pathogens (Lamp et al. 1999). Electrons from the initial redox reaction catalyzed by PFOR are transferred to ferredoxin, which has a sufficiently low redox potential to reductively activate metronidazole to its toxic radical metabolites. Through a series of electron transfer reactions, different radical intermediates are formed that act primarily as alkylating agents that disrupt DNA synthesis and growth of invading pathogens (Dingsdag et al. 2018). The different radical metabolites of metronidazole have been illustrated in
Figure 8 below. Strains of
G. duodenalis and
E. histolytica with decreased or absent expression of
pfor have been shown to exhibit increased metronidazole resistance (Dan et al. 2000). However, some recent studies suggest the involvement of alternative enzymes that also play central roles in the activation of, and resistance to, metronidazole (Leitsch et al. 2011, Ansell et al. 2017, Müller et al. 2019, Lopes-Oliveira et al. 2020). For example,
T. vaginalis has been shown in some studies to display a complete resistance to metronidazole only after both PFOR and NAD-dependent malic enzyme are inactivated in the hydrogenosome (Rasoloson et al. 2002, Hrdý, Cammack et al. 2005).
The initial step in the bio-activation of metronidozole involves the transfer of one electron from ferredoxin to metronidazole to form its radical anion. In the presence of molecular oxygen the radical anion is re-oxidized to the parent compound in a futile cycle, hence, the overall bio-activation reaction is stalled at this stage. In anaerobic conditions, the radical anion recieves a further electron to form the 5-nitroso intermediate. In a subsequent reduction reaction involving the transfer of two electrons, the 5-nitroso intermediate is converted to a 5-(N–hydroxy)-amino intermediate, which then undergoes a spotaneous ring fision to produce hydroxethyl oxamic acid and acetamide. These fission products are the proposed end products of the bioactivation process that contribute immensely to the toxicity of metronidazole in invading pathogens.
2.1.2. Interaction between nitazoxanide and the PFOR of C. parvum
Nitazoxanide is a nitrothiazolyl-salicylamide derivative which exhibits broad-spectrum antimicrobial activity against a variety of pathogens including anaerobic and microaerophilic bacteria, protozoa, helminths, and viruses (Shakya et al. 2018, Bharti et al. 2021). Recent preclinical studies have demonstrated potential antineoplastic activity of nitazoxanide that can be further explored in the field of oncology (Pal et al. 2020, Abd El-Fadeal et al. 2021, Kiehl et al. 2021). The antiviral activity of nitazoxanide against the SARS-CoV-2 infection is currently being explored (Meneses et al. 2020, Lokhande et al. 2021, Rocco et al. 2021). Interestingly, nitazoxanide interacts with PFOR in a completely different manner than metronidazole. Some in-vitro studies postulate an interference of the redox function of PFOR by nitazoxanide (Hoffman et al. 2007, Scior et al. 2015). It deactivates the oxidative decarboxylation of pyruvate along with the electron transfer machinery that produces reduced ferredoxin in the presence of the coenzyme thiamine pyrophosphate (TPP) (Hoffman et al. 2007). This is the generally accepted mechanism by which it inactivates the metabolism, growth and survival of C. parvum (Gargala 2008). A similar mechanism of action is postulated for the interaction between the PFOR of some anaerobic bacteria and nitazoxanide (Warren et al. 2012, Kennedy et al. 2016).