Submitted:
24 November 2023
Posted:
24 November 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
- Resource and Strategic Planning: Predicting trial durations helps ensure optimal distribution of personnel and funds, minimizing inefficiencies. Furthermore, this foresight enables organizations to make informed decisions about trial prioritization, resource allocation, and initiation timelines [3,4].
- Patient Involvement & Safety: Estimating trial durations provides patients with clarity on their commitment, which safeguards their well-being and promotes informed participation [5].
- Transparent Relations with Regulators: Providing predictions on trial durations, whether below or above the average, fosters open communication with regulatory authorities. This strengthens compliance, builds trust, and establishes transparent relationships among all stakeholders [6].
2. Background
- Pioneering Work in Duration Prediction: Our model stands as a trailblazing effort in the domain, bridging the existing gap in duration prediction applications and establishing benchmarks for future research.
- Diverse Modeling: We extensively reviewed eight machine learning models, highlighting the Random Forest model for its unparalleled efficiency in predicting durations.
- Comprehensive Variable Exploration: Our model incorporates varied variables, from enrollment metrics to study patterns, enhancing its predictive capabilities.
- Insight into Data Volume: Beyond mere predictions, we delve into determining the optimal data volume required for precise forecasting.
- In-Depth Model Probability: Apart from binary predictions, our model associates higher probabilities with longer average durations, along with a 95% CI. This precision offers a comprehensive range of potential trial durations, aiding informed decision-making and strategic planning.
- Broad Applicability: With proven efficacy in lung cancer trials, our model showcases its potential use across various oncology areas.
3. Materials and Methods
3.1. Dataset
3.2. Data Preprocessing
3.3. Data Exploration and Feature Engineering
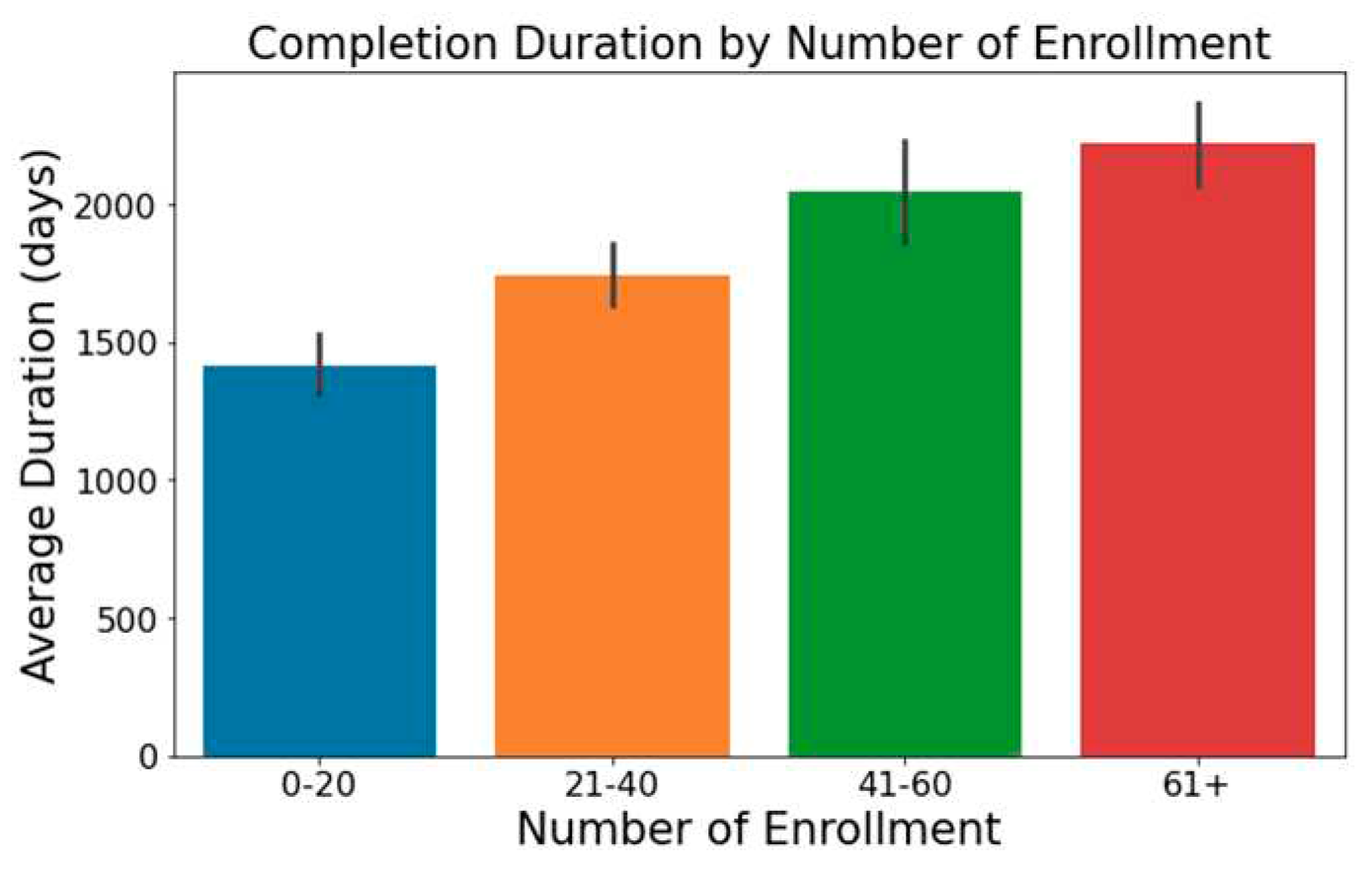
- Trials with increased enrollment often exhibit longer durations, as illustrated in Figure 1.
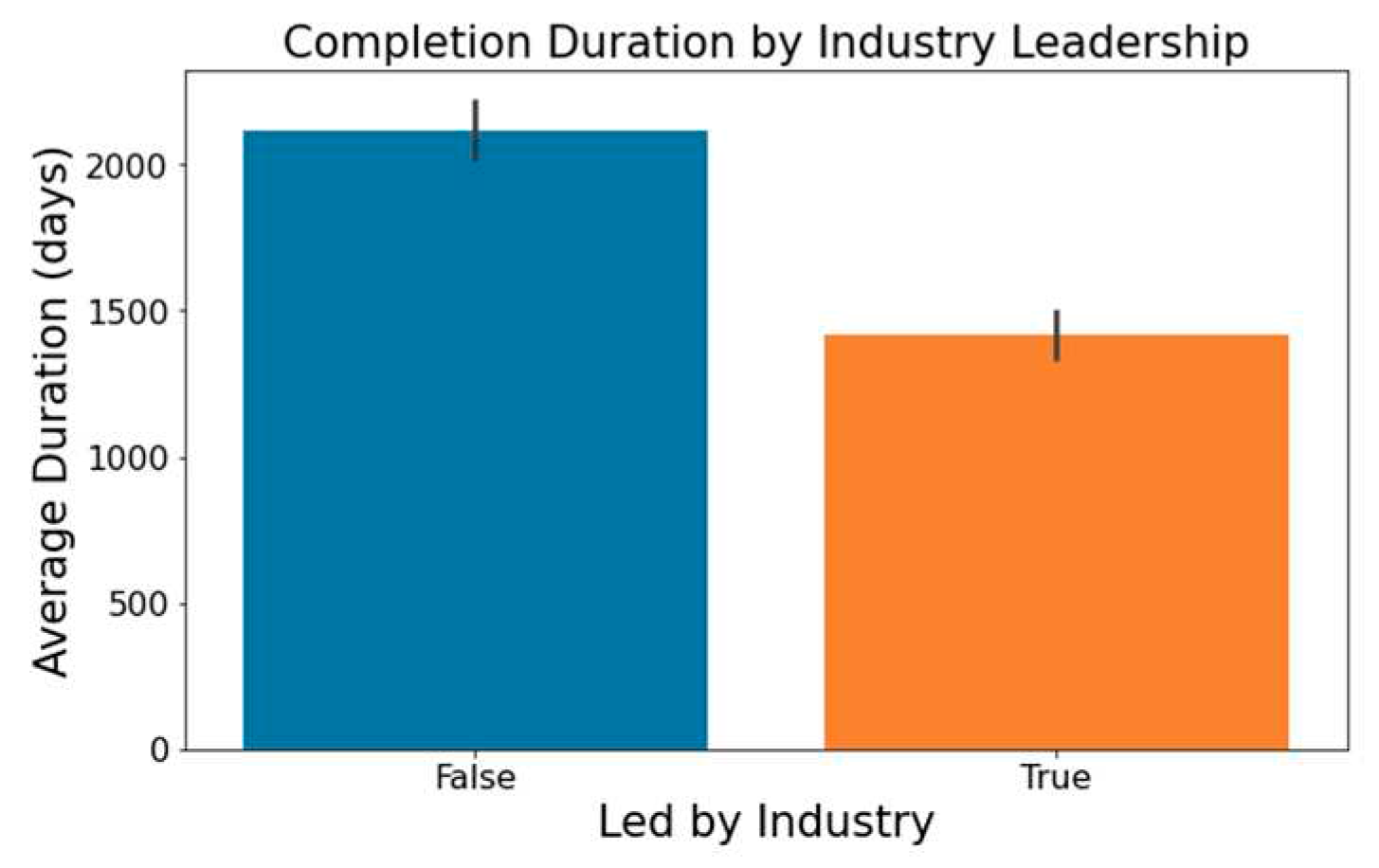
- Figure 2 highlights that industry-led trials tend to wrap up more swiftly than non-industry-led ones.
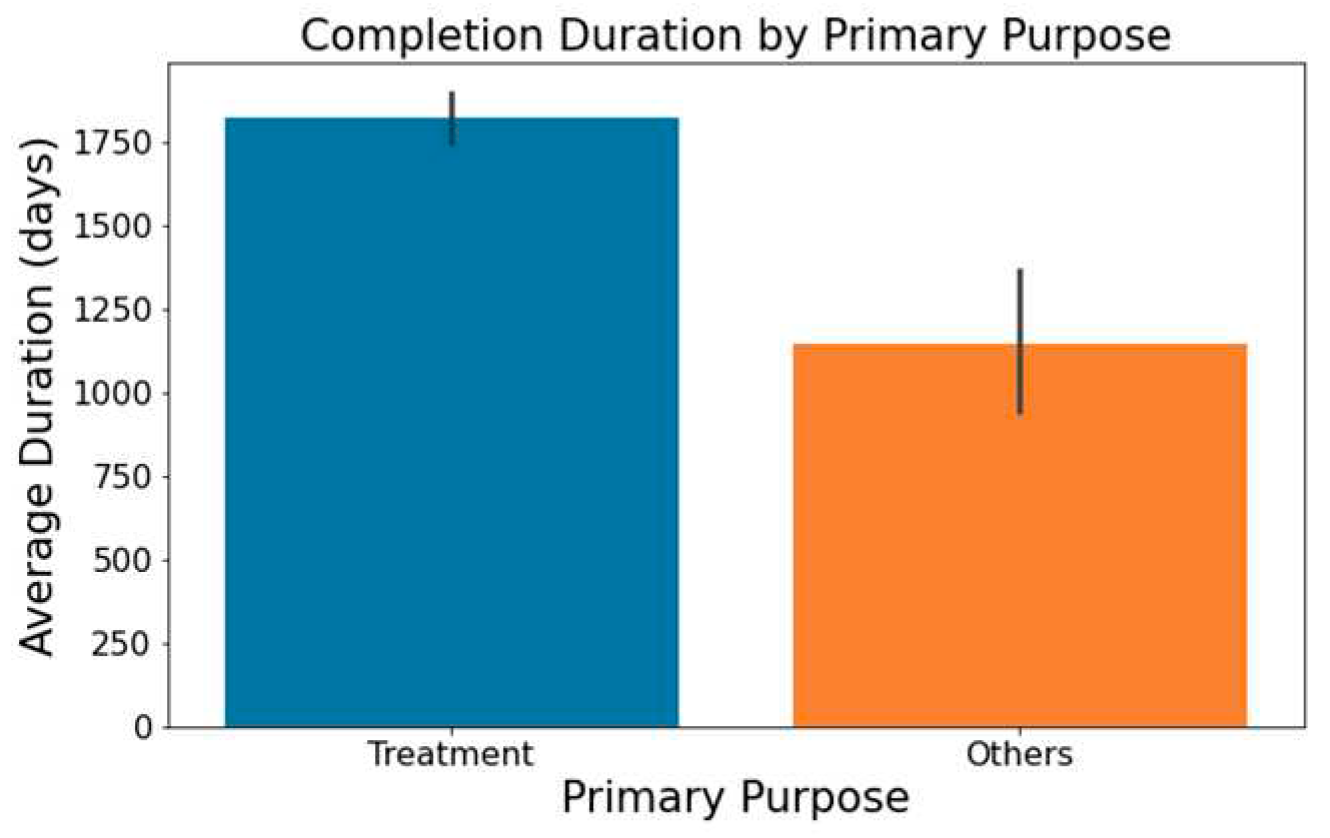
- As showcased in Figure 5, trials with a primary emphasis on 'Treatment' typically have longer durations than those aimed at 'Supportive Care,' 'Diagnostics,' 'Prevention,' or other areas.
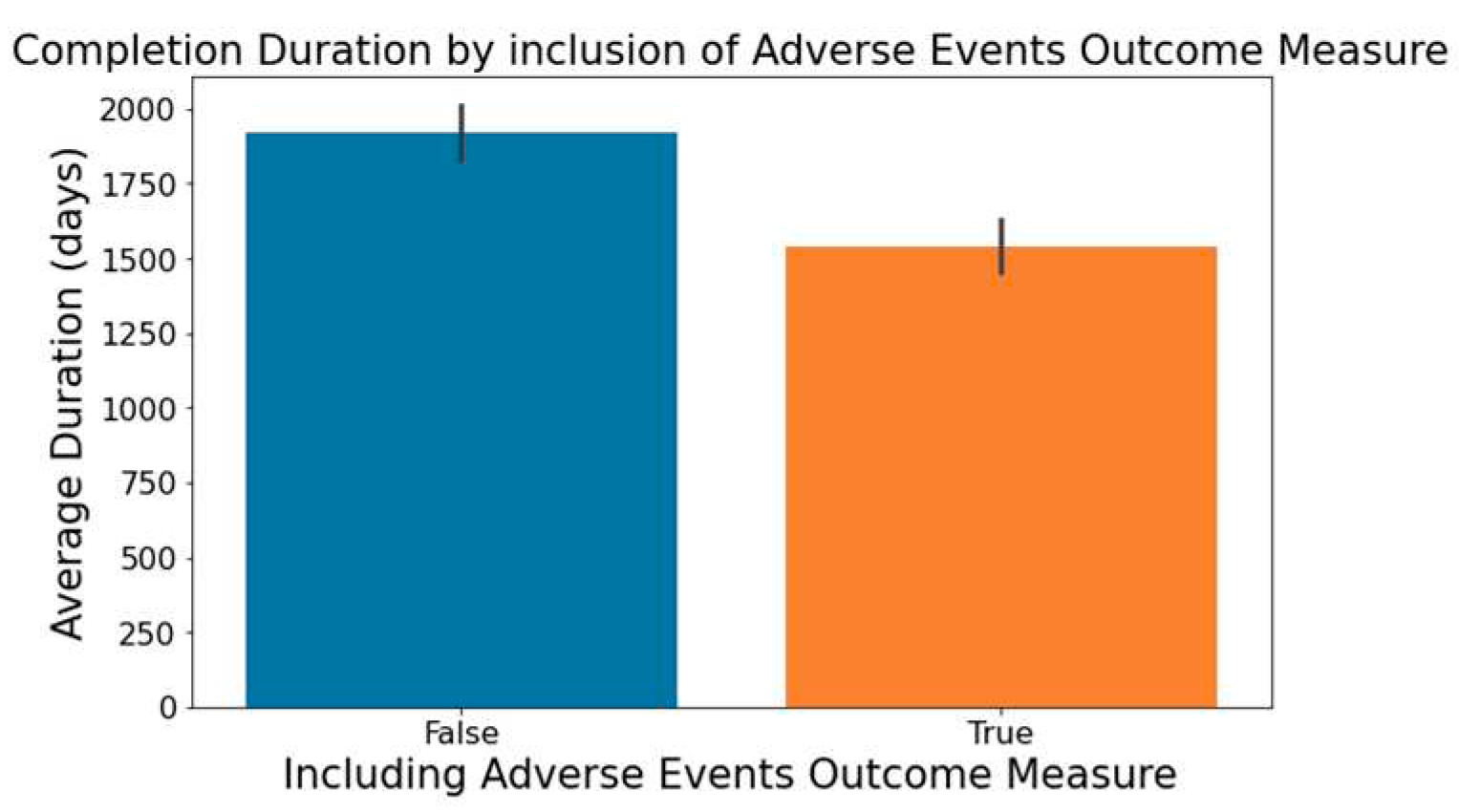
- Figure 6 demonstrates that trials focusing on the measurement of adverse events within the 'Outcome Measures' column tend to be completed faster.
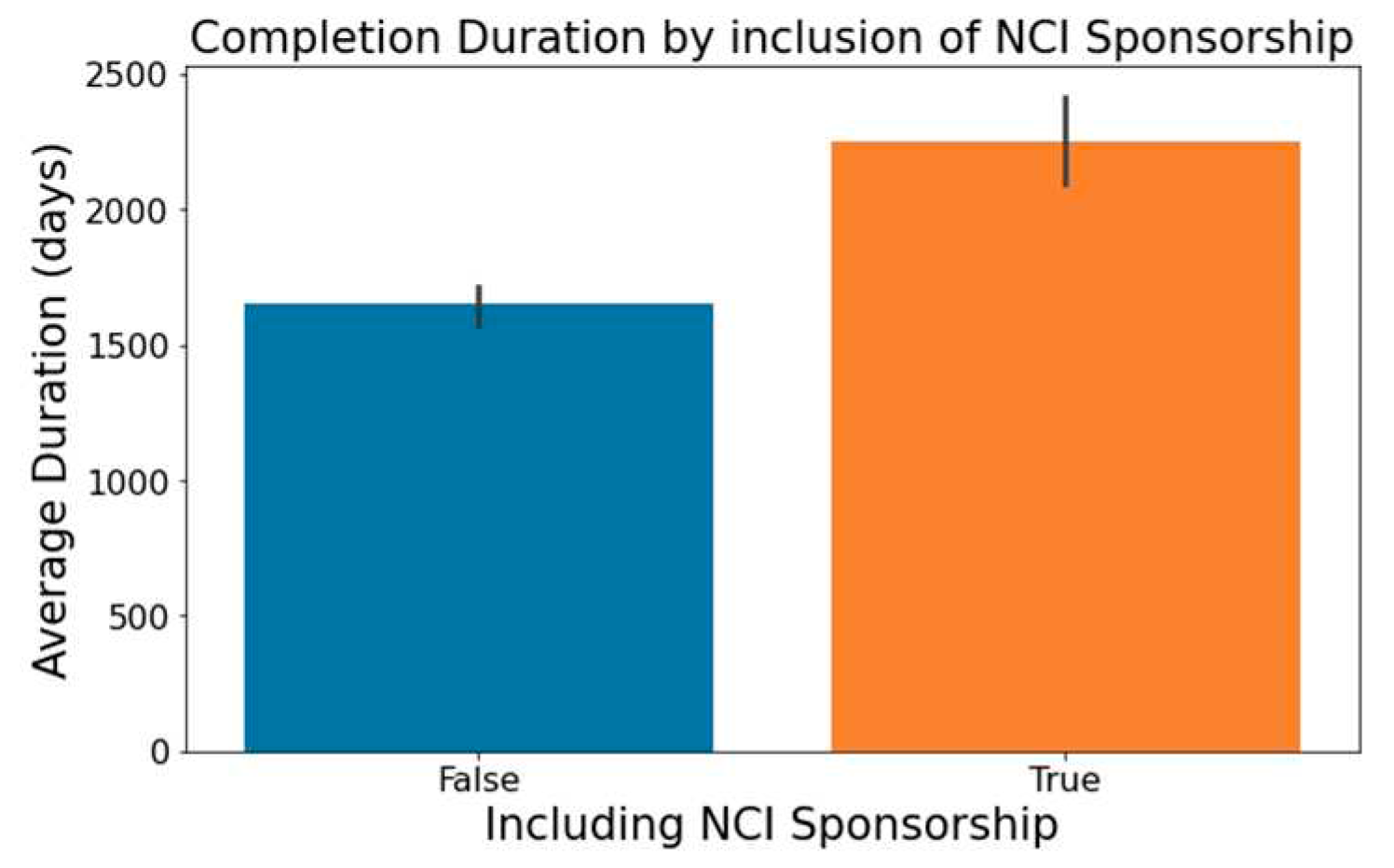
- Trials indicating 'National Cancer Institute (NCI)' in the 'Sponsor/Collaborators' column are observed to have lengthier durations, a trend captured in Figure 7.
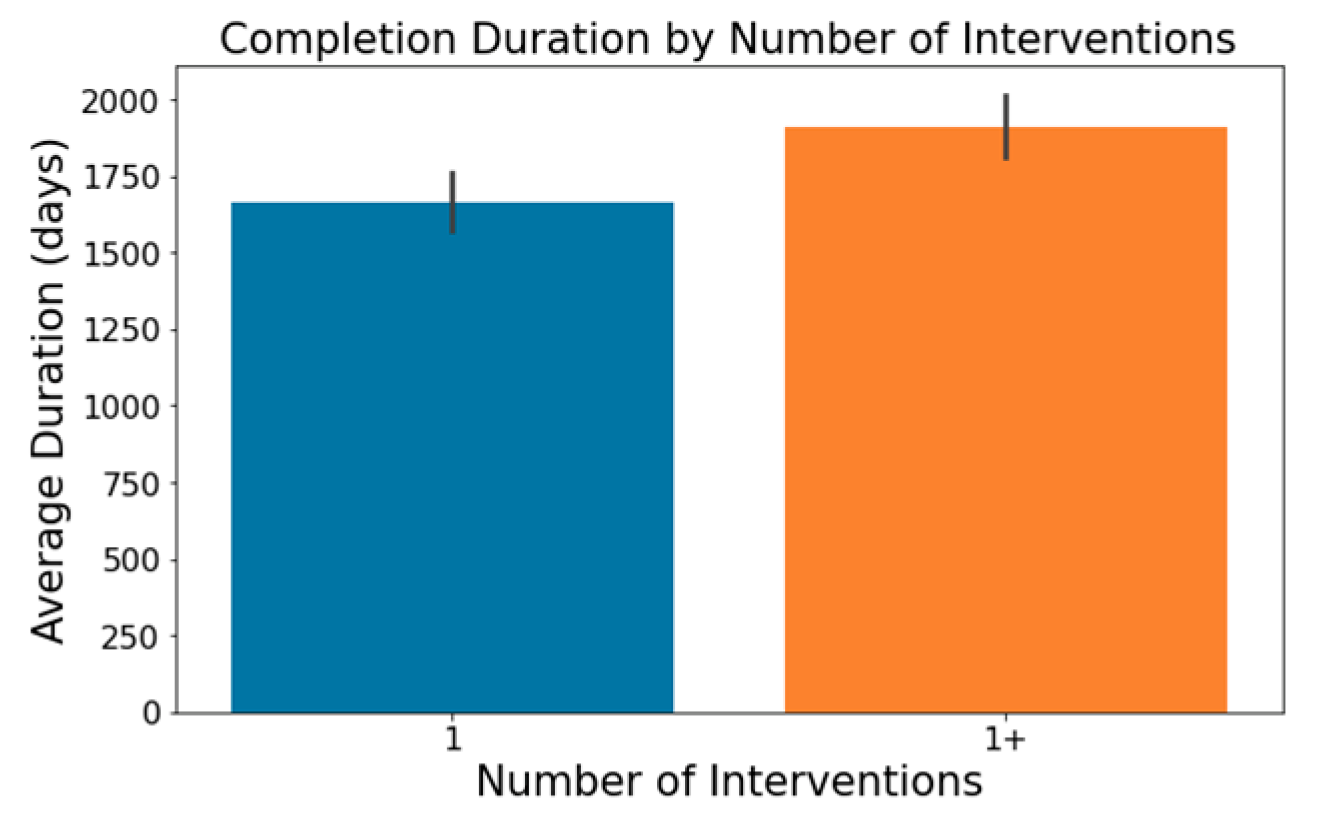
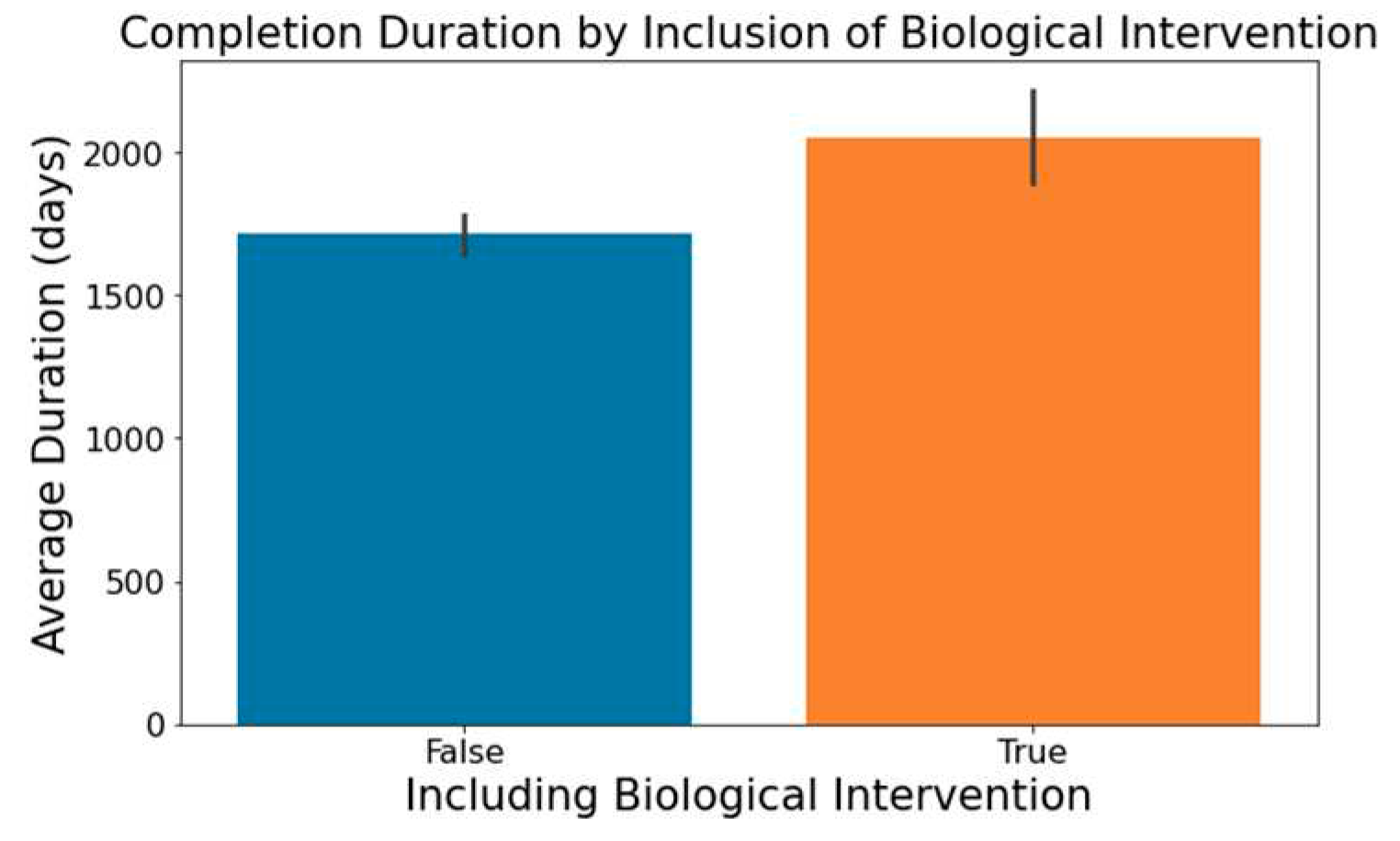
- The involvement of biological interventions in trials, represented in the ‘Interventions’ column, often results in extended durations, as seen in Figure 8.
3.4. Machine Learning Models and Evaluation Metrics
3.4.1. Logistic Regression (LR)
3.4.2. K-Nearest Neighbors (KNN)
3.4.3. Decision Tree (DT)
3.4.4. Random Forest (RF) & 3.4.5. XGBoost (XGB)
3.4.6. Linear Discriminant Analysis (LDA) & 3.4.7. Gaussian Naïve Bayes (Gaussian NB)
3.4.8. Multi-Layer Perceptron (MLP)
- Accuracy measures the fraction of correct predictions (See Equation (1)).
- ROC visually represents classifier performance by plotting recall against the false positive rate ((See Equation (2)) across diverse thresholds. This visual representation is condensed into a metric via the AUC; a value between 0 and 1, where 1 signifies flawless classification
- Precision gauges the reliability of positive classifications, shedding light on the inverse of the false positive rate (See Equation (3)).
- Recall (or sensitivity) denotes the fraction of actual positives correctly identified, emphasizing the influence of false negatives (See Equation (4)).
- F1-score provides a balance between precision and recall, acting as their harmonic mean (See Equation (5)).
4. Results and Discussion
4.1. Sample Characteristics
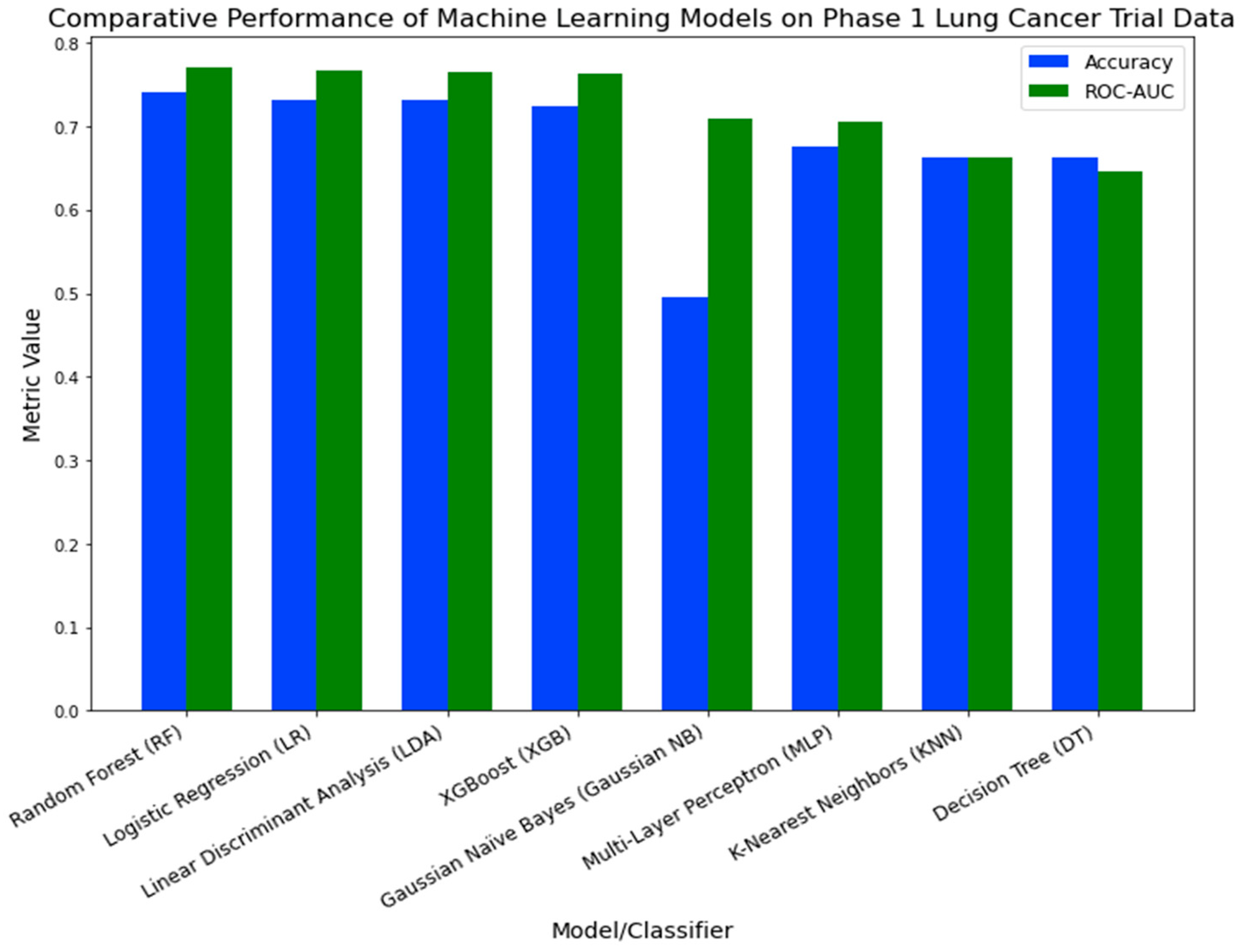
4.2 Machine Learning Classification
4.3. Random Forest Model Validation
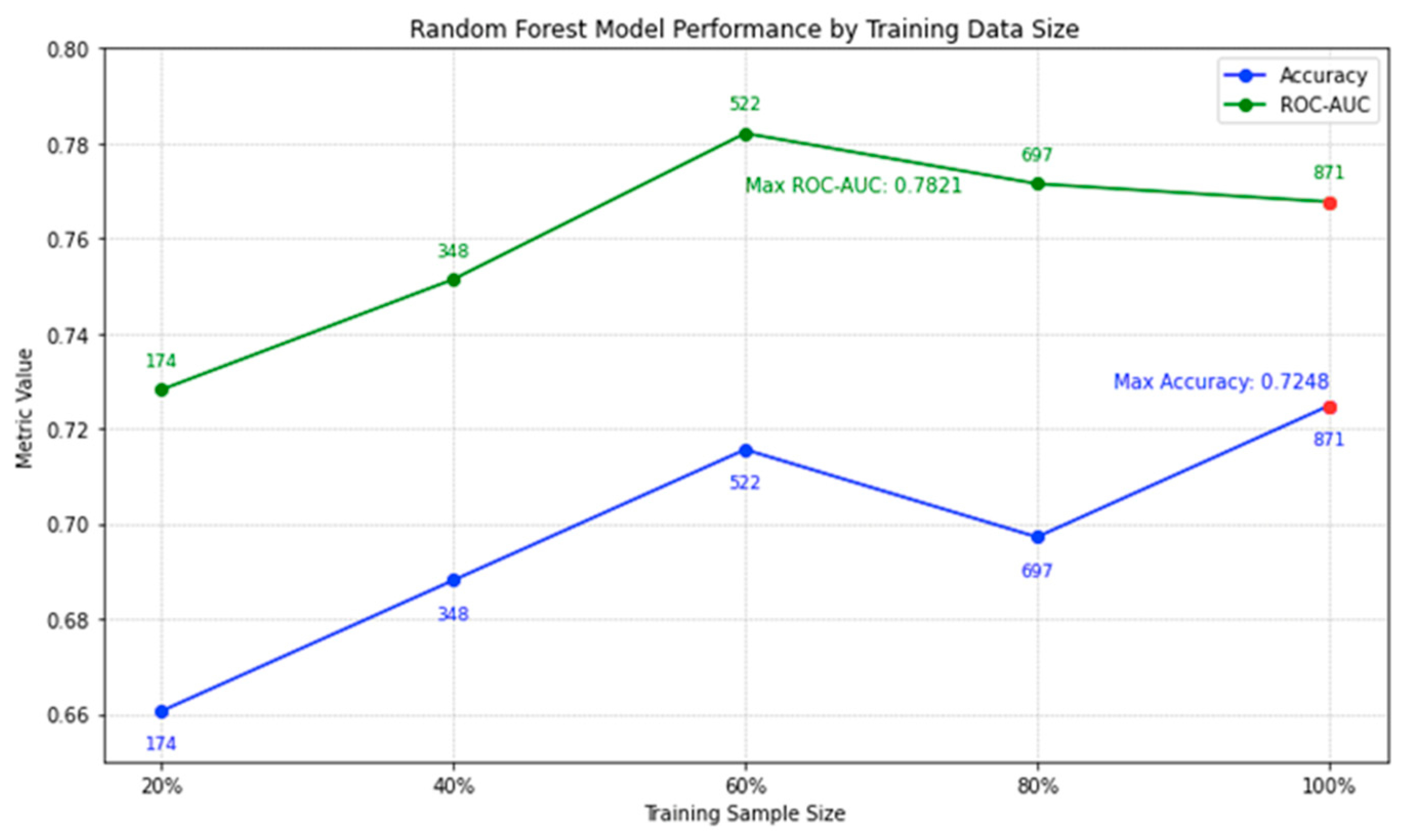
4.3.1. Impact of Varying Training Data Sizes on Model Performance
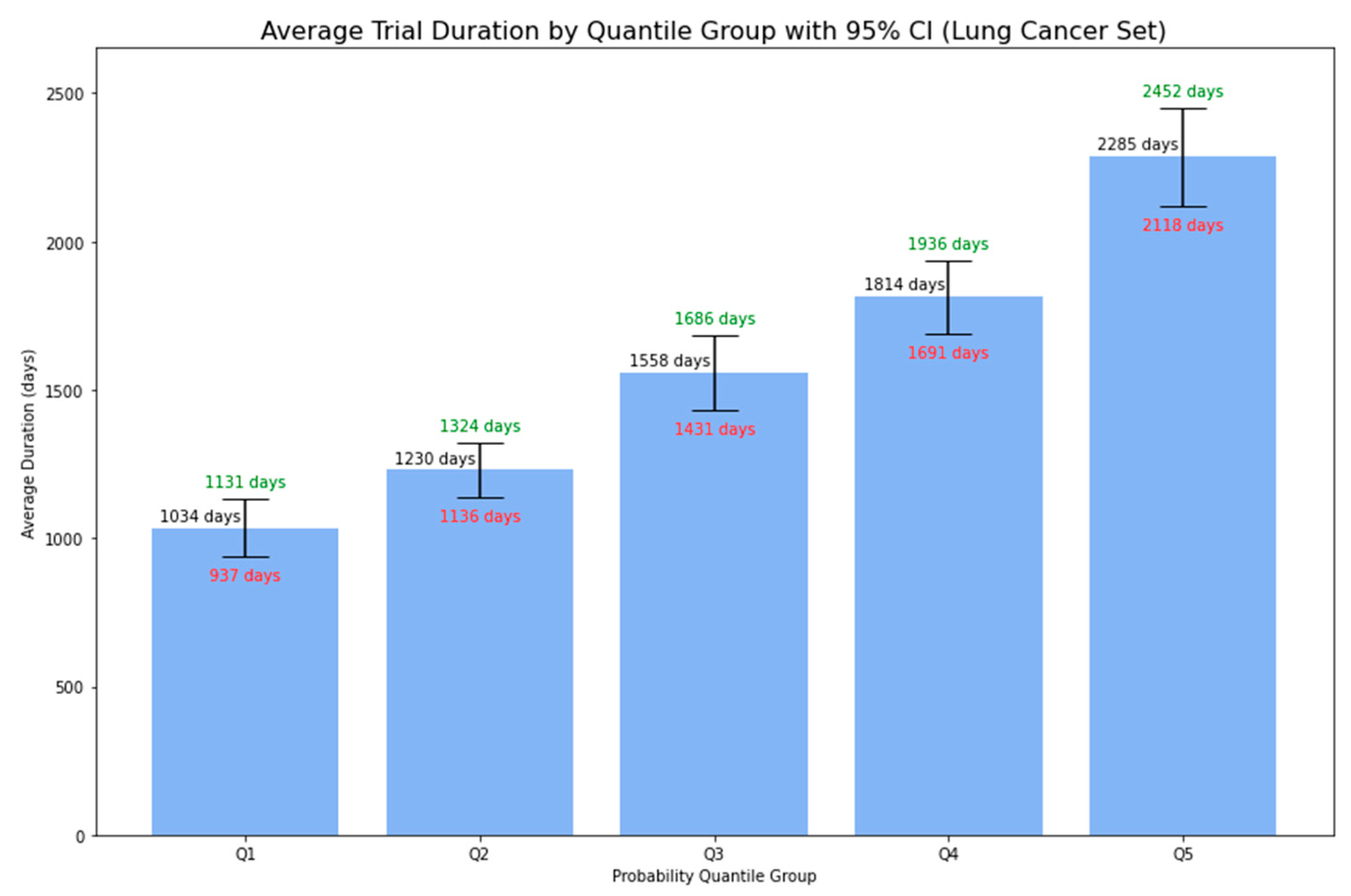
4.3.2. External Validation using Phase 1 Lung Cancer Trial Data
5. Conclusion
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- R. L. Siegel, K. D. Miller, N. S. Wagle, and A. Jemal, “Cancer statistics, 2023,” Ca Cancer J Clin, vol. 73, no. 1, pp. 17–48, 2023.
- T. G. Roberts et al., “Trends in the risks and benefits to patients with cancer participating in phase 1 clinical trials,” Jama, vol. 292, no. 17, pp. 2130–2140, 2004. [CrossRef]
- E. H. Weissler et al., “The role of machine learning in clinical research: transforming the future of evidence generation,” Trials, vol. 22, no. 1, p. 537, Dec. 2021. [CrossRef]
- K. Wu et al., “Machine Learning Prediction of Clinical Trial Operational Efficiency,” AAPS J., vol. 24, no. 3, p. 57, May 2022. [CrossRef]
- T. L. Beauchamp and J. F. Childress, Principles of biomedical ethics. Oxford University Press, USA, 2001. Accessed: Oct. 30, 2023. [Online]. Available: https://books.google.com/books?hl=en&lr=&id=_14H7MOw1o4C&oi=fnd&pg=PR9&dq=Beauchamp,+T.+L.,+%26+Childress,+J.+F.+(2013).+Principles+of+biomedical+ethics+(7th+ed.).+New+York:+Oxford+University+Press.&ots=1x_n4OBqWq&sig=pCzR4XfW0iDFmXEFsOajo6dGdU4.
- D. A. Dri, M. Massella, D. Gramaglia, C. Marianecci, and S. Petraglia, “Clinical Trials and Machine Learning: Regulatory Approach Review,” Rev. Recent Clin. Trials, vol. 16, no. 4, pp. 341–350, 2021. [CrossRef]
- R. K. Harrison, “Phase II and phase III failures: 2013–2015,” Nat. Rev. Drug Discov., vol. 15, no. 12, Art. no. 12, Dec. 2016. [CrossRef]
- Uniform, “How to avoid costly Clinical Research delays | Blog,” MESM. Accessed: Oct. 30, 2023. [Online]. Available: https://www.mesm.com/blog/tips-to-help-you-avoid-costly-clinical-research-delays/.
- E. W. Steyerberg, Clinical Prediction Models: A Practical Approach to Development, Validation, and Updating. in Statistics for Biology and Health. Cham: Springer International Publishing, 2019. [CrossRef]
- D. J. Sargent, B. A. Conley, C. Allegra, and L. Collette, “Clinical trial designs for predictive marker validation in cancer treatment trials,” J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol., vol. 23, no. 9, pp. 2020–2027, Mar. 2005. [CrossRef]
- I. Kola and J. Landis, “Can the pharmaceutical industry reduce attrition rates?,” Nat. Rev. Drug Discov., vol. 3, no. 8, Art. no. 8, Aug. 2004. [CrossRef]
- E. W. Steyerberg and Y. Vergouwe, “Towards better clinical prediction models: seven steps for development and an ABCD for validation,” Eur. Heart J., vol. 35, no. 29, pp. 1925–1931, Aug. 2014. [CrossRef]
- S. Mandrekar and D. Sargent, “Clinical Trial Designs for Predictive Biomarker Validation: One Size Does Not Fit All,” J. Biopharm. Stat., vol. 19, pp. 530–42, Feb. 2009. [CrossRef]
- P. Blanche, J.-F. Dartigues, and H. Jacqmin-Gadda, “Estimating and comparing time-dependent areas under receiver operating characteristic curves for censored event times with competing risks,” Stat. Med., vol. 32, no. 30, pp. 5381–5397, Dec. 2013. [CrossRef]
- P. J. Rousseeuw and A. M. Leroy, Robust regression and outlier detection. John wiley & sons, 2005. Accessed: Oct. 30, 2023. [Online]. Available: https://books.google.com/books?hl=en&lr=&id=woaH_73s-MwC&oi=fnd&pg=PR13&dq=Rousseeuw,+P.J.,+Leroy,+A.M.+(1987).+Robust+Regression+and+Outlier+Detection.+John+Wiley+%26+Sons.&ots=TCuOR_zkjR&sig=pwLEHKv7QboOplfEIV0LO6POvdY.
- T. Hastie, J. Friedman, and R. Tibshirani, The Elements of Statistical Learning. in Springer Series in Statistics. New York, NY: Springer New York, 2001. [CrossRef]
- C. M. Bishop and N. M. Nasrabadi, Pattern recognition and machine learning, vol. 4. Springer, 2006. Accessed: Oct. 30, 2023. [Online]. Available: https://link.springer.com/book/9780387310732.
- J. Fox, Applied regression analysis and generalized linear models, 2nd ed. in Applied regression analysis and generalized linear models, 2nd ed. Thousand Oaks, CA, US: Sage Publications, Inc, 2008, pp. xxi, 665.
- E. Schwager et al., “Utilizing machine learning to improve clinical trial design for acute respiratory distress syndrome,” Npj Digit. Med., vol. 4, no. 1, Art. no. 1, Sep. 2021. [CrossRef]
- E. Kavalci and A. Hartshorn, “Improving clinical trial design using interpretable machine learning based prediction of early trial termination,” Sci. Rep., vol. 13, no. 1, Art. no. 1, Jan. 2023. [CrossRef]
- S. Harrer, P. Shah, B. Antony, and J. Hu, “Artificial Intelligence for Clinical Trial Design,” Trends Pharmacol. Sci., vol. 40, no. 8, pp. 577–591, Aug. 2019. [CrossRef]
- T. Cai et al., “Improving the Efficiency of Clinical Trial Recruitment Using an Ensemble Machine Learning to Assist With Eligibility Screening,” ACR Open Rheumatol., vol. 3, no. 9, pp. 593–600, 2021. [CrossRef]
- J. Vazquez, S. Abdelrahman, L. M. Byrne, M. Russell, P. Harris, and J. C. Facelli, “Using supervised machine learning classifiers to estimate likelihood of participating in clinical trials of a de-identified version of ResearchMatch,” J. Clin. Transl. Sci., vol. 5, no. 1, p. e42, Jan. 2021. [CrossRef]
- A. M. Chekroud et al., “Cross-trial prediction of treatment outcome in depression: a machine learning approach,” Lancet Psychiatry, vol. 3, no. 3, pp. 243–250, Mar. 2016. [CrossRef]
- A. V. Schperberg, A. Boichard, I. F. Tsigelny, S. B. Richard, and R. Kurzrock, “Machine learning model to predict oncologic outcomes for drugs in randomized clinical trials,” Int. J. Cancer, vol. 147, no. 9, pp. 2537–2549, 2020. [CrossRef]
- L. Tong, J. Luo, R. Cisler, and M. Cantor, “Machine Learning-Based Modeling of Big Clinical Trials Data for Adverse Outcome Prediction: A Case Study of Death Events,” in 2019 IEEE 43rd Annual Computer Software and Applications Conference (COMPSAC), Jul. 2019, pp. 269–274. [CrossRef]
- E. Batanova, I. Birmpa, and G. Meisser, “Use of Machine Learning to classify clinical research to identify applicable compliance requirements,” Inform. Med. Unlocked, vol. 39, p. 101255, Jan. 2023. [CrossRef]
- "ClinicalTrials.gov," National Library of Medicine, 2023. [Online]. Available: https://clinicaltrials.gov/. [Accessed: July 25, 2023].
- M. Honnibal and I. Montani, “spaCy 2: Natural language understanding with Bloom embeddings, convolutional neural networks and incremental parsing,” 2017.
- A. Yadav, H. Shokeen, and J. Yadav, “Disjoint Set Union for Trees,” in 2021 12th International Conference on Computing Communication and Networking Technologies (ICCCNT), IEEE, 2021, pp. 1–6. Accessed: Oct. 30, 2023. [Online]. Available: https://ieeexplore.ieee.org/abstract/document/9580066/.
- L. Breiman, J. Friedman, R. Olshen, and C. Stone, “Classification and regression trees–crc press,” Boca Raton Fla., 1984.
- F. Pedregosa et al., “Scikit-learn: Machine learning in Python,” J. Mach. Learn. Res., vol. 12, no. Oct, pp. 2825–2830, 2011.
- T. Chen and C. Guestrin, “XGBoost: A Scalable Tree Boosting System,” in Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, in KDD ’16. New York, NY, USA: ACM, 2016, pp. 785–794. [CrossRef]
- M. K. Hasan, M. A. Alam, S. Roy, A. Dutta, M. T. Jawad, and S. Das, “Missing value imputation affects the performance of machine learning: A review and analysis of the literature (2010–2021),” Informatics in Medicine Unlocked, vol. 27, 2021. [CrossRef]
- Y. Wu, Q. Zhang, Y. Hu, K. Sun-Woo, X. Zhang, H. Zhu, L. jie, and S. Li, “Novel binary logistic regression model based on feature transformation of XGBoost for type 2 Diabetes Mellitus prediction in healthcare systems,” Future Generation Computer Systems, vol. 129, pp. 1-12, 2022. [CrossRef]
- N. S. Rajliwall, R. Davey, and G. Chetty, “Cardiovascular Risk Prediction Based on XGBoost,” in 2018 5th Asia-Pacific World Congress on Computer Science and Engineering (APWC on CSE), Nadi, Fiji, 2018, pp. 246-252. [CrossRef]
- B. Long, F. Tan, and M. Newman, “Ensemble DeBERTa Models on USMLE Patient Notes Automatic Scoring using Note-based and Character-based approaches,” Advances in Engineering Technology Research, vol. 6, no. 1, pp. 107-107, 2023. [CrossRef]
- V. Barnett and T. Lewis, Outliers in statistical data, vol. 3. Wiley New York, 1994. Accessed: Oct. 30, 2023. [Online]. Available: https://scholar.archive.org/work/l4rvge57snh7fjjzpc5idiyxj4/access/wayback/http://tocs.ulb.tu-darmstadt.de:80/214880745.pdf.
- K. Maheswari, P. Packia Amutha Priya, S. Ramkumar, and M. Arun, “Missing Data Handling by Mean Imputation Method and Statistical Analysis of Classification Algorithm,” in EAI International Conference on Big Data Innovation for Sustainable Cognitive Computing, A. Haldorai, A. Ramu, S. Mohanram, and C. C. Onn, Eds., in EAI/Springer Innovations in Communication and Computing. Cham: Springer International Publishing, 2020, pp. 137–149. [CrossRef]
| Rank | 47 |
|---|---|
| NCT Number | NCT02220842 |
| Title | A Safety and Pharmacology Study of Atezolizumab (MPDL3280A) Administered With Obinutuzumab or Tazemetostat in Participants With Relapsed/Refractory Follicular Lymphoma and Diffuse Large B-cell Lymphoma |
| Acronym | |
| Status | Completed |
| Study Results | No Results Available |
| Conditions | Lymphoma |
| Interventions | Drug: Atezolizumab|Drug: Obinutuzumab|Drug: Tazemetostat |
| Outcome Measures | Percentage of Participants With Dose Limiting Toxicities (DLTs)|Recommended Phase 2 Dose (RP2D) of Atezolizumab|Obinutuzumab Minimum Serum Concentration (Cmin)|Percentage of Participants With Adverse Events (AEs) Graded According to the National Cancer Institute (NCI) Common Terminology Criteria for Adverse Events version 4.0 (CTCAE v4.0)... |
| Sponsor/Collaborators | Hoffmann-La Roche |
| Gender | All |
| Age | 18 Years and older   (Adult, Older Adult) |
| Phases | Phase 1 |
| Enrollment | 96 |
| Funded Bys | Industry |
| Study Type | Interventional |
| Study Designs | Allocation: Non-Randomized|Intervention Model: Parallel Assignment|Masking: None (Open Label)|Primary Purpose: Treatment |
| Other IDs | GO29383|2014-001812-21 |
| Start Date | 18-Dec-14 |
| Primary Completion Date | 21-Jan-20 |
| Completion Date | 21-Jan-20 |
| First Posted | 20-Aug-14 |
| Results First Posted | |
| Last Update Posted | 27-Jan-20 |
| Locations | City of Hope National Medical Center, Duarte, California, United States|Fort Wayne Neurological Center, Fort Wayne, Indiana, United States|Hackensack University Medical Center, Hackensack, New Jersey, United States… |
| Study Documents | |
| URL | https://ClinicalTrials.gov/show/NCT02220842 |
| Feature Name | Explanation |
|---|---|
| Enrollment | Number of trial participants |
| Industry-led | Trial led by the industry (True/False) |
| Location Count | Number of trial locations |
| Measures Count | Number of outcome measures |
| Condition Count | Number of medical conditions |
| Intervention Count | Number of interventions |
| NCI Sponsorship | Sponsorship includes NCI (True/False) |
| AES Outcome Measure | Outcome measure includes adverse events (True/False) |
| Open Masking Label | Trial uses open masking label (True/False) |
| Biological Intervention | Intervention type includes biological (True/False) |
| Efficacy Keywords | Title includes efficacy-related keywords (True/False) |
| Random Allocation | Patient allocation is random (True/False) |
| US-led | Trial primarily in the US (True/False) |
| Procedure Intervention | Intervention type includes procedure (True/False) |
| Overall Survival Outcome Measure | Outcome measure includes overall survival rate (True/False) |
| Drug Intervention | Intervention type includes drugs (True/False) |
| MTD Outcome Measure | Outcome measure includes maximally tolerated dose (True/False) |
| US-included | Trial location includes the US (True/False) |
| DOR Outcome Measure | Outcome measure includes duration of response (True/False) |
| Prevention Purpose | Primary purpose is prevention (True/False) |
| AES Outcome Measure (Lead) | Leading outcome measure is adverse events (True/False) |
| DLT Outcome Measure | Outcome measure includes dose-limiting toxicity (True/False) |
| Treatment Purpose | Primary purpose is treatment (True/False) |
| DLT Outcome Measure (Lead) | Leading outcome measure is dose-limiting toxicity (True/False) |
| MTD Outcome Measure (Lead) | Leading outcome measure is maximally tolerated dose (True/False) |
| Radiation Intervention | Intervention type includes radiation (True/False) |
| Tmax Outcome Measure | Outcome measure includes time of Cmax (True/False) |
| Cmax Outcome Measure | Outcome measure includes maximum measured concentration (True/False) |
| Non-Open Masking Label | Trial use non-open masking label (True/False) |
| Crossover Assignment | Patient assignment is crossover (True/False) |
| Characteristics | Training/Cross-Validation Sets (n=871) | Testing Set (n=218) |
|---|---|---|
| Percentage of Trials Exceeding 5-Year Completion Time (Target) | 40% | 40% |
| Mean Trial Participant Enrollment | 49 | 50 |
| Percentage of Industry-led Trials | 46% | 48% |
| Average Number of Trial Locations | 6 | 6 |
| Average Outcome Measures Count | 6 | 6 |
| Average Medical Conditions Addressed | 4 | 4 |
| Average Interventions per Trial | 3 | 2 |
| Percentage of NCI-Sponsored Trials | 23% | 24% |
| Percentage of Trials with AES Outcome Measure | 34% | 34% |
| Percentage of Trials with Open Label Masking | 91% | 92% |
| Percentage of Titles Suggesting Efficacy | 50% | 51% |
| Percentage of Trials Involving Biological Interventions | 23% | 20% |
| Percentage of Randomly Allocated Patient Trials | 24% | 27% |
| Models/Classifier | Accuracy | ROC-AUC | Precision | Recall | F1-Score |
|---|---|---|---|---|---|
| XGBoost (XGB) | 0.7442±0.0384 | 0.7854±0.0389 | 0.7009±0.0439 | 0.6286±0.0828 | 0.6614±0.0633 |
| Random Forest (RF) | 0.7371±0.0389 | 0.7755±0.0418 | 0.6877±0.0403 | 0.6286±0.0969 | 0.6544±0.0667 |
| Logistic Regression (LR) | 0.7118±0.0324 | 0.7760±0.0282 | 0.6525±0.0487 | 0.6171±0.0506 | 0.6323±0.0367 |
| Linear Discriminant Analysis (LDA) | 0.7072±0.0393 | 0.7567±0.0365 | 0.6457±0.0545 | 0.6114±0.0388 | 0.6272±0.0412 |
| Multi-Layer Perceptron (MLP) | 0.6717±0.0302 | 0.7071±0.0593 | 0.6133±0.0423 | 0.4914±0.0984 | 0.5414±0.0684 |
| Gaussian Naïve Bayes (Gaussian NB) | 0.5293±0.0169 | 0.6980±0.0274 | 0.4571±0.0096 | 0.9086±0.0194 | 0.6081±0.0097 |
| K-Nearest Neighbors (KNN) | 0.6223±0.0475 | 0.6487±0.0445 | 0.5385±0.0762 | 0.4286±0.0619 | 0.4786±0.0661 |
| Decision Tree (DT) | 0.6464±0.0252 | 0.6363±0.0317 | 0.5567±0.0295 | 0.5771±0.0780 | 0.5651±0.0502 |
| Model/Classifier | Accuracy | ROC-AUC | Precision | Recall | F1-Score |
|---|---|---|---|---|---|
| Random Forest (RF) | 0.7248 | 0.7677 | 0.675 | 0.6136 | 0.6429 |
| XGBoost (XGB) | 0.6881 | 0.7574 | 0.6282 | 0.5568 | 0.5904 |
| Model/Classifier | Parameter Adjustment |
| Random Forest (RF) | maxDepth: 20; minSamplesSplit: 10; numTress: 100; bootstrap: False; seed: 42 |
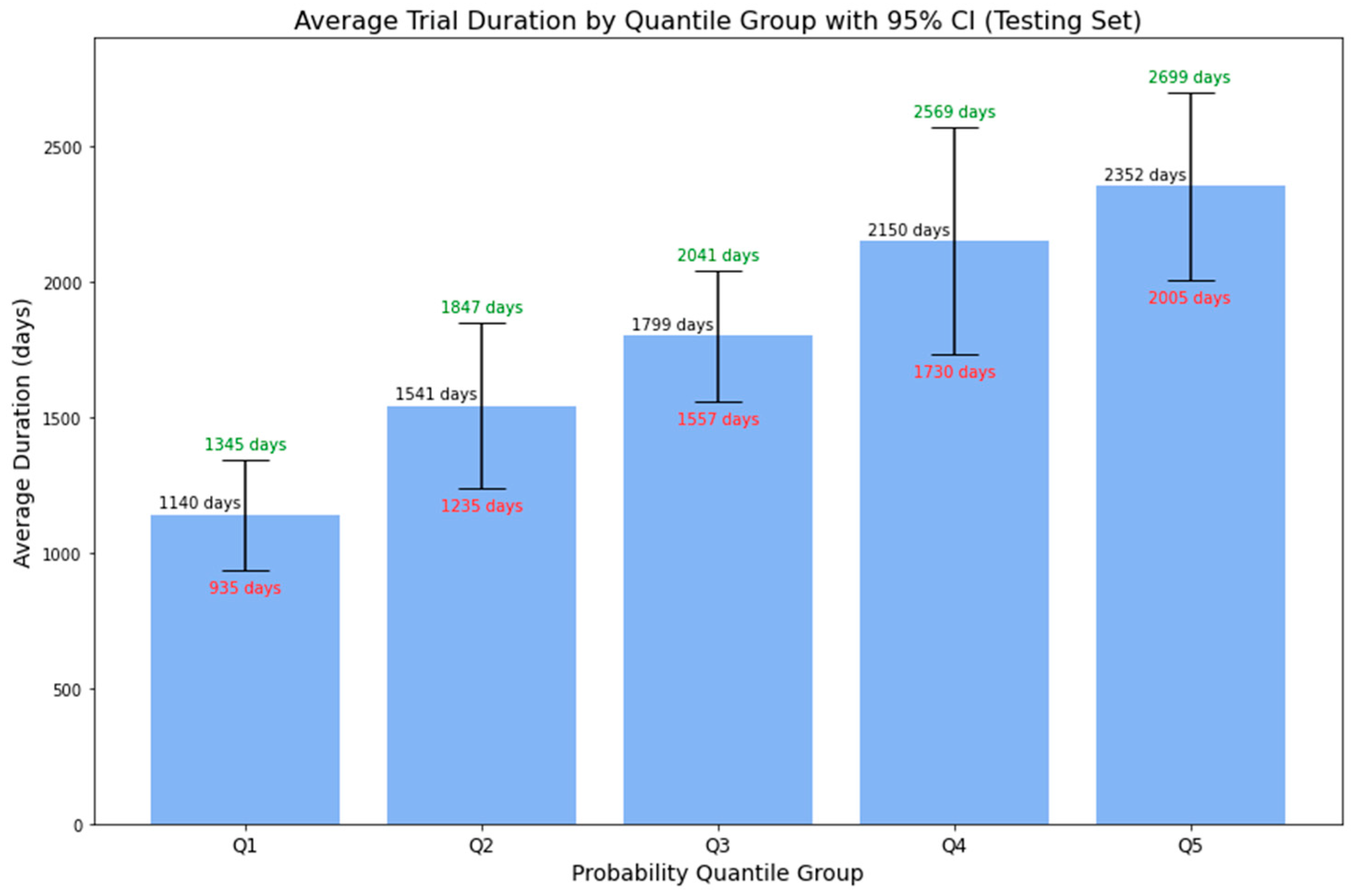
| Probability Quantile Group | Probability Range | Average Duration | Lower Bound (95% CI) | Upper Bound (95% CI) |
|---|---|---|---|---|
| Q1 | 0 to 0.1624 | 1140 days | 935 days | 1345 days |
| Q2 | 0.1624 to 0.3039 | 1541 days | 1235 days | 1847 days |
| Q3 | 0.3039 to 0.4697 | 1799 days | 1557 days | 2041 days |
| Q4 | 0.4697 to 0.6291 | 2150 days | 1730 days | 2569 days |
| Q5 | 0.6291 to 1 | 2352 days | 2005 days | 2699 days |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




