Submitted:
20 October 2023
Posted:
23 October 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
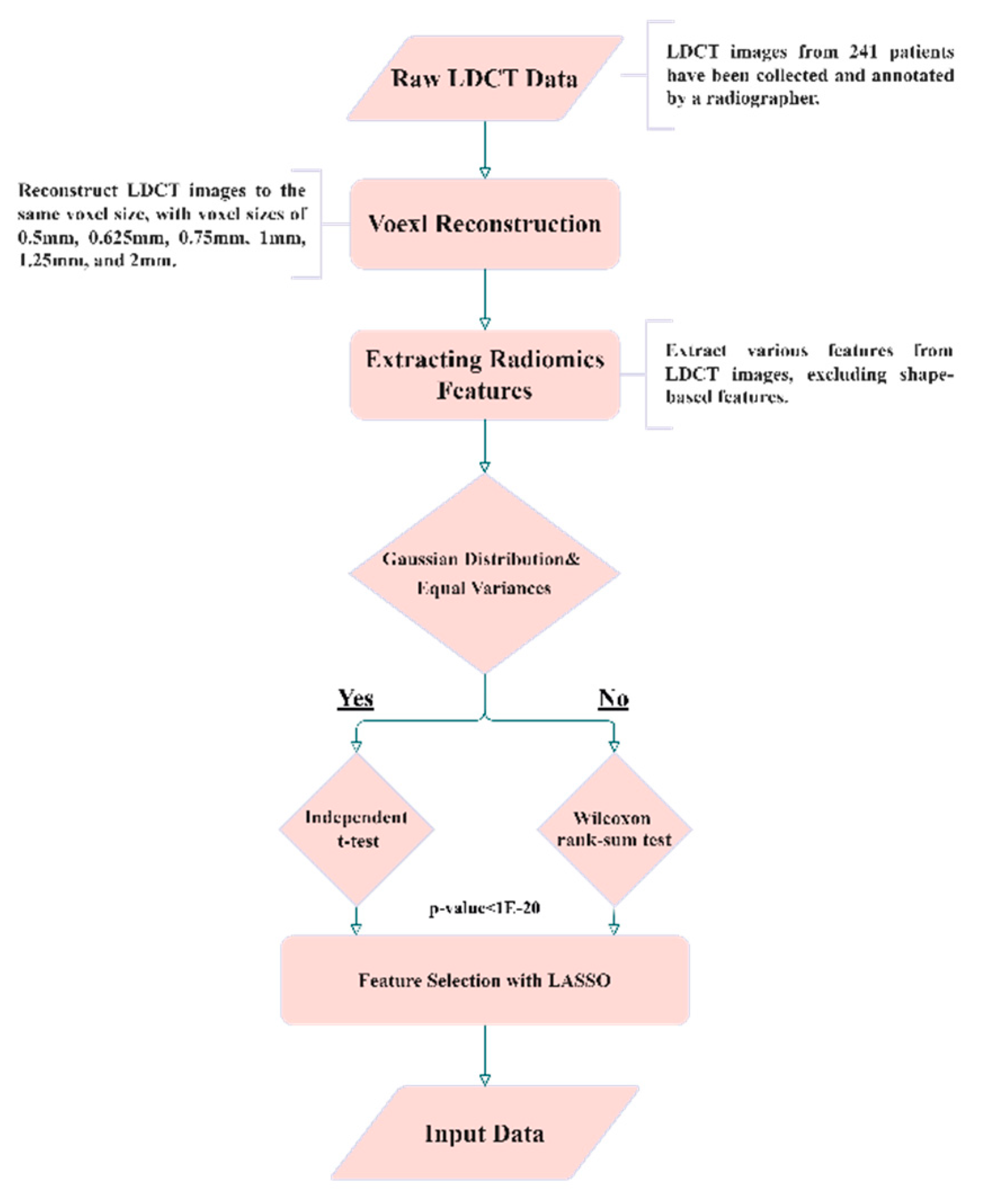
Dataset in House and Annotation
Isotropic Voxel Normalization and Image Reconstruction
Radiomics and Feature Selection
Support Vector Machine (SVM) and Hyperparameter Optimization
K-Fold Cross-Validation and Model Performance Evaluation
3. Results
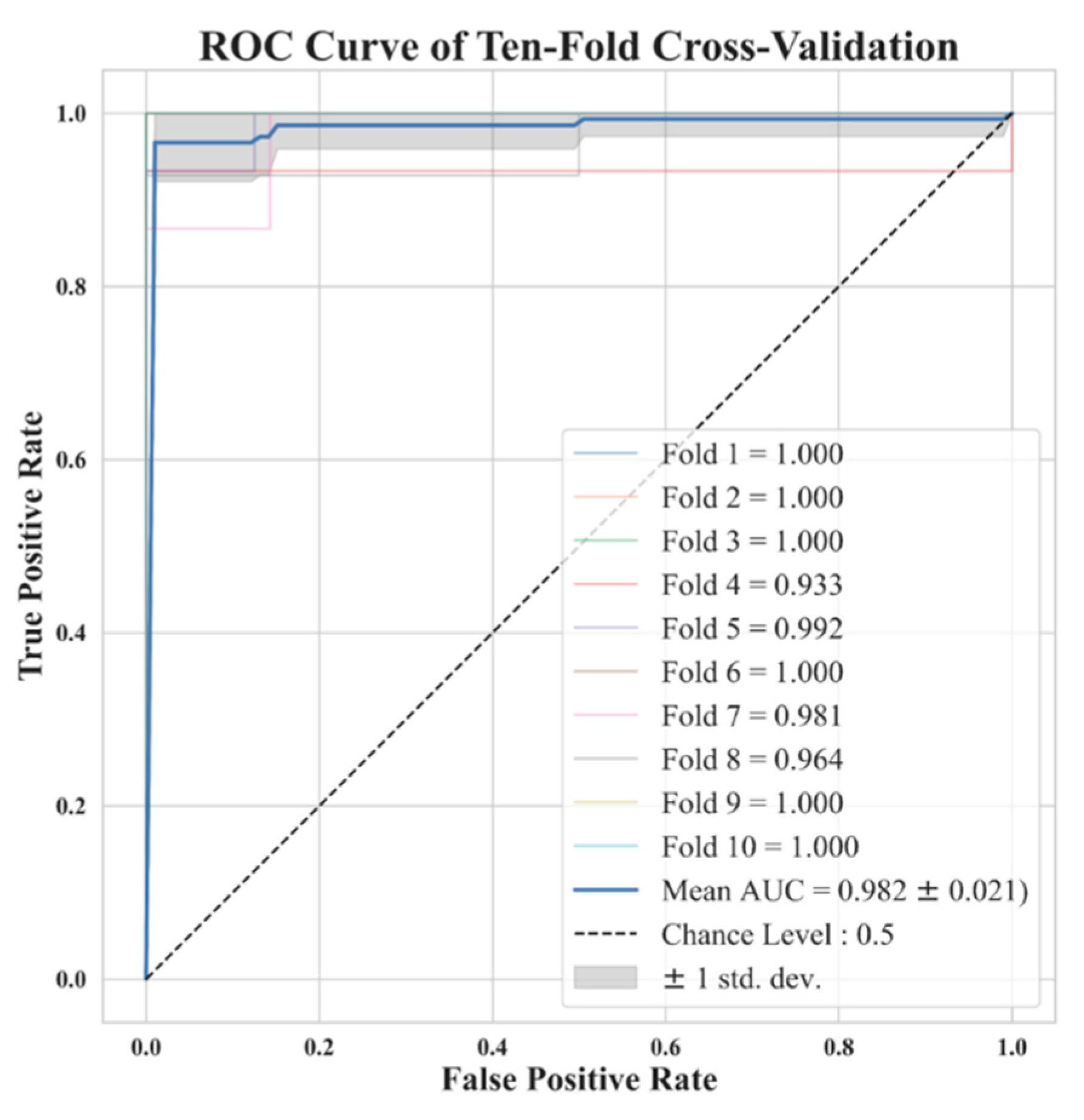
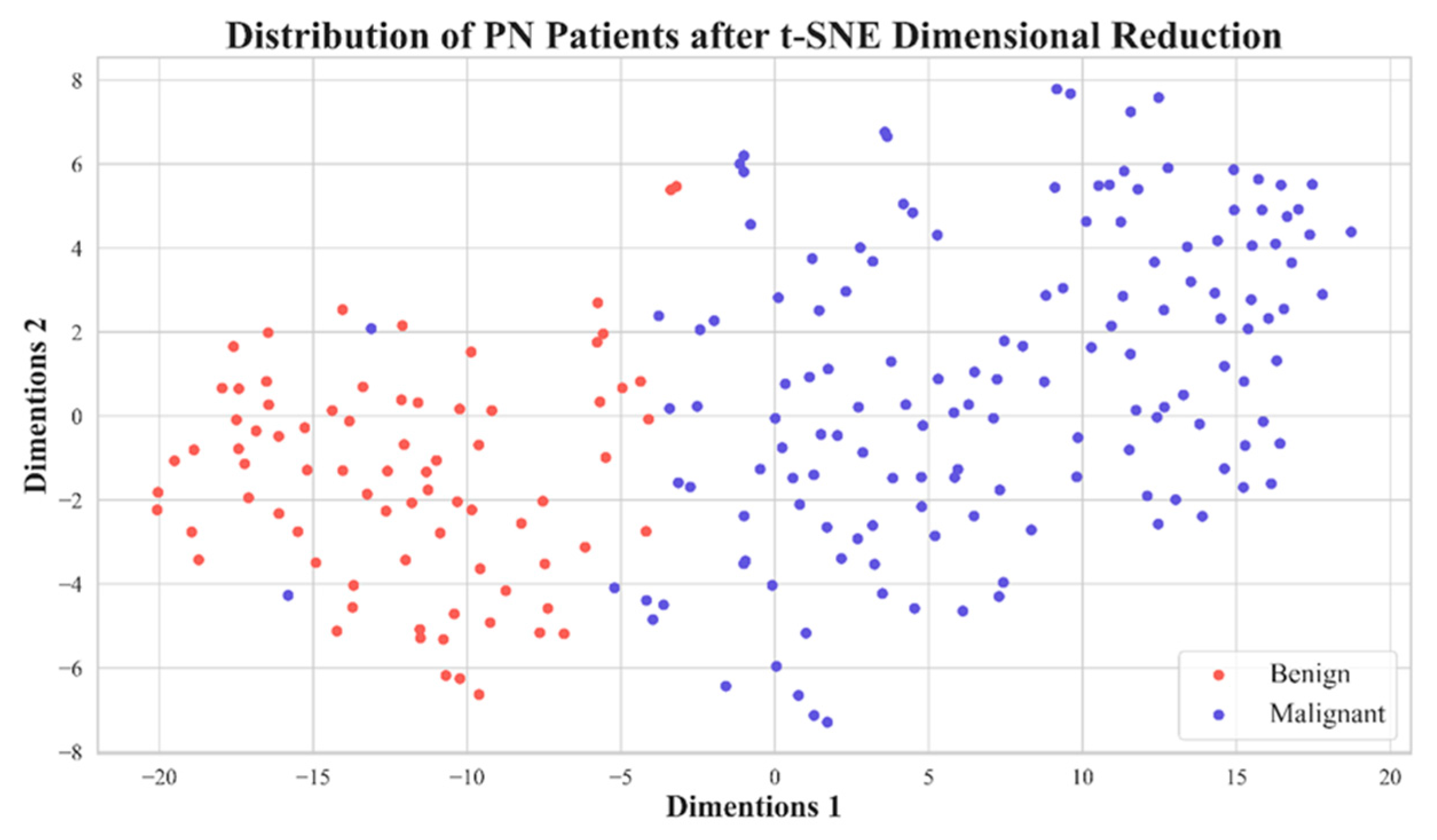
4. Discussions
5. Conclusion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
Appendix A
| Feature Description | Types |
|---|---|
| original_shape_Elongation | Shape-Based |
| original_shape_Flatness | Shape-Based |
| original_shape_LeastAxisLength | Shape-Based |
| original_shape_MajorAxisLength | Shape-Based |
| original_shape_Maximum2DDiameterColumn | Shape-Based |
| original_shape_Maximum2DDiameterRow | Shape-Based |
| original_shape_Maximum2DDiameterSlice | Shape-Based |
| original_shape_Maximum3DDiameter | Shape-Based |
| original_shape_MeshVolume | Shape-Based |
| original_shape_MinorAxisLength | Shape-Based |
| original_shape_Sphericity | Shape-Based |
| original_shape_SurfaceArea | Shape-Based |
| original_shape_SurfaceVolumeRatio | Shape-Based |
| original_shape_VoxelVolume | Shape-Based |
References
- Siegel, R.L.; Miller, K.D.; Wagle, N.S.; Jemal, A. Cancer statistics, 2023. Ca Cancer J Clin 2023, 73, 17–48. [Google Scholar] [CrossRef]
- Herbst, R.S.; Morgensztern, D.; Boshoff, C. The biology and management of non-small cell lung cancer. Nature 2018, 553, 446–454. [Google Scholar] [CrossRef] [PubMed]
- Knight, S.B.; Crosbie, P.; Balata, H.; Chudziak, J.; Hussell, T.; Dive, C. Progress and prospects of early detection in lung cancer. Open Biol. 2017.
- Jemal, A.; Fedewa, S.A. Lung cancer screening with low-dose computed tomography in the United States—2010 to 2015. JAMA oncology 2017, 3, 1278–1281. [Google Scholar] [CrossRef]
- Armato III, S.G.; McLennan, G.; Bidaut, L.; McNitt-Gray, M.F.; Meyer, C.R.; Reeves, A.P.; Zhao, B.; Aberle, D.R.; Henschke, C.I.; Hoffman, E.A. The lung image database consortium (LIDC) and image database resource initiative (IDRI): a completed reference database of lung nodules on CT scans. Medical physics 2011, 38, 915–931. [Google Scholar] [CrossRef]
- Cha, M.J.; Lee, K.S.; Kim, H.S.; Lee, S.W.; Jeong, C.J.; Kim, E.Y.; Lee, H.Y. Improvement in imaging diagnosis technique and modalities for solitary pulmonary nodules: from ground-glass opacity nodules to part-solid and solid nodules. Expert Review of Respiratory Medicine 2016, 10, 261–278. [Google Scholar] [CrossRef]
- Swensen, S.J.; Viggiano, R.W.; Midthun, D.E.; Müller, N.L.; Sherrick, A.; Yamashita, K.; Naidich, D.P.; Patz, E.F.; Hartman, T.E.; Muhm, J.R. Lung nodule enhancement at CT: multicenter study. Radiology 2000, 214, 73–80. [Google Scholar] [CrossRef]
- Chelala, L.; Hossain, R.; Kazerooni, E.A.; Christensen, J.D.; Dyer, D.S.; White, C.S. Lung-RADS version 1.1: challenges and a look ahead, from the AJR special series on radiology reporting and data systems. American Journal of Roentgenology 2021, 216, 1411–1422. [Google Scholar] [CrossRef]
- Swensen, S.J.; Silverstein, M.D.; Edell, E.S.; Trastek, V.F.; Aughenbaugh, G.L.; Ilstrup, D.M.; Schleck, C.D. Solitary pulmonary nodules: clinical prediction model versus physicians. Proceedings of Mayo Clinic Proceedings; pp. 319–329.
- Khan, T.; Usman, Y.; Abdo, T.; Chaudry, F.; Keddissi, J.I.; Youness, H.A. Diagnosis and management of peripheral lung nodule. Annals of Translational Medicine 2019, 7. [Google Scholar] [CrossRef]
- Gillies, R.J.; Kinahan, P.E.; Hricak, H. Radiomics: images are more than pictures, they are data. Radiology 2016, 278, 563–577. [Google Scholar] [CrossRef]
- Doi, K. Current status and future potential of computer-aided diagnosis in medical imaging. The British journal of radiology 2005, 78, s3–s19. [Google Scholar] [CrossRef] [PubMed]
- Dey, R.; Lu, Z.; Hong, Y. Diagnostic classification of lung nodules using 3D neural networks. Proceedings of 2018 IEEE 15th international symposium on biomedical imaging (ISBI 2018); pp. 774–778.
- Huang, H.; Wu, R.; Li, Y.; Peng, C. Self-supervised transfer learning based on domain adaptation for benign-malignant lung nodule classification on thoracic CT. IEEE Journal of Biomedical and Health Informatics 2022, 26, 3860–3871. [Google Scholar] [CrossRef]
- Kang, G.; Liu, K.; Hou, B.; Zhang, N. 3D multi-view convolutional neural networks for lung nodule classification. PloS one 2017, 12, e0188290. [Google Scholar] [CrossRef]
- Kumar, D.; Wong, A.; Clausi, D.A. Lung nodule classification using deep features in CT images. In Proceedings of the 2015 12th conference on computer and robot vision; 2015; pp. 133–138. [Google Scholar]
- Liu, K.; Kang, G. Multiview convolutional neural networks for lung nodule classification. International Journal of Imaging Systems and Technology 2017, 27, 12–22. [Google Scholar] [CrossRef]
- Lu, X.; Nanehkaran, Y.A.; Karimi Fard, M. A method for optimal detection of lung cancer based on deep learning optimized by marine predators algorithm. Computational Intelligence and Neuroscience 2021, 2021. [Google Scholar] [CrossRef]
- Mehta, K.; Jain, A.; Mangalagiri, J.; Menon, S.; Nguyen, P.; Chapman, D.R. Lung nodule classification using biomarkers, volumetric radiomics, and 3D CNNs. Journal of Digital Imaging 2021, 1–20. [Google Scholar] [CrossRef]
- Saihood, A.; Karshenas, H.; Nilchi, A.R.N. Deep fusion of gray level co-occurrence matrices for lung nodule classification. Plos one 2022, 17, e0274516. [Google Scholar] [CrossRef]
- Shen, W.; Zhou, M.; Yang, F.; Yang, C.; Tian, J. Multi-scale convolutional neural networks for lung nodule classification. Proceedings of Information Processing in Medical Imaging: 24th International Conference, IPMI 2015, Sabhal Mor Ostaig, Isle of Skye, UK, 2015, Proceedings 24, June 28-July 3; pp. 588–599.
- Shen, W.; Zhou, M.; Yang, F.; Yu, D.; Dong, D.; Yang, C.; Zang, Y.; Tian, J. Multi-crop convolutional neural networks for lung nodule malignancy suspiciousness classification. Pattern Recognition 2017, 61, 663–673. [Google Scholar] [CrossRef]
- Tomassini, S.; Falcionelli, N.; Sernani, P.; Burattini, L.; Dragoni, A.F. Lung nodule diagnosis and cancer histology classification from computed tomography data by convolutional neural networks: A survey. Computers in Biology and Medicine 2022, 146, 105691. [Google Scholar] [CrossRef] [PubMed]
- Halder, A.; Chatterjee, S.; Dey, D. Adaptive morphology aided 2-pathway convolutional neural network for lung nodule classification. Biomedical Signal Processing and Control 2022, 72, 103347. [Google Scholar] [CrossRef]
- Kim, H.; Goo, J.M.; Ohno, Y.; Kauczor, H.-U.; Hoffman, E.A.; Gee, J.C.; Van Beek, E.J. Effect of reconstruction parameters on the quantitative analysis of chest computed tomography. Journal of thoracic imaging 2019, 34, 92–102. [Google Scholar] [CrossRef]
- Lu, Y.; Fontaine, K.; Germino, M.; Mulnix, T.; Casey, M.E.; Carson, R.E.; Liu, C. Investigation of sub-centimeter lung nodule quantification for low-dose PET. IEEE Transactions on Radiation and Plasma Medical Sciences 2017, 2, 41–50. [Google Scholar] [CrossRef]
- Tonekaboni, S.; Joshi, S.; McCradden, M.D.; Goldenberg, A. What clinicians want: contextualizing explainable machine learning for clinical end use. In Proceedings of the Machine learning for healthcare conference; 2019; pp. 359–380. [Google Scholar]
- Nioche, C.; Orlhac, F.; Boughdad, S.; Reuzé, S.; Goya-Outi, J.; Robert, C.; Pellot-Barakat, C.; Soussan, M.; Frouin, F.; Buvat, I. LIFEx: a freeware for radiomic feature calculation in multimodality imaging to accelerate advances in the characterization of tumor heterogeneity. Cancer research 2018, 78, 4786–4789. [Google Scholar] [CrossRef]
- Heeren, T.; D'Agostino, R. Robustness of the two independent samples t-test when applied to ordinal scaled data. Statistics in medicine 1987, 6, 79–90. [Google Scholar] [CrossRef]
- Rosner, B.; Glynn, R.J.; Ting Lee, M.L. Incorporation of clustering effects for the Wilcoxon rank sum test: a large-sample approach. Biometrics 2003, 59, 1089–1098. [Google Scholar] [CrossRef]
- Das, K.R.; Imon, A. A brief review of tests for normality. American Journal of Theoretical and Applied Statistics 2016, 5, 5–12. [Google Scholar]
- Schultz, B.B. Levene's test for relative variation. Systematic Zoology 1985, 34, 449–456. [Google Scholar] [CrossRef]
- Fonti, V.; Belitser, E. Feature selection using lasso. VU Amsterdam research paper in business analytics 2017, 30, 1–25. [Google Scholar]
- Van der Maaten, L.; Hinton, G. Visualizing data using t-SNE. Journal of machine learning research 2008, 9. [Google Scholar]
- Suthaharan, S.; Suthaharan, S. Support vector machine. Machine learning models and algorithms for big data classification: thinking with examples for effective learning 2016, 207-235.
- Cervantes, J.; Garcia-Lamont, F.; Rodríguez-Mazahua, L.; Lopez, A. A comprehensive survey on support vector machine classification: Applications, challenges and trends. Neurocomputing 2020, 408, 189–215. [Google Scholar] [CrossRef]
- Frazier, P.I. A tutorial on Bayesian optimization. arXiv 2018, arXiv:1807.02811. [Google Scholar]
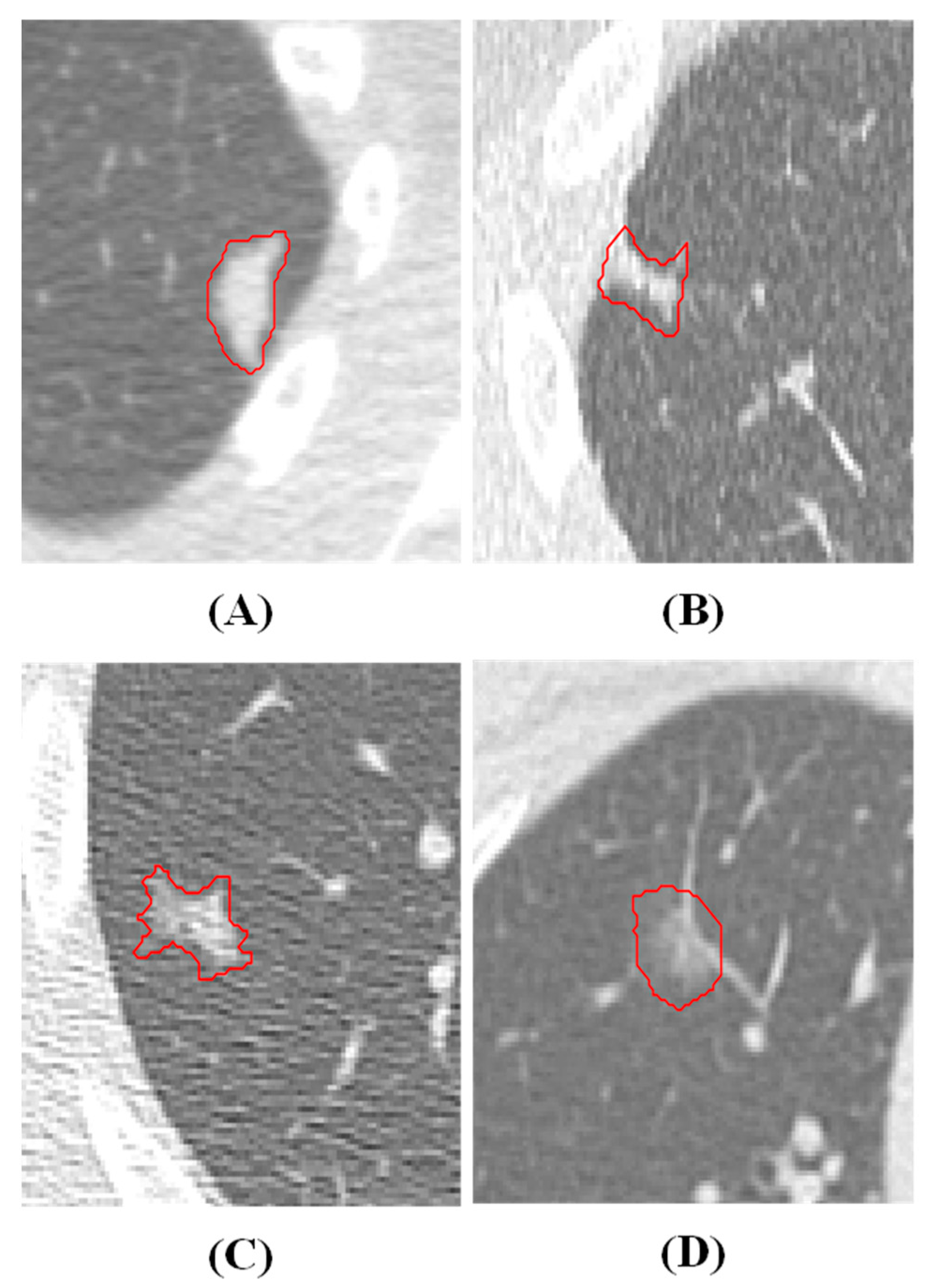
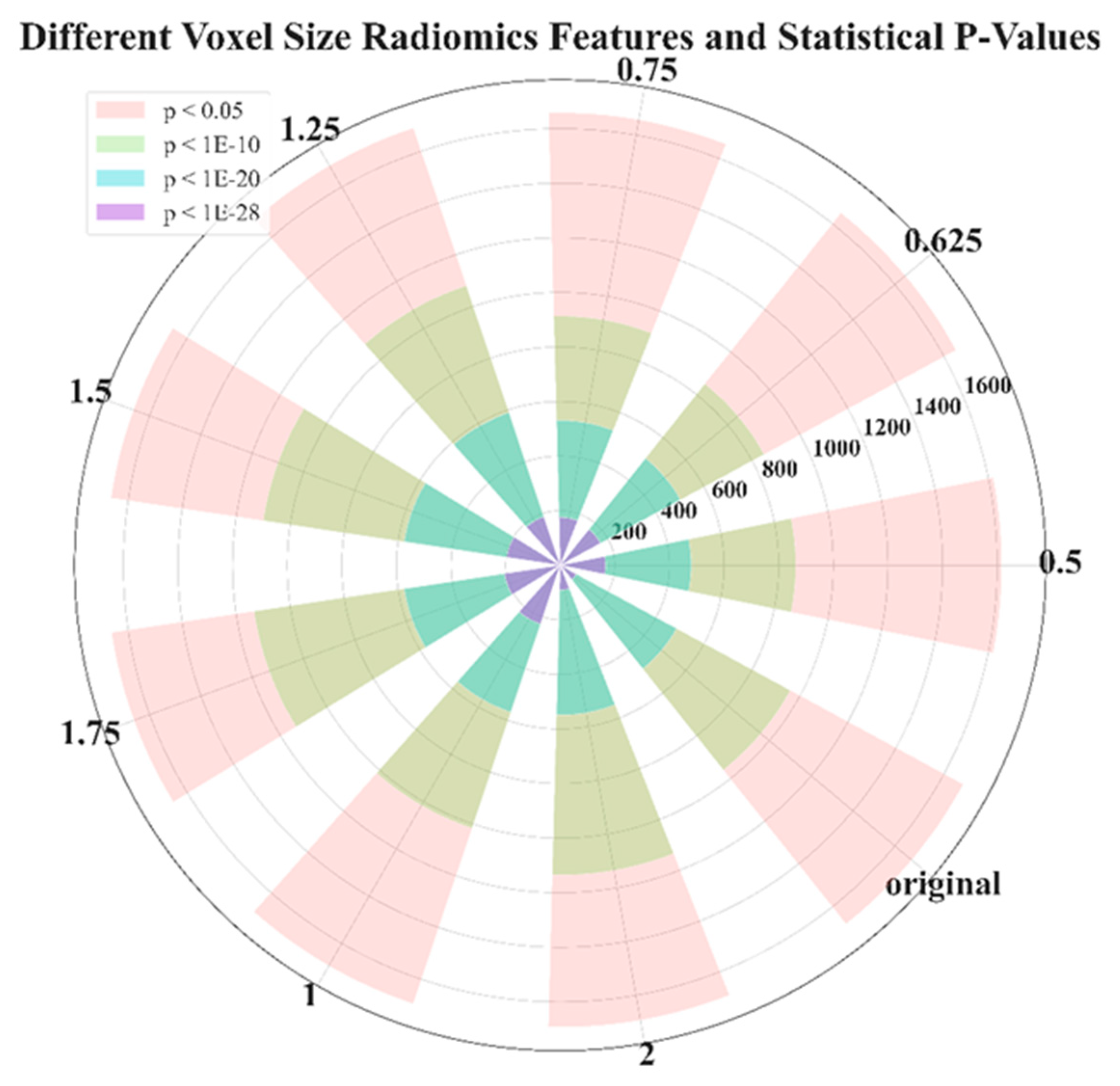
| VOXEL SIZE | 0.5 | 0.625 | 0.75 | 1 | 1.25 | 1.5 | 1.75 | 2 | ORIGINAL |
|---|---|---|---|---|---|---|---|---|---|
| P<0.05 | 1617 | 1650 | 1657 | 1694 | 1690 | 1663 | 1661 | 1692 | 1680 |
| P<1E-10 | 863 | 850 | 913 | 1016 | 1081 | 1100 | 1134 | 1135 | 959 |
| P<1E-20 | 480 | 501 | 531 | 568 | 590 | 578 | 578 | 549 | 485 |
| P<1E-28 | 166 | 168 | 175 | 227 | 187 | 198 | 206 | 91 | 67 |
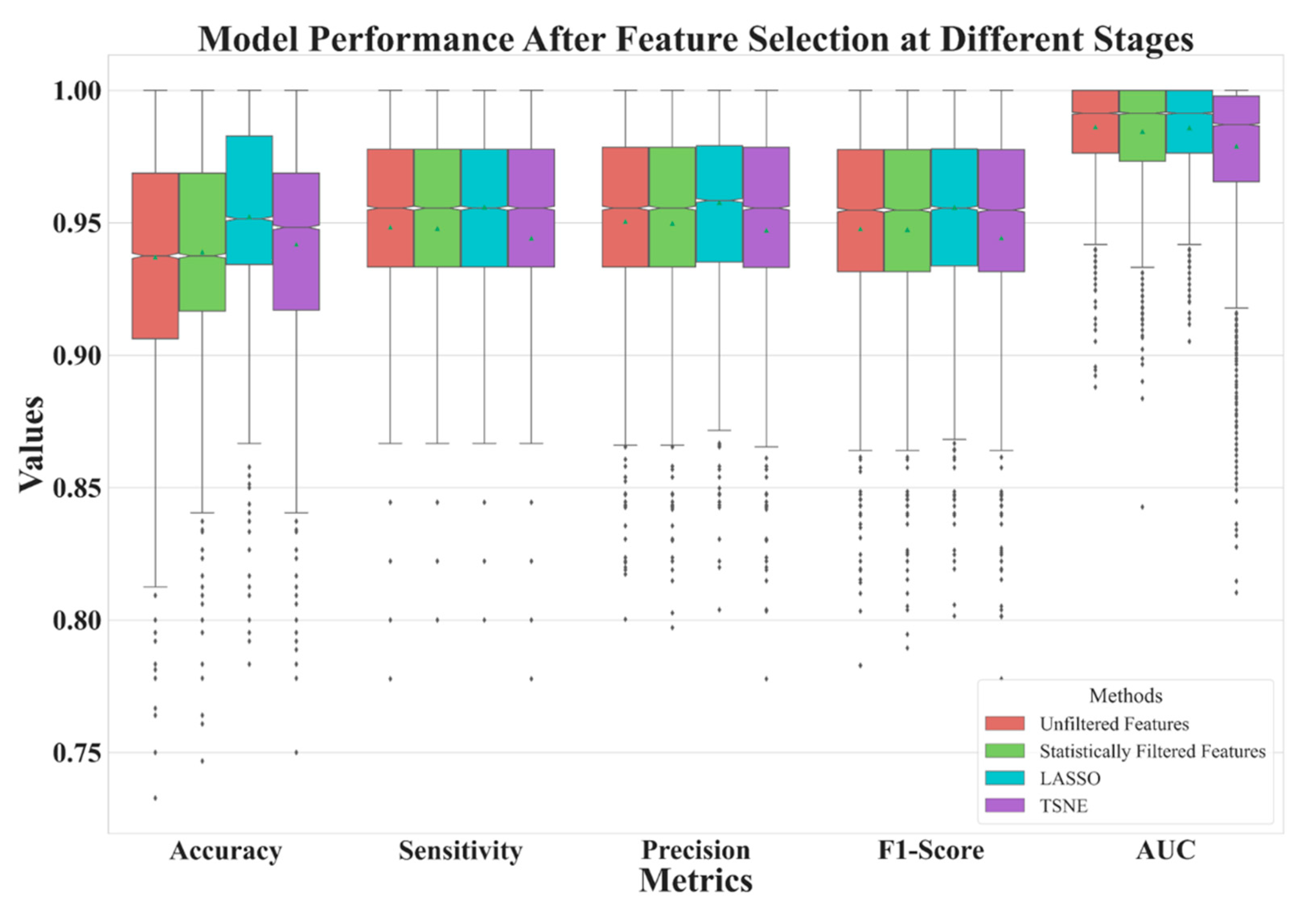
| Unfiltered Features | Statistically Filtered Features | LASSO | t-SNE | |
|---|---|---|---|---|
| Feature Number | 2061 | 480 | 11 | 2 |
| Accuracy | AUC | Sensitivity | Precision | F1 Score | |
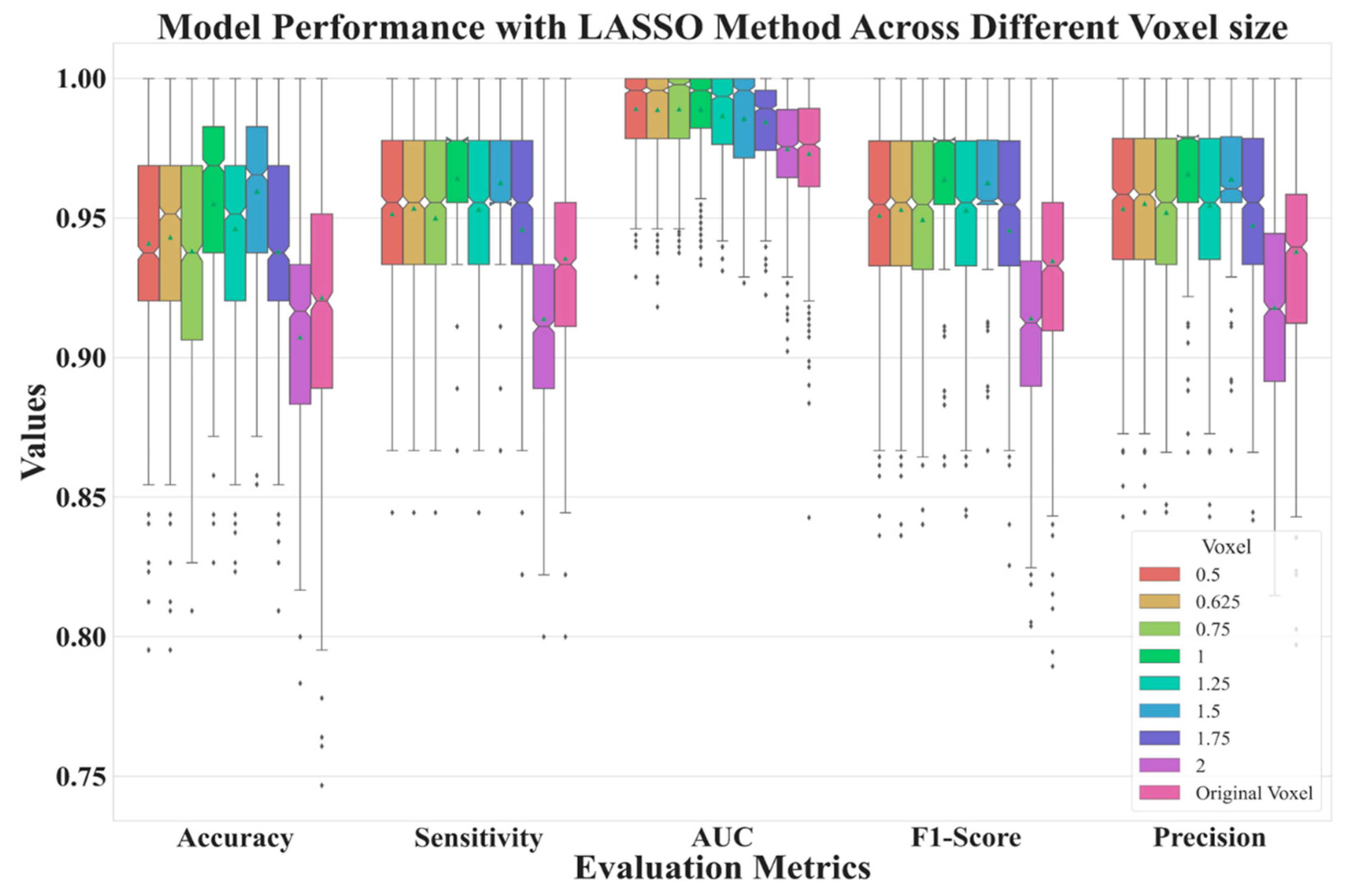
|---|---|---|---|---|---|
| 0.5 | 0.9409 | 0.9891 | 0.9514 | 0.9533 | 0.9509 |
| 0.625 | 0.9431 | 0.9887 | 0.9535 | 0.9551 | 0.9530 |
| 0.75 | 0.9350 | 0.9890 | 0.9481 | 0.9501 | 0.9473 |
| 1 | 0.9531 | 0.9890 | 0.9624 | 0.9640 | 0.9620 |
| 1.25 | 0.9467 | 0.9866 | 0.9532 | 0.9548 | 0.9530 |
| 1.5 | 0.9596 | 0.9855 | 0.9619 | 0.9633 | 0.9619 |
| 1.75 | 0.9371 | 0.9844 | 0.9452 | 0.9468 | 0.9449 |
| 2 | 0.9073 | 0.9747 | 0.9156 | 0.9197 | 0.9159 |
| Original | 0.9223 | 0.9731 | 0.9357 | 0.9381 | 0.9349 |
| Halder et al.2021[24] | 0.9610 | 0.9936 | 0.9685 | ||
| Mehta et al.2021[19] | 0.8659 | ||||
| Shen et al.2017[22] | 0.8612 | ||||
| Lu et al. 2021[18] | 0.934 | 0.984 |
| Feature Description | Types |
|---|---|
| original_gldm_SmallDependenceLowGrayLevelEmphasis | Texture |
| log-sigma-2-0-mm-3D_glcm_DifferenceEntropy | Texture |
| log-sigma-2-0-mm-3D_gldm_SmallDependenceEmphasis | Texture |
| log-sigma-3-0-mm-3D_glszm_ZonePercentage | Texture |
| lbp-2D_gldm_DependenceNonUniformityNormalized | Texture |
| lbp-3D-m1_gldm_DependenceNonUniformityNormalized | Texture |
| lbp-3D-m2_gldm_DependenceNonUniformityNormalized | Texture |
| log-sigma-2-0-mm-3D_firstorder_Mean | First order |
| lbp-3D-m1_firstorder_Skewness lbp-3D- | First order |
| wavelet-LLH_firstorder_Mean | First order |
| wavelet-LHL_firstorder_Mean | First order |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




