Submitted:
16 August 2023
Posted:
17 August 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Related Work
2.1. Traditional Experimental Method
2.1.1. Experimental Method
2.1.2. Computer Aided Method
2.2. Machine Learning Method
2.2.1. Traditional Machine Learning Methods
2.2.2. Deep Learning Method
2.2.3. Integrated Learning Method
3. Data Preprocessing
3.1 Descriptive Statistics
3.2 Elimination of Low Variance Characteristics
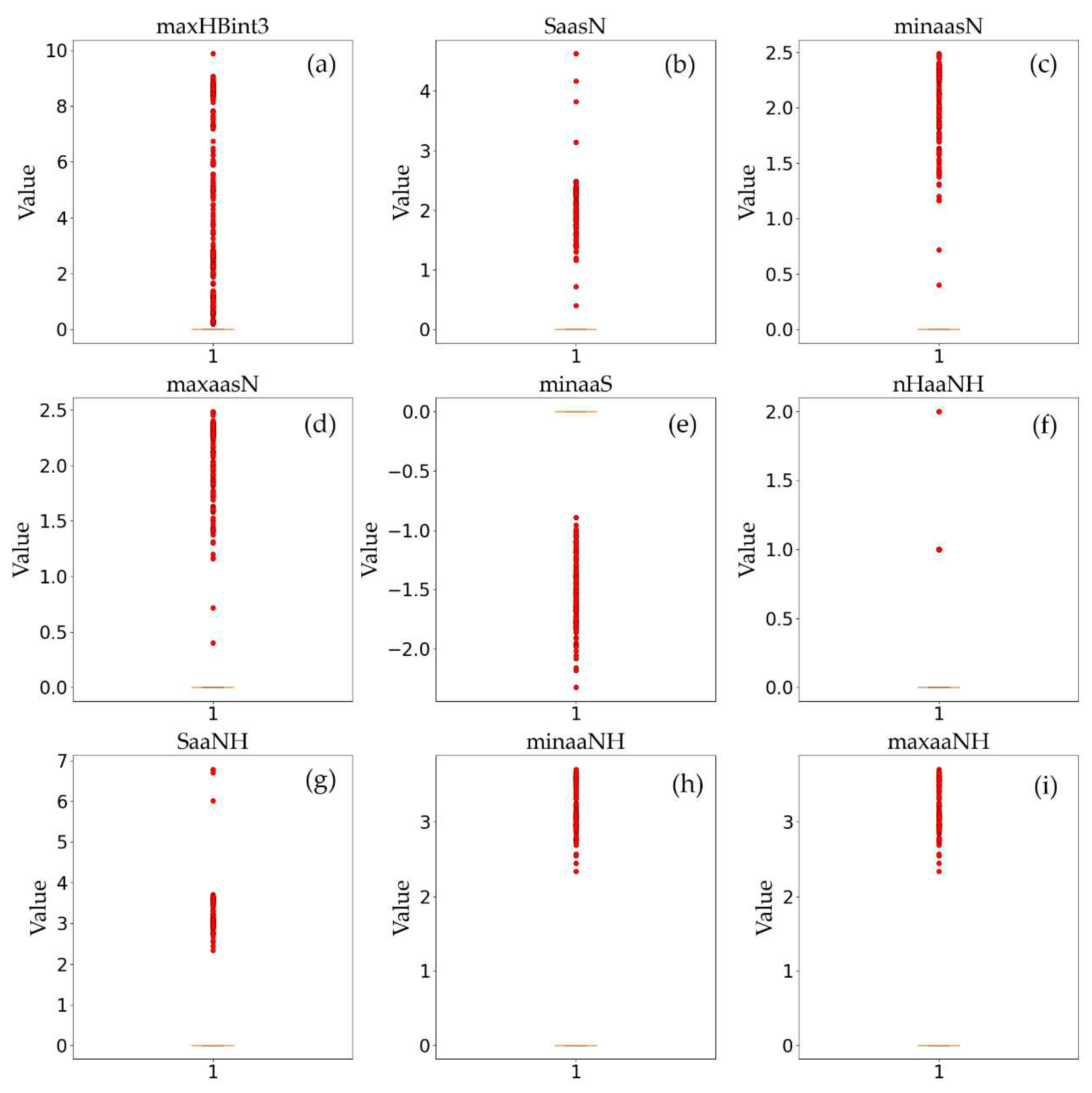
3.3 Diagnosis of Abnormal Variables
4. Methodology
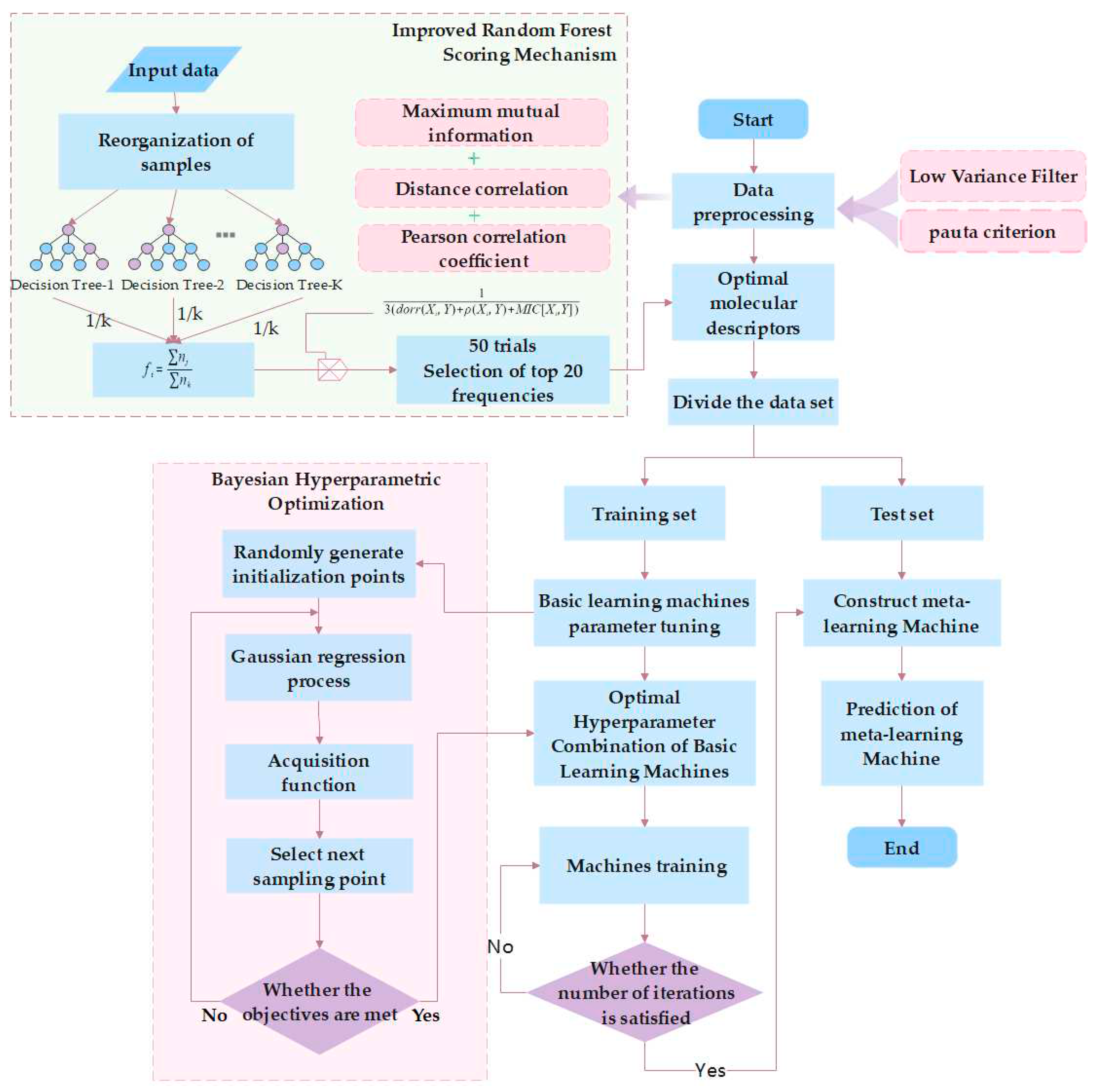
4.1 Improved Random Forest Feature Selection Method.
4.1.1. The Original Random Forest Method
4.1.2. Maximum Mutual Information Coefficient
4.1.3. Pearson Correlation Coefficient
4.1.4. Distance Correlation Coefficient
4.1.5. Improved Random Forest Method
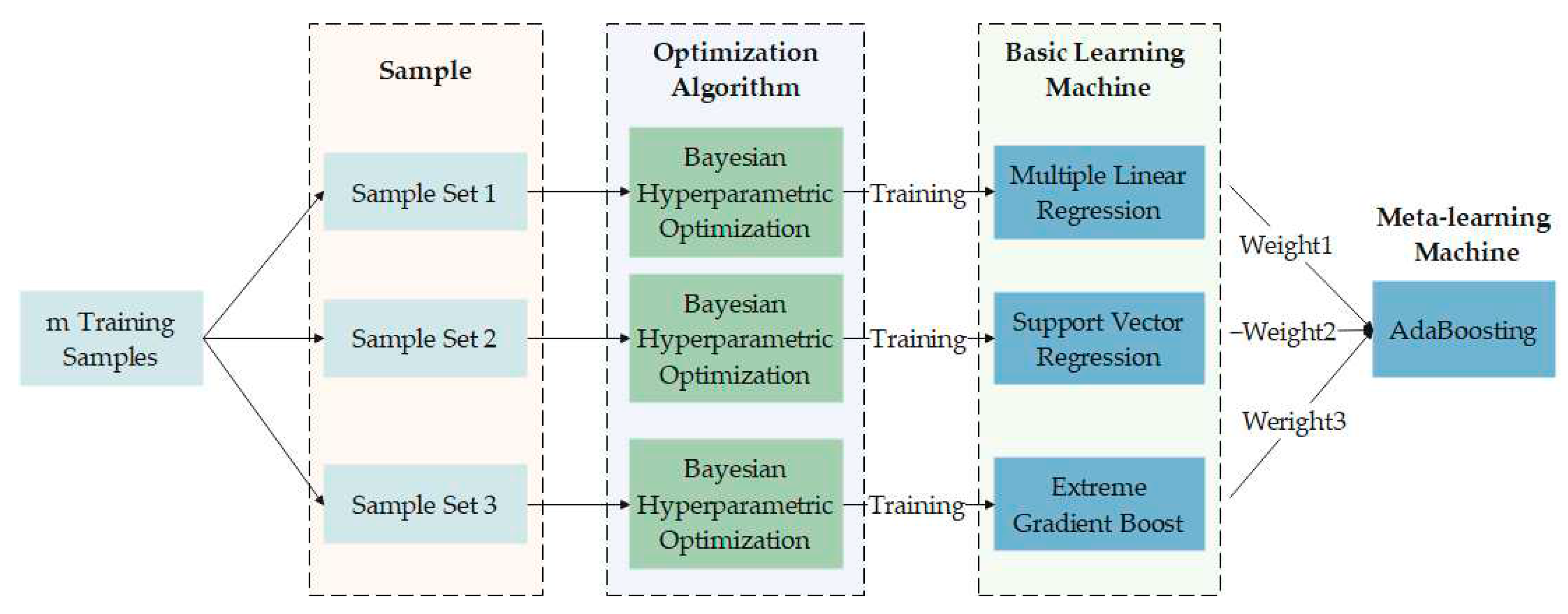
4.2. Establishment of the BHO-AdaBoosting Model
4.2.1. Extreme Gradient Boosting (XGBoost)
4.2.2. Multiple Linear Regression (MLR) Model
4.2.3. Support Vector Machine Regression (SVR)
4.2.4. Bayesian Hyperparameter Optimization
5. Experimental Results
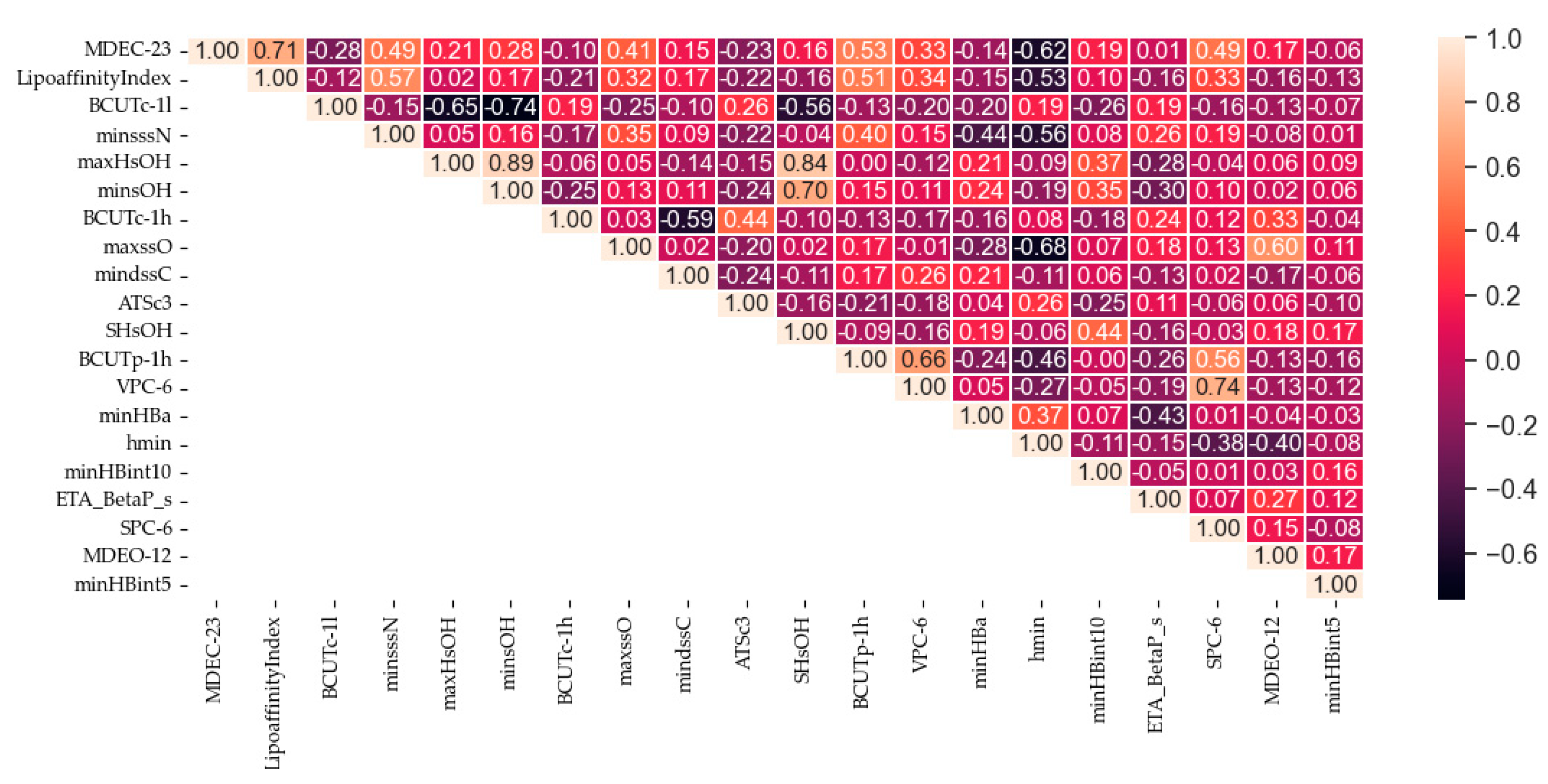
5.1. Results of the Improved Random Forest Feature Selection
5.2. Quantitative Prediction of ERα Biological Activity
5.2.1. Results of Bayesian Hyperparameter Optimization
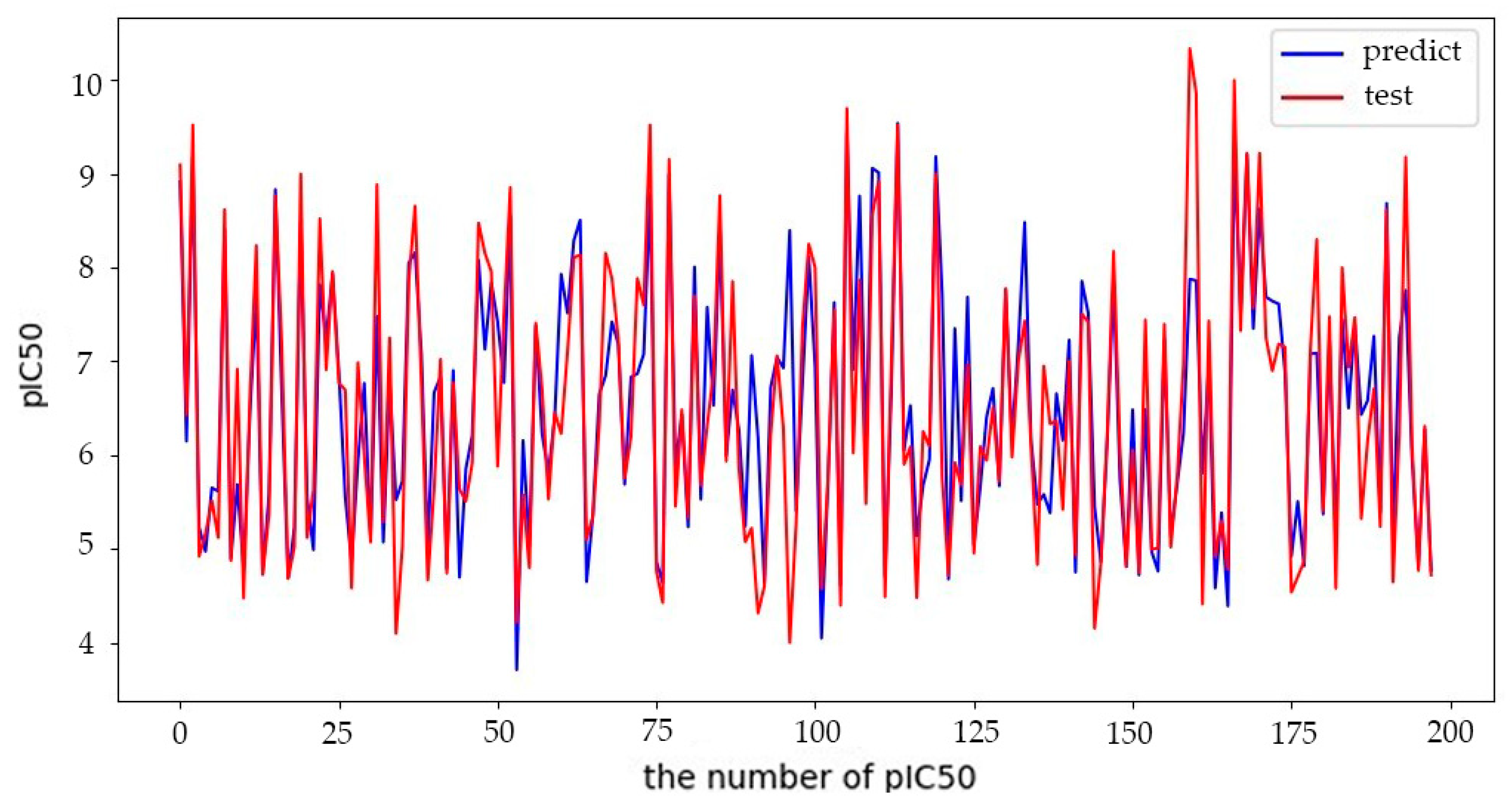
5.2.2. Quantitative Prediction Results of BHO-AdaBoosting
6. Discussion
6.1. Comparative Experimental Results and Error Analysis
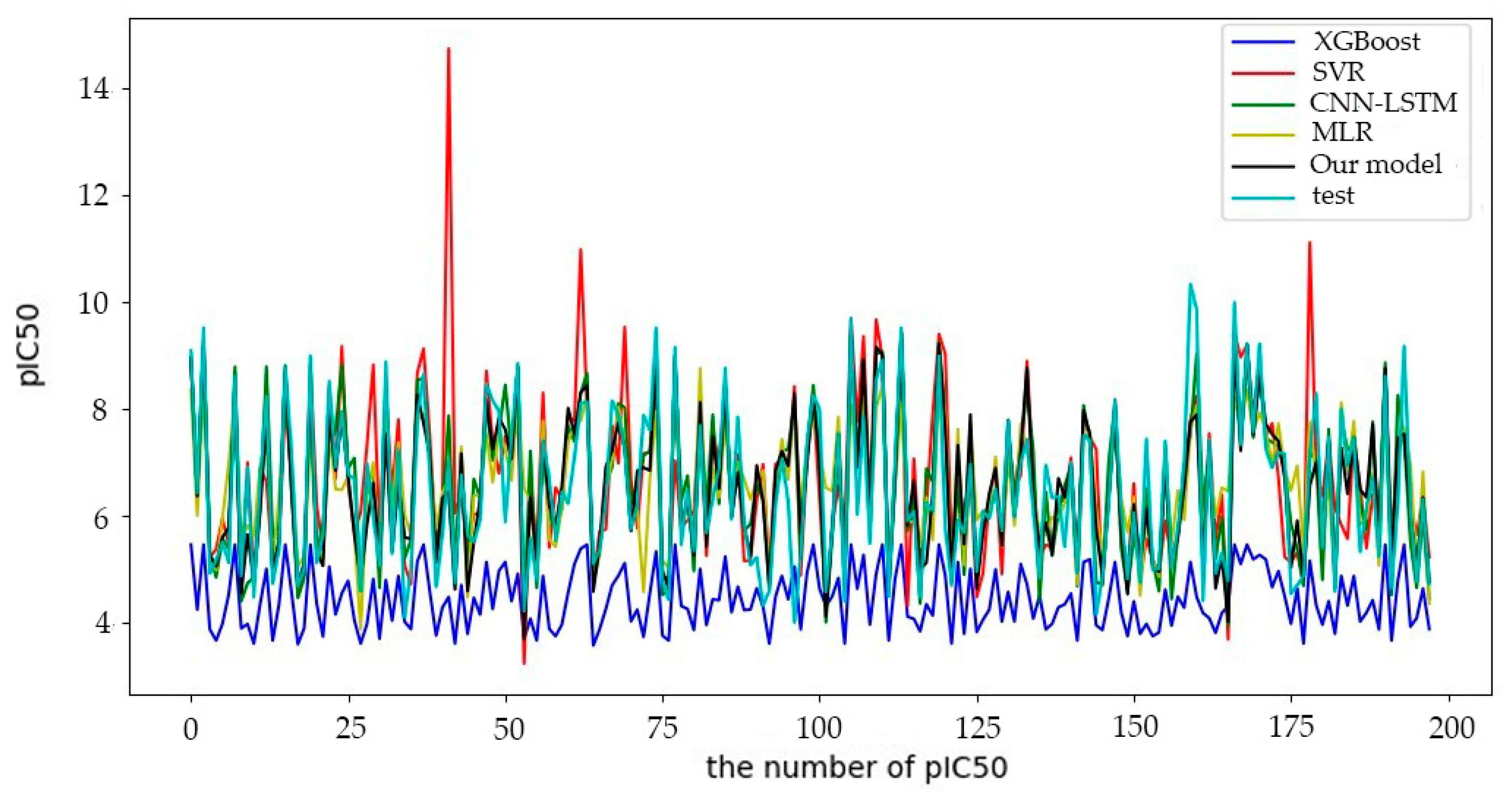
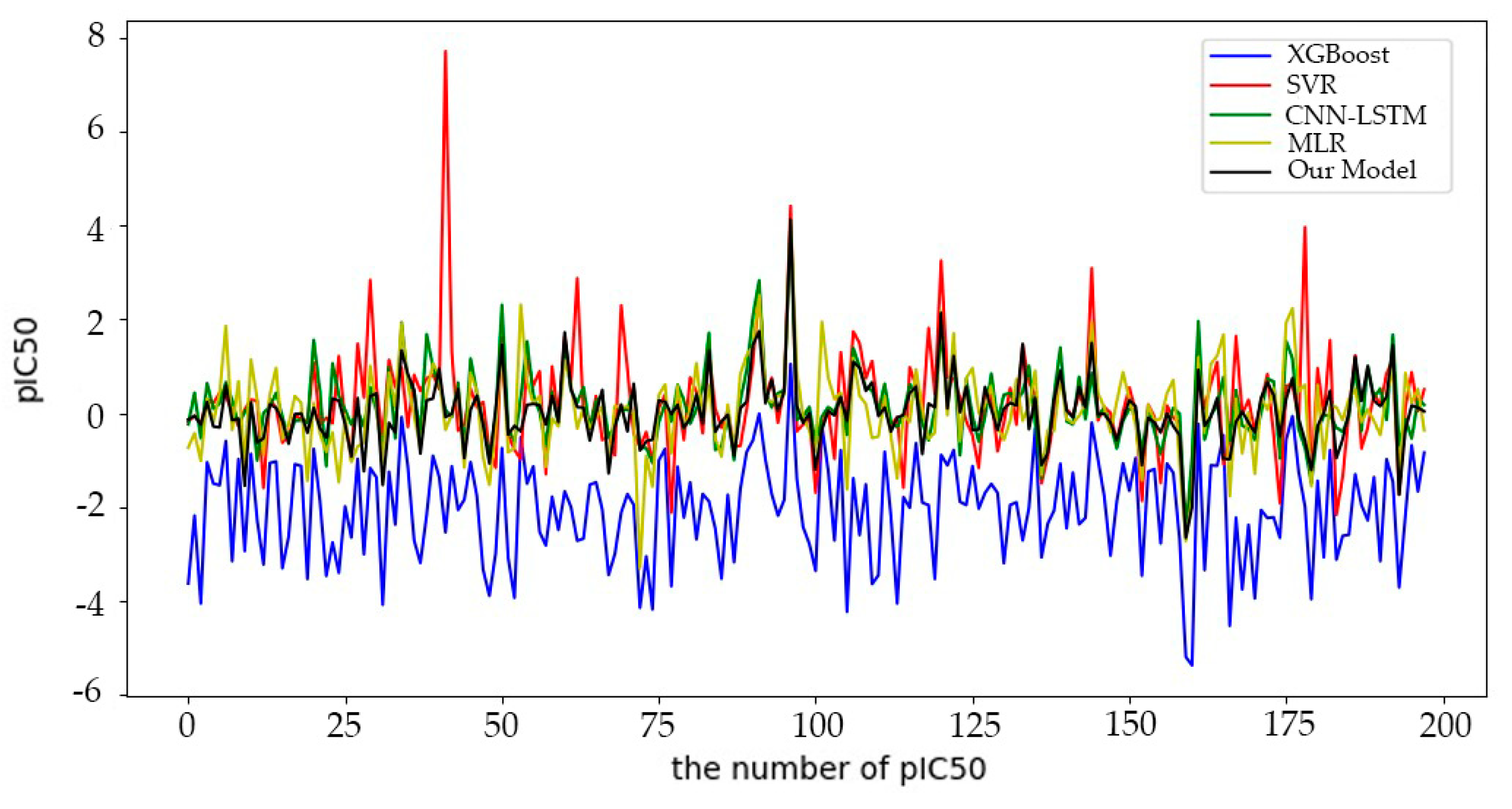
6.1.1. Comparative Experimental Results
6.2. Prediction for 50 Candidate Compounds
7. Conclusion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Siegel, R.L.; Miller, K.D.; Fuchs, H.E.; Jemal, A. Cancer Statistics 2021. CA Cancer J Clin. 2021, 71, 7–33. [Google Scholar] [CrossRef]
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef] [PubMed]
- Siegel, R.L.; Miller, K.D.; Fuchs, H.E.; Jemal, A. Cancer statistics 2022. CA Cancer J Clin. 2022, 72, 7–33. [Google Scholar] [CrossRef]
- Ferlay J, Ervik M, Lam F, Colombet M, Mery L, Piñeros M, Znaor A, Soerjomataram I, Bray F (2020). Global Cancer Observatory: Cancer Today. Lyon, France: International Agency for Research on Cancer. Available from: https://gco.iarc.fr/today, accessed []. 20 July.
- Hayashi N; Kumamaru H; Isozumi U; Aogi K; Asaga S; Iijima K; Kadoya T; Kojima Y; Kubo M; Miyashita M; Miyata H; Nagahashi M; Niikura N; Ogo E; Tamura K; Tanakura K; Yamamoto Y; Yoshida M; Imoto S; Jinno H. Annual report of the Japanese Breast Cancer Registry for 2017. Breast Cancer. 2020, 27, 803–809. [CrossRef]
- Lei, S.; Zheng, R.; Zhang, S.; Wang, S.; Chen, R.; Sun, K.; Zeng, H.; Zhou, J.; Wei, W. Global patterns of breast cancer incidence and mortality: A population-based cancer registry data analysis from 2000 to 2020. Cancer Commun (Lond). 2021, 41, 1183–1194. [Google Scholar] [CrossRef] [PubMed]
- Dickson, R.B.; Lippman, M.E. Control of human breast cancer by estrogen, growth factors, and oncogenes. Cancer Treat Res. 1988, 40, 119–65. [Google Scholar] [CrossRef]
- Jacquemier, J.D.; Hassoun, J.; Torrente, M.; Martin, P.M. Distribution of estrogen and progesterone receptors in healthy tissue adjacent to breast lesions at various stages--immunohistochemical study of 107 cases. Breast Cancer Res Treat. 1990, 15, 109–17. [Google Scholar] [CrossRef] [PubMed]
- Tekmal, R. R.; Liu, Y. G.; Nair, H. B.; Jones, J.; Perla, R. P.; Lubahn, D. B. ; Korach,K. S.; Kirma, N. Estrogen receptor alpha is required for mammary development and the induction of mammary hyperplasia and epigenetic alterations in the aromatase transgenic mice[J]. Journal of Steroid Biochemistry and Molecular Biology, 2005, 95, 9–15. [Google Scholar] [CrossRef]
- Lee, J.J.K.; Jung, Y.L.; Cheong, T.C.; Espejo Valle-Inclan, J.; Chu, C.; Gulhan, D. C.; Ljungström, V.; Jin, H.; Viswanadham, V.V.; Watson, E.V. ; Cortés-Ciriano,I. Nature 618. 2023, 1024–1032. [Google Scholar] [CrossRef]
- Chen, J.Q.; Russo, J. ERα-negative and triple negative breast cancer: molecular features and potential therapeutic approaches. Biochimica et Biophysica Acta (BBA)-Reviews on Cancer. 2009, 1796, 162–175. [Google Scholar] [CrossRef]
- Franke, C.M.; Gu, V.W.; Grimm, B.G.; Cassady, V. C.; White, J. R.; Weigel, R. J.; & Kulak, M. V. ; & Kulak, M. V. TFAP2C regulates carbonic anhydrase XII in human breast cancer. Oncogene. 2020, 39, 1290–1301. [Google Scholar] [CrossRef]
- Molehin, D.; Castro-Piedras, I.; Sharma, M.; Sennoune, S. R.; Arena, D.; Manna, P. R.; Pruitt, K. Aromatase acetylation patterns and altered activity in response to sirtuin inhibition. Molecular Cancer Research. 2018, 16, 1530–1542. [CrossRef] [PubMed]
- Rong, M.; Li, Y.; Guo, X.; Zong, T.; Ma, Z.; Li, P. An ISSA-RF Algorithm for Prediction Model of Drug Compound Molecules Antagonizing ERα Gene Activity. Oncologie. 2022, 24, 309–327. [Google Scholar] [CrossRef]
- Chen, F.; Yin, X.; Wang, Y.; Lv, Y.; Sheng, S.; Ouyang, S.; Zhong, Y. Pharmacokinetics, Tissue Distribution, and Druggabil5ty Prediction of the Natural Anticancer Active Compound Cytisine N-Isoflavones Combined with Computer Simulation. Biological & pharmaceutical bulletin.2020,43.976-984. [CrossRef]
- Wang, R. Comparison of Decision Tree, Random Forest and Linear Discriminant analysis models in breast cancer prediction. Journal of Physics: Conference Series, 0120; 43. [Google Scholar] [CrossRef]
- Hopkins, A.L. Network pharmacology. Nat Biotechnol. 2007, 25, 1110–1111. [Google Scholar] [CrossRef] [PubMed]
- Betz, M. ;Saxena,K. ;Schwalbe, H. Biomolecular NMR: a chaperone to drug discovery. Current Opinion in Chemical Biology. 2006, 10, 219–225. [Google Scholar] [CrossRef]
- Mansa, F. Simon G. Assessing inhibitory activity of probiotic culture supernatants against Pseudomonas aeruginosa: a comparative methodology between agar diffusion, broth culture and microcalorimetry. World journal of microbiology & biotechnology. 2019, 35, 49. [Google Scholar] [CrossRef]
- Stefan, L. Optimizing the hit-to-lead process using SPR analysis. Assay and drug development technologies. 2004, 2, 407–415. [Google Scholar] [CrossRef]
- Taoka, K. I.; Kawahara, I.; Shinya, S.; Harada, K. I.; Yamashita, E.; Shimatani, Z.; Furuita, K.; Tomoaki, M.; Tokitaka, M.; Terada, R.; Nakagawa, A.; Fujiwara, T.; Tsuji, H.; Kojima, C. Multifunctional chemical inhibitors of the florigen activation complex discovered by structure-based high-throughput screening. The Plant journal : for cell and molecular biology. 2022, 112, 1337–1349. [Google Scholar] [CrossRef]
- Hansch, C.; Maloney, P.; Fujita, T. Correlation of Bioactivityof Phenoxyacetic Acids with Hammett Substituent Constants and Partition Coefficients. Nature. 1962, 194, 178–180. [Google Scholar] [CrossRef]
- Mansouri, K; Kleinstreuer, N; Abdelaziz, AM; Alberga, D; Alves, VM; Andersson, PL; Andrade, CH; Bai, F; Balabin, I; Ballabio, D; Benfenati, E; Bhhatarai, B; Boyer, S et al. CoMPARA: Collaborative Modeling Project for Androgen Receptor Activity. Environ Health Perspect. 2020, 128, 27002. [CrossRef]
- Putri, D. E. K.; Pranowo, H. D.; Wijaya, A. R.; Suryani, N.; Utami, M.; Suma, A. A. T. ; Chung,W. J.; Almutairi,S.M.; Hussein,D.S.;Rasheed,R.A.; Ranganathan, V. The predicted models of anti-colon cancer and anti-hepatoma activities of substituted 4-anilino coumarin derivatives using quantitative structure-activity relationship (QSAR). Journal of King Saud University - Science. 2022, 34, 101837. [Google Scholar] [CrossRef]
- Wang, P.; Shehu, A.I.; Ma, X. The Opportunities of Metabolomics in Drug Safety Evaluation. Current pharmacology reports. 2017, 3, 10–15. [Google Scholar] [CrossRef] [PubMed]
- Gohlke, H.; Hendlich, M.; Klebe, G. Knowledge-based scoring function to predict protein-ligand interactions. J Mol Biol. 2000, 295, 337–56. [Google Scholar] [CrossRef] [PubMed]
- Jones, G.; Willett, P.; Glen, R. C.; Leach, A. R.; Taylor, R. Development and validation of a genetic algorithm for flexible docking. Journal of molecular biology. 1997, 267, 727–748. [Google Scholar] [CrossRef] [PubMed]
- Head, R. D.; Smythe, M. L.; Oprea, T. I.; Waller, C. L.; Green, S. M.; Marshall, G. R. VALIDATE: A New Method for the Receptor-Based Prediction of Binding Affinities of Novel Ligands. Journal of the American Chemical Society. 1996, 118, 3959–3969. [Google Scholar] [CrossRef]
- Yang, Y.; Chen, S.; Wang, Q.; Niu, M. M.; Qu, Y.; Zhou, Y. Identification of novel and potent dual-targeting HDAC1/SPOP inhibitors using structure-based virtual screening, molecular dynamics simulation and evaluation of in vitro and in vivo antitumor activity. Frontiers in Pharmacology. 2023, 14, 1208740. [Google Scholar] [CrossRef]
- Xiao, X.; Qin, M.; Zhang, F.; Su, Y.; Zhou, B.; Zhou, Z. Understanding the Mechanism of Activation/Deactivation of GLP-1R via Accelerated Molecular Dynamics Simulation. Australian Journal of Chemistry. 2021, 74, 211–218. [Google Scholar] [CrossRef]
- Stephenson, N.; Shane, E.; Chase, J.; Rowland, J.; Ries, D.; Justice, N. ;Leong. Z;Chan,L.;Cao, R. Survey of machine learning techniques in drug discovery. Current Drug Metabolism. 2019, 20, 185–193. [Google Scholar] [CrossRef]
- Martinčič, R.; Kuzmanovski, I.; Wagner, A. Novič M. Development of models for prediction of the antioxidant activity of derivatives of natural compounds. Anal Chim Acta. 2015, 868, 23–35. [Google Scholar] [CrossRef]
- Dutt, R.; Madan, A.K. Development and application of novel molecular descriptors for predicting bioactivity. Medicinal Chemistry Research. 2017, 26, 1988–2006. [Google Scholar] [CrossRef]
- Lane, T. R.; Urbina, F. , Rank, L. ; Gerlach, J.; Riabova, O.; Lepioshkin, A.; Kazakova,E.; Vocat,A.; Tkachenko,V.; Cole, S.; Makarov,V.; Ekins, S. Machine Learning Models for Mycobacterium tuberculosis In Vitro Activity: Prediction and Target Visualization. Molecular pharmaceutics. 2021, 19, 674–689. [Google Scholar] [CrossRef] [PubMed]
- Hrynkiewicz, M.; Iwaniak, A.; Bucholska, J.; Minkiewicz, P.; Darewicz, M. Structure⁻Activity Prediction of ACE Inhibitory/Bitter Dipeptides-A Chemometric Approach Based on Stepwise Regression. Molecules. 2019, 24, 950. [Google Scholar] [CrossRef]
- Kato, Y.; Hamada, S.; Goto, H. Validation Study of QSAR/DNN Models Using the Competition Datasets. Mol Inform. 2020, 39, e1900154. [Google Scholar] [CrossRef]
- Cai, C.; Guo, P.; Zhou, Y.; Zhou, J.; Wang, Q.; Zhang, F.; Fang, J.; Cheng, F. Deep Learning-Based Prediction of Drug-Induced Cardiotoxicity. J Chem Inf Model. 2019, 59, 1073–1084. [Google Scholar] [CrossRef] [PubMed]
- Luo, H.; Xiang, Y.; Fang, X.; Lin, W.; Wang, F.; Wu, H.; Wang, H. BatchDTA: implicit batch alignment enhances deep learning-based drug-target affinity estimation. Brief Bioinform. 2022, 23, bbac260. [Google Scholar] [CrossRef]
- Hentabli, H.; Bengherbia, B.; Saeed, F.; Salim, N.; Nafea, I.; Toubal, A.; Nasser, M. Convolutional Neural Network Model Based on 2D Fingerprint for Bioactivity Prediction. International Journal of Molecular Sciences. 2022, 23, 13230. [Google Scholar] [CrossRef] [PubMed]
- Dadfar, E.; Shafiei, F.; Isfahani, T.M. Structural Relationship Study of Octanol-Water Partition Coefficient of Some Sulfa Drugs Using GA-MLR and GA-ANN Methods. Current Computer - Aided Drug Design. 2020, 16, 207–221. [Google Scholar] [CrossRef]
- Zhang, G.; Dai, Z.; Dai, X. A Novel Hybrid CNN-SVR for CRISPR/Cas9 Guide RNA Activity Prediction. Front Genet. 2020, 10, 1303. [Google Scholar] [CrossRef]
- Xing, G.; Liang, L.; Deng, C.; Hua, Y.; Chen, X.; Yang, Y.; Liu, H.; Lu, T.; Chen, Y.; Zhang, Y. Activity Prediction of Small Molecule Inhibitors for Antirheumatoid Arthritis Targets Based on Artificial Intelligence. ACS Comb Sci. 2020, 22, 873–886. [Google Scholar] [CrossRef]
- Badura, A.; Krysiński, J.; Nowaczyk, A.; Buciński, A. Application of artificial neural networks to the prediction of antifungal activity of imidazole derivatives against Candida albicans. Chemometrics and Intelligent Laboratory Systems. 2022, 222, 104501. [Google Scholar] [CrossRef]
- Afolabi, L. T.; Saeed, F.; Hashim, H.; Petinrin, O. O. Ensemble learning method for the prediction of new bioactive molecules. PloS one, 2018, 13, e0189538. [Google Scholar] [CrossRef]
- Tavakoli, H.; Ghasemi, B.J. An improved ensemble learning machine for bioactivityprediction of tyrosine kinase inhibitors. Journal of Chemometrics. 2015, 29, 213–223. [Google Scholar] [CrossRef]
- Zhong, J.; Xuan, W.; Lu, S.; Cui, S.; Zhou, Y.; Tang, M. ; Qu,X. ; Lu,W.; Huo,H.;Zhang,C.; Zhang, N.; Niu, B. Discovery of ANO1 Inhibitors based on Machine learning and molecule docking simulation approaches. European journal of pharmaceutical sciences : official journal of the European Federation for Pharmaceutical Sciences. 2023, 184, 106408. [Google Scholar] [CrossRef]
- Wu, Z.; Lei, T.; Shen, C.; Wang, Z.; Cao, D.; Hou, T. ADMET evaluation in drug discovery. 19. Reliable prediction of human cytochrome P450 inhibition using artificial intelligence approaches. Journal of chemical information and modeling. 2019, 59, 4587–4601. [Google Scholar] [CrossRef]
- Wishart DS, Feunang YD, Guo AC, Lo EJ, Marcu A, Grant JR, Sajed T, Johnson D, Li C, Sayeeda Z, Assempour N, Iynkkaran I, Liu Y, Maciejewski A, Gale N, Wilson A, Chin L, Cummings R, Le D, Pon A, Knox C, Wilson M. DrugBank 5.0: a major update to the DrugBank database for 2018. Nucleic Acids Res. 2017 Nov 8. [CrossRef]
- Jaganathan, K; Tayara, H; Chong, KT. An Explainable Supervised Machine Learning Model for Predicting Respiratory Toxicity of Chemicals Using Optimal Molecular Descriptors. Pharmaceutics. 2022, 14, 832. [CrossRef]
- Xia, J; Zhang, J; Wang, Y; Han, L; Yan, H. WC-KNNG-PC: Watershed clustering based on k-nearest-neighbor graph and Pauta Criterion. Pattern Recognition. 2022, 121, 108177. [CrossRef]
- Bahl, L.; Brown, P.; De Souza, P.; Mercer, R. Maximum mutual information estimation of hidden Markov model parameters for speech recognition. ICASSP '86. IEEE International Conference on Acoustics, Speech, and Signal Processing, Tokyo, Japan, 1986, pp. -52. [CrossRef]
- Benesty, J.; Chen, J.; Huang, Y.; Cohen, I. Pearson Correlation Coefficient. 2009; Volume 2, pp. 1-4.
- Székely, G.J.; Rizzo, M.L.; Bakirov, N.K. Measuring and Testing Dependence by Correlation of Distance. The Annals of Statistics. 2008,35, 2769-2794. [CrossRef]
- Schapire, R.E. Explaining AdaBoost. In Empirical Inference: Festschrift in Honor of Vladimir N. Vapnik, Schölkopf, B., Luo, Z., Vovk, V., Eds.; Springer Berlin Heidelberg: Berlin, Heidelberg, 2013. [Google Scholar]
- Chen, T.; Guestrin, C. XGBoost: A Scalable Tree Boosting System. In Proceedings of the Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, San Francisco, California, USA, 2016; pp. 785–794.
- Haas, BC; Goetz, AE; Bahamonde, A; McWilliams, JC; Sigman, MS. Predicting relative efficiency of amide bond formation using multivariate linear regression. Proc Natl Acad Sci U S A. 2022, 119, e2118451119. [CrossRef]
- Darwiche M, Mokhiamar O.
- J]. Alexandria Engineering Journal, 2022, 61, 6119–6128. [CrossRef]
- Shahriari,B; Swersky,K; Wang,Z; Adams, R. P.; de Freitas, N. Taking the human out of the loop: A review of Bayesian Optimization. Proceedings of the IEEE. 2016, 104, 148–175. [Google Scholar] [CrossRef]
- Python Release 3.9.6. Available online: https://www.python.org/downloads/release/python-396/ (accessed on 20 May 2023).
- Yan, Y; He, M; Zhao, L; Wu, H; Zhao, Y; Han, L; Wei, B; Ye, D; Lv, X; Wang, Y; Yao, W; Zhao, H; Chen, B; Jin, Z; Wen, J; Zhu, Y; Yu, T; Jin, F; Wei, M. A novel HIF-2α targeted inhibitor suppresses hypoxia-induced breast cancer stemness via SOD2-mtROS-PDI/GPR78-UPRER axis. Cell Death Differ. 2022, 29, 1769–1789. [CrossRef]
| Molecular descriptor | Minimum value | Maximum value | Mean value | Standard deviation |
|---|---|---|---|---|
| pIC50 | 2.456 | 10.34 | 6.59 | 1.42 |
| nAcid | 0.00 | 4.00 | 0.11 | 0.35 |
| ALogP | -23.11 | 5.18 | 1.11 | 1.43 |
| ALogp2 | 0.00 | 533.84 | 3.29 | 12.83 |
| AMR | 54.07 | 517.43 | 116.56 | 31.57 |
| . | . | . | . | . |
| . | . | . | . | . |
| . | . | . | . | . |
| WPATH | 349.00 | 301690.00 | 2709.62 | 7194.53 |
| WPOL | 14.00 | 230.00 | 46.28 | 13.29 |
| XLogP | -3.59 | 14.28 | 2.97 | 1.62 |
| Zagreb | 6.00 | 748.00 | 150.72 | 41.45 |
| Ranking | Feature name | Contribution degree | Ranking | Feature name | Contribution degree |
|---|---|---|---|---|---|
| 1 | MDEC-23 | 0.3578 | 11 | SHsOH | 0.0281 |
| 2 | LipoaffinityIndex | 0.0802 | 12 | BCUTp-1h | 0.0259 |
| 3 | BCUTc-1l | 0.0761 | 13 | VPC-6 | 0.0247 |
| 4 | minsssN | 0.0548 | 14 | minHBa | 0.0214 |
| 5 | maxHsOH | 0.0530 | 15 | hmin | 0.0210 |
| 6 | minsOH | 0.0406 | 16 | minHBint10 | 0.0203 |
| 7 | BCUTc-1h | 0.0371 | 17 | ETA_BetaP_s | 0.0190 |
| 8 | maxssO | 0.0355 | 18 | SPC-6 | 0.0178 |
| 9 | mindssC | 0.0287 | 19 | MDEO-12 | 0.0152 |
| 10 | ATSc3 | 0.0282 | 20 | minHBint5 | 0.0146 |
| Model | Hyperparameter | Meaning | Range |
|---|---|---|---|
| XGBoost | n_estimators | Decision tree quantity | [50,100,150,200] |
| max_depth | Maximum depth of the tree | (1,10)evenly distributed, step size is 1. | |
| learning_rate | Learning rate | (10-6,1)logarithmically uniform distribution. | |
| subsample | Ratio of subsamples to training samples | (0.5,1)evenly distributed. | |
| Colsample_bytree | Feature sampling rate | (0.5,1)evenly distributed. | |
| MLR | fit_intercept | Whether to fit the intercept | [True, False] |
| SVR | C | Regularization parameter | (10-6,1)logarithmically uniform distribution. |
| gamma | Kernel value range | (10-6,1)logarithmically uniform distribution. | |
| kernel | Kernel type | ['linear', 'rbf'] |
| Model | Parameter setting |
|---|---|
| MLR | default parameter |
| SVR | kernel='rbf', C=1e3, gamma=0.1 |
| XGBoost | objective='reg:squarederror', colsample_bytree=0.3, |
| learning_rate=0.1, | |
| max_depth=5, | |
| alpha=10 | |
| n_estimators=10 | |
| CNN-LSTM | a 1D convolutional layer was established that receives input features of 64 size and holds hidden states of 50 size. |
| Model | RMSE | MAE | R2 |
|---|---|---|---|
| MLR | 0.9416 | 0.7002 | 0.5955 |
| SVR | 1.1727 | 0.7698 | 0.3726 |
| XGBoost | 1.9316 | 1.6702 | 0.1591 |
| CNN-LSTM | 0.7486 | 0.5171 | 0.7443 |
| BHO-AdaBoosting | 0.6920 | 0.4837 | 0.8155 |
| Ranking | MF | pIC50 | Ranking | MF | pIC50 |
|---|---|---|---|---|---|
| 1 | C25H22O3 | 8.583 | 26 | C24H19FO5 | 6.890 |
| 2 | C25H19FO3S | 7.953 | 27 | C51H67N3O10 | 6.885 |
| 3 | C29H33NO2 | 7.859 | 28 | C29H34N2O4 | 6.878 |
| 4 | C31H24FNO3 | 7.733 | 29 | C65H107N21O16 | 6.871 |
| 5 | C36H33FN2O3 | 7.708 | 30 | C65H107N21O16 | 6.871 |
| 6 | C34H30O8S | 7.602 | 31 | C23H17FO3 | 6.854 |
| 7 | C36H33FN2O3 | 7.583 | 32 | C26H23FO5 | 6.847 |
| 8 | C29H27FN2O3 | 7.527 | 33 | C65H107N21O16 | 6.826 |
| 9 | C27H30NO4 | 7.510 | 34 | C64H105N21O16 | 6.822 |
| 10 | C26H22O5 | 7.435 | 35 | C32H34O4 | 6.808 |
| 11 | C30H29FN2O3 | 7.401 | 36 | C62H105N21O16 | 6.764 |
| 12 | C31H23FO3 | 7.384 | 37 | C19H26OS | 6.757 |
| 13 | C30H23FO2 | 7.380 | 38 | C63H101N19O17 | 6.723 |
| 14 | C25H20O4 | 7.348 | 39 | C18H24OS | 6.690 |
| 15 | C31H33FN2O3 | 7.344 | 40 | C27H21FO4 | 6.541 |
| 16 | C29H28FNO3 | 7.327 | 41 | C22H31NO3 | 6.494 |
| 17 | C25H19FO4 | 7.326 | 42 | C21H29NO3 | 6.343 |
| 18 | C29H26FNO3 | 7.239 | 43 | C28H26ClN3O3 | 6.289 |
| 19 | C31H38N2O5 | 7.144 | 44 | C29H28ClN3O3 | 5.978 |
| 20 | C52H71N3O10 | 7.135 | 45 | C26H25ClN4O2 | 5.686 |
| 21 | C26H21FO3 | 7.106 | 46 | C23H26ClN3O3 | 5.544 |
| 22 | C25H22O6 | 7.054 | 47 | C21H22ClN3O3 | 5.411 |
| 23 | C29H33NO2 | 7.048 | 48 | C23H24ClN3O3 | 5.396 |
| 24 | C16H12Cl2M2O2 | 6.969 | 49 | C23H24ClN5O2 | 5.386 |
| 25 | C31H38N2O4 | 6.911 | 50 | C23H27ClN4O2 | 5.358 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




