Submitted:
03 February 2023
Posted:
03 February 2023
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results and Discussion
2.1. Hydrogen bonds analyses based on trajectories of classical MD
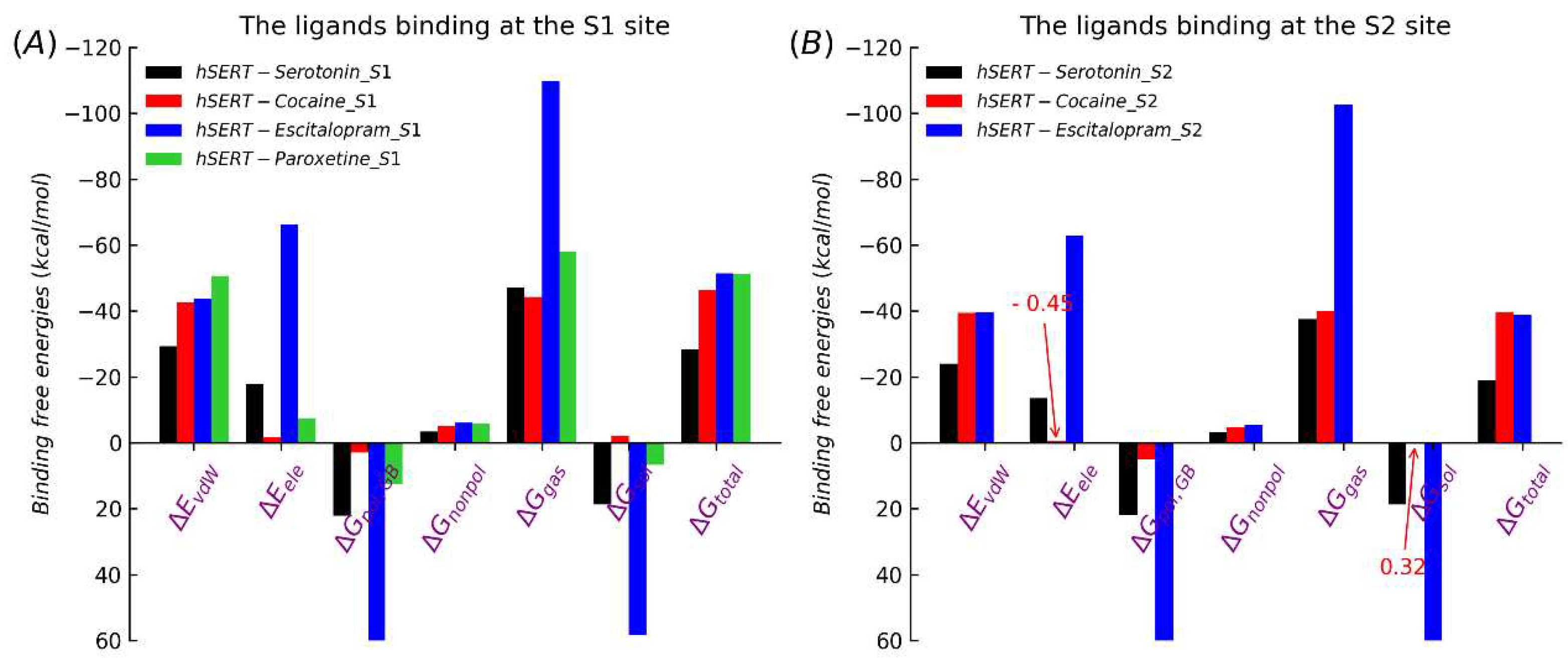
2.2. Binding free energies at the equilibrium states
2.3. Potential of mean forces along the unbinding pathways
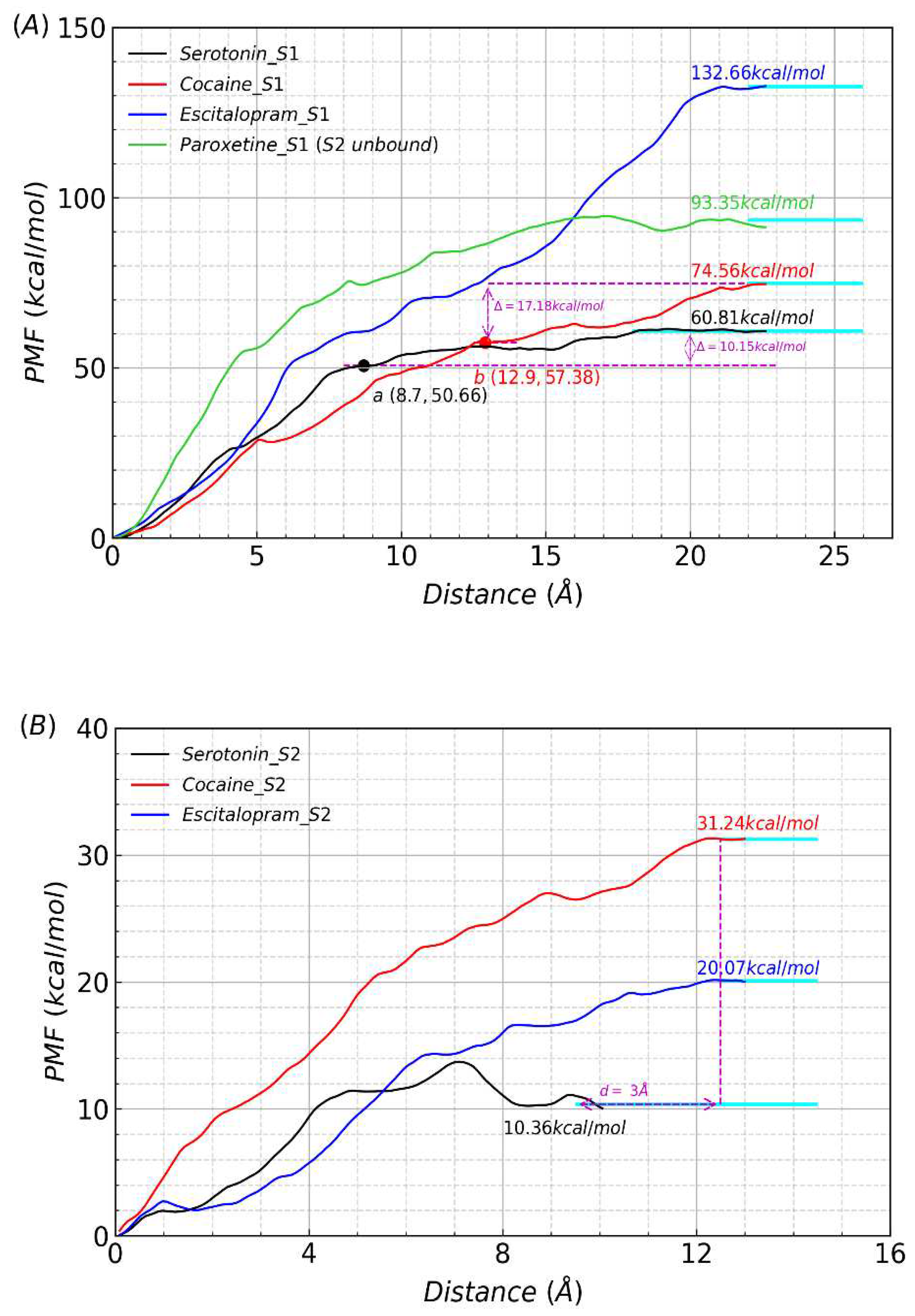
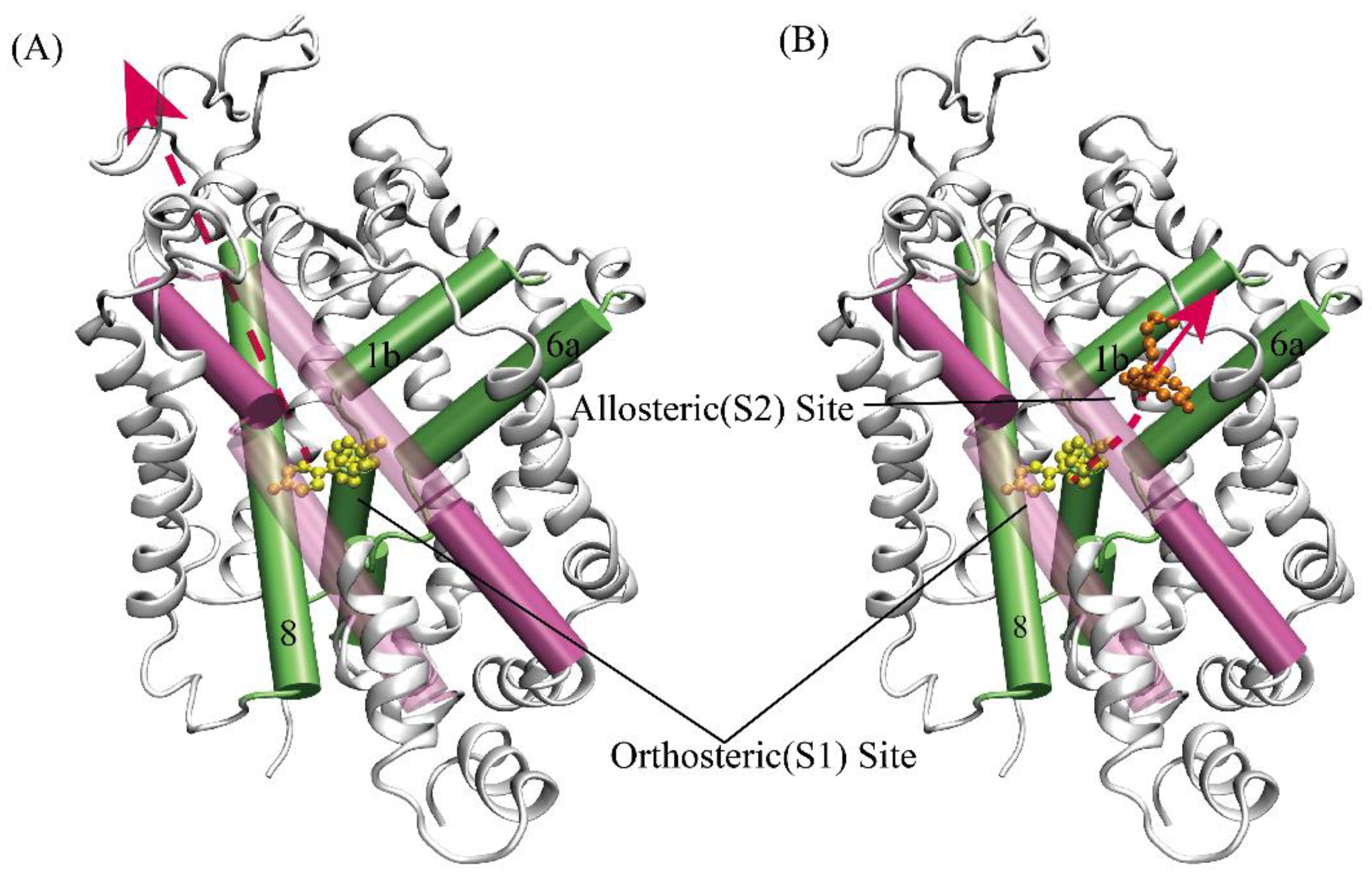
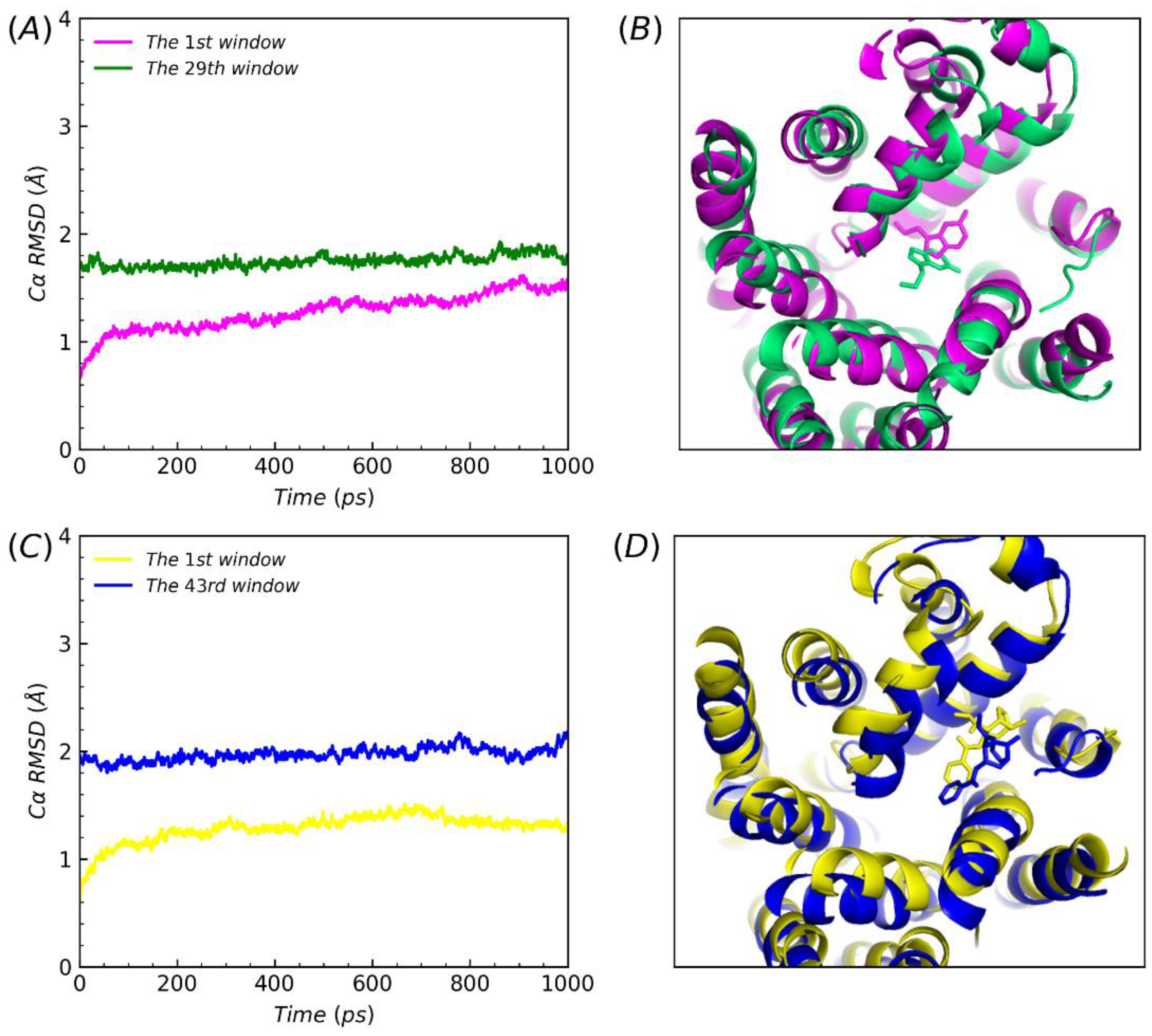
2.4. Structural analysis and dissociative process of PMF simulations
3. Materials and Methods
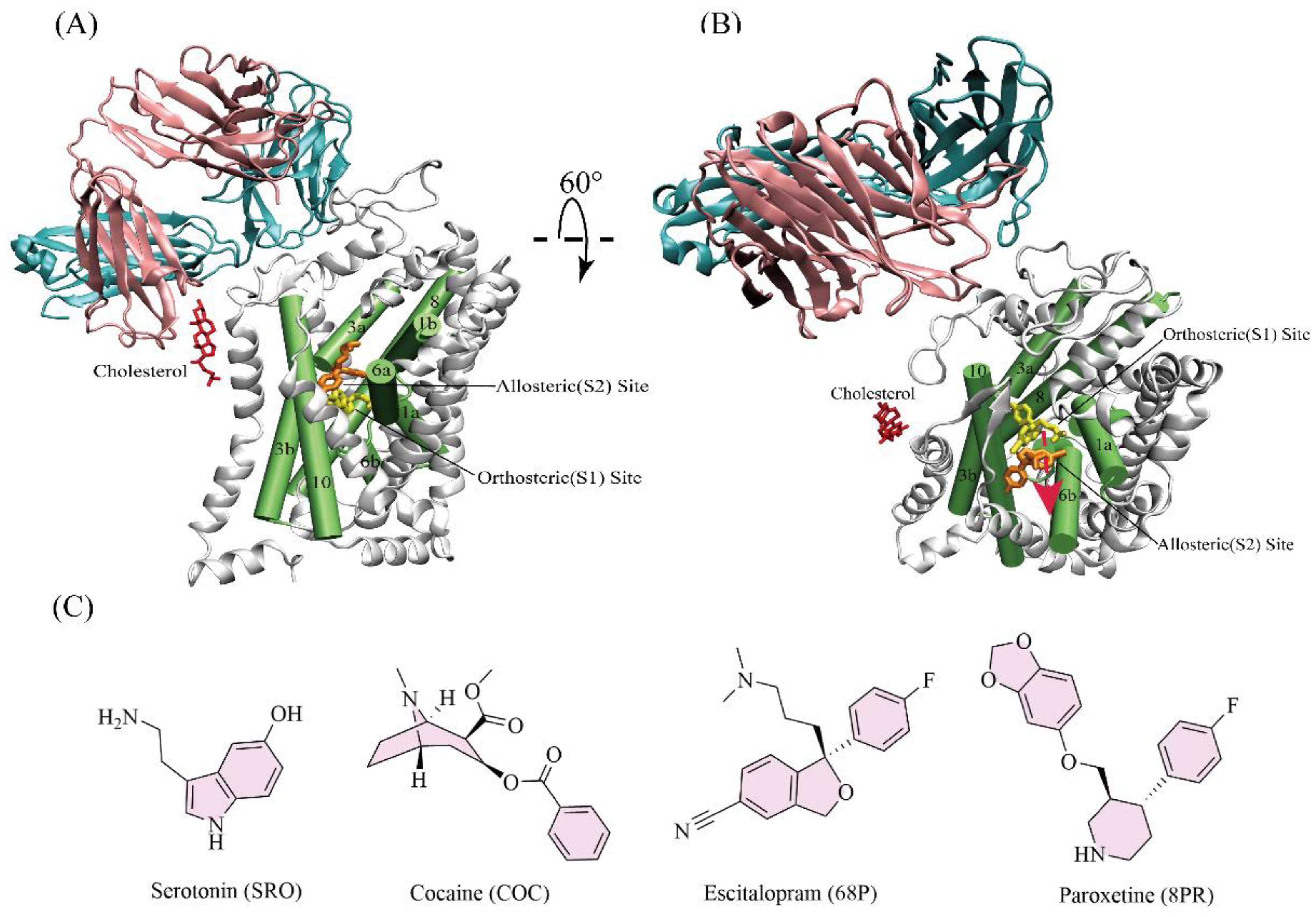
3.1. Systems Preparation
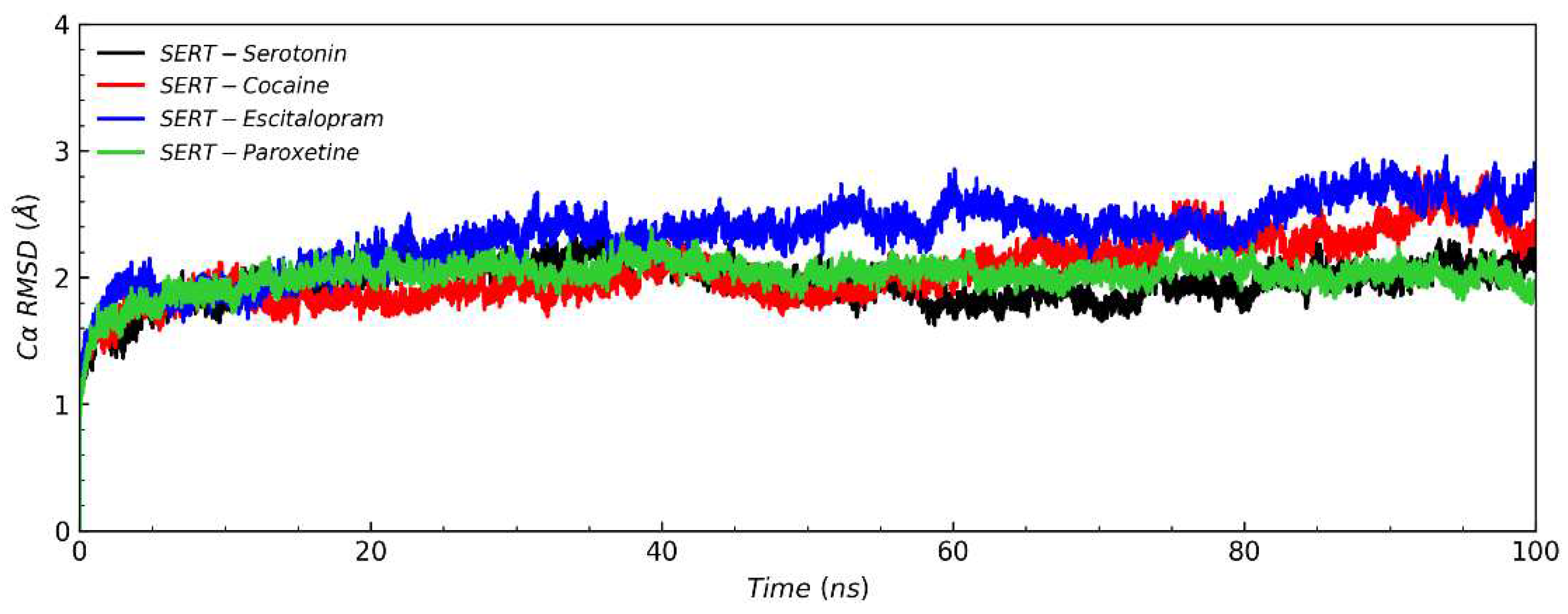
3.2. Conventional Molecular Dynamics Simulation
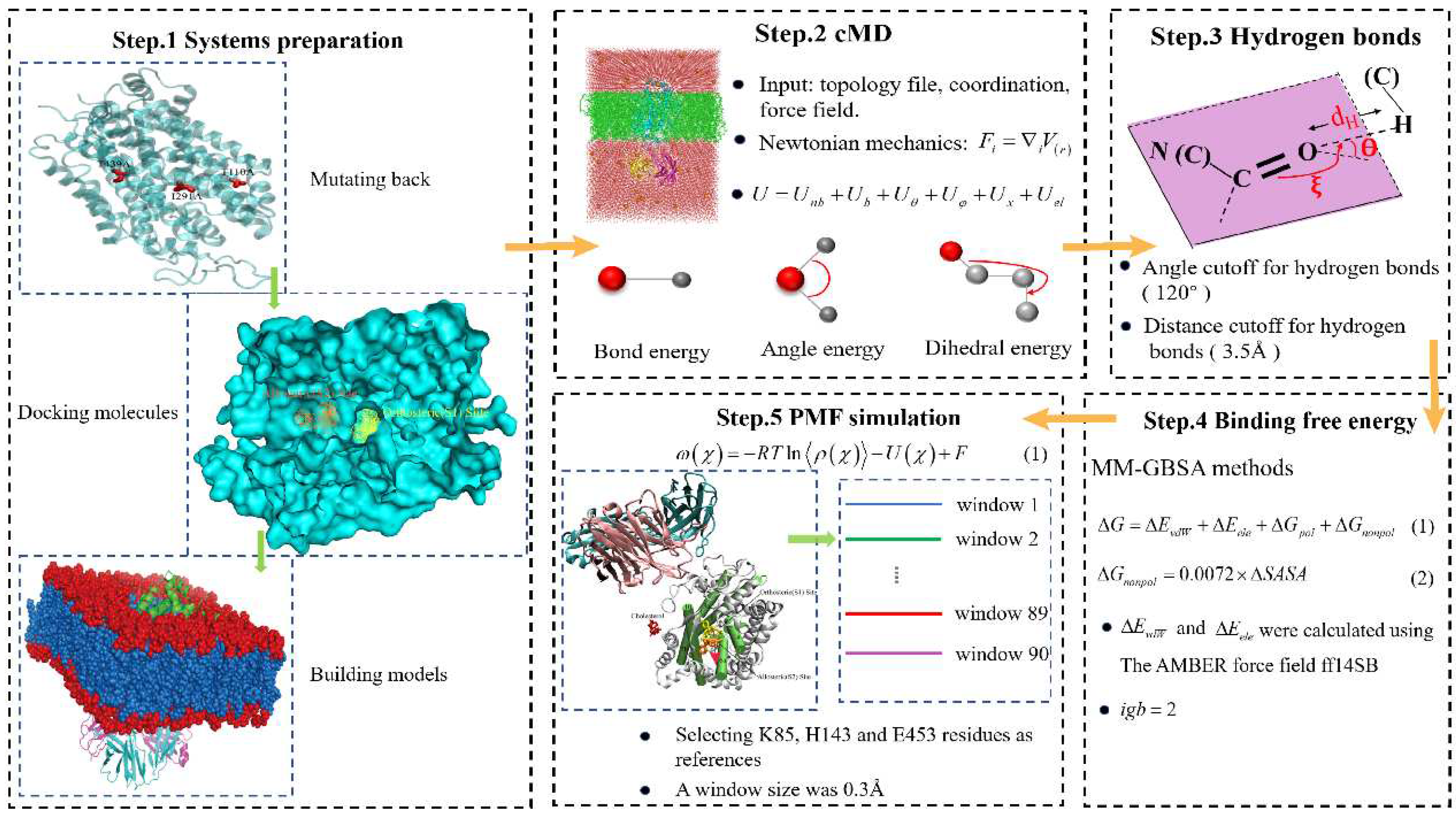
3.3. Molecular Mechanics/Generalized Born Surface Area (MM/GBSA)
3.4. Steered Molecular Dynamics Simulation
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Sepanlou, S.G.; Safiri, S.; Bisignano, C.; Ikuta, K.S.; Merat, S.; Saberifiroozi, M.; Poustchi, H.; Tsoi, D.; Colombara, D.V.; Abdoli, A. The global, regional, and national burden of cirrhosis by cause in 195 countries and territories, 1990–2017: a systematic analysis for the Global Burden of Disease Study 2017. The Lancet gastroenterology & hepatology 2020, 5, 245–266. [Google Scholar]
- Coccaro, E.F. New Hope for Patients with Major Depressive Disorder? 2019, 381, 980-981. [CrossRef]
- Recourt, K.; de Boer, P.; Zuiker, R.; Luthringer, R.; Kent, J.; van der Ark, P.; Van Hove, I.; van Gerven, J.; Jacobs, G.; van Nueten, L. The selective orexin-2 antagonist seltorexant (JNJ-42847922/MIN-202) shows antidepressant and sleep-promoting effects in patients with major depressive disorder. Translational psychiatry 2019, 9, 1–10. [Google Scholar]
- Saltiel, P.F.; Silvershein, D.I. Major depressive disorder: mechanism-based prescribing for personalized medicine. Neuropsychiatric disease and treatment 2015, 11, 875. [Google Scholar] [CrossRef] [PubMed]
- Krishnan, V.; Nestler, E.J. The molecular neurobiology of depression. Nature 2008, 455, 894–902. [Google Scholar] [CrossRef] [PubMed]
- Kupfer, D.J.; Frank, E.; Phillips, M.L. Major depressive disorder: new clinical, neurobiological, and treatment perspectives. The Lancet 2012, 379, 1045–1055. [Google Scholar] [CrossRef] [PubMed]
- Caspi, A.; Sugden, K.; Moffitt, T.E.; Taylor, A.; Craig, I.W.; Harrington, H.; McClay, J.; Mill, J.; Martin, J.; Braithwaite, A. Influence of life stress on depression: moderation by a polymorphism in the 5-HTT gene. Science 2003, 301, 386–389. [Google Scholar] [CrossRef]
- Fuller, R.W.; Wong, D.T. Serotonin uptake and serotonin uptake inhibition. Annals of the New York Academy of Sciences 1990. [Google Scholar] [CrossRef] [PubMed]
- Murphy, D.L.; Lerner, A.; Rudnick, G.; Lesch, K.-P. Serotonin transporter: gene, genetic disorders, and pharmacogenetics. Molecular interventions 2004, 4, 109. [Google Scholar] [CrossRef]
- Chen, N. Reith ME, and Quick MW. Synaptic uptake and beyond: the sodium-and chloride-dependent neurotransmitter transporter family SLC6. Pflügers Arch 2004, 447, 519-531. [CrossRef]
- Sitte, H.H.; Freissmuth, M. Amphetamines, new psychoactive drugs and the monoamine transporter cycle. Trends in pharmacological sciences 2015, 36, 41–50. [Google Scholar] [CrossRef]
- Niello, M.; Gradisch, R.; Loland, C.J.; Stockner, T.; Sitte, H.H. Allosteric modulation of neurotransmitter transporters as a therapeutic strategy. Trends in Pharmacological Sciences 2020, 41, 446–463. [Google Scholar] [CrossRef]
- Rothman, R.B.; Partilla, J.S.; Baumann, M.H.; Lightfoot-Siordia, C.; Blough, B.E. Studies of the biogenic amine transporters. 14. Identification of low-efficacy “partial” substrates for the biogenic amine transporters. Journal of Pharmacology and Experimental Therapeutics 2012, 341, 251–262. [Google Scholar] [CrossRef] [PubMed]
- Hasenhuetl, P.S.; Bhat, S.; Freissmuth, M.; Sandtner, W. Functional selectivity and partial efficacy at the monoamine transporters: a unified model of allosteric modulation and amphetamine-induced substrate release. Molecular Pharmacology 2019, 95, 303–312. [Google Scholar] [CrossRef] [PubMed]
- Xue, W.; Fu, T.; Deng, S.; Yang, F.; Yang, J.; Zhu, F. Molecular mechanism for the allosteric inhibition of the human serotonin transporter by antidepressant escitalopram. ACS chemical neuroscience 2022, 13, 340–351. [Google Scholar] [CrossRef] [PubMed]
- Yang, D.; Gouaux, E. Illumination of serotonin transporter mechanism and role of the allosteric site. Science Advances 2021, 7, eabl3857. [Google Scholar] [CrossRef]
- Chan, M.C.; Selvam, B.; Young, H.J.; Procko, E.; Shukla, D. The substrate import mechanism of the human serotonin transporter. Biophysical journal 2022, 121, 715–730. [Google Scholar] [CrossRef] [PubMed]
- Gether, U.; Andersen, P.H.; Larsson, O.M.; Schousboe, A. Neurotransmitter transporters: molecular function of important drug targets. Trends in pharmacological sciences 2006, 27, 375–383. [Google Scholar] [CrossRef] [PubMed]
- Kristensen, A.S.; Andersen, J.; Jørgensen, T.N.; Sørensen, L.; Eriksen, J.; Loland, C.J.; Strømgaard, K.; Gether, U. SLC6 neurotransmitter transporters: structure, function, and regulation. Pharmacological reviews 2011, 63, 585–640. [Google Scholar] [CrossRef] [PubMed]
- Hahn, M.; Blakely, R. Monoamine transporter gene structure and polymorphisms in relation to psychiatric and other complex disorders. The pharmacogenomics journal 2002, 2, 217–235. [Google Scholar] [CrossRef] [PubMed]
- Severinsen, K.; Koldsø, H.; Thorup, K.A.V.; Schjøth-Eskesen, C.; Møller, P.T.; Wiborg, O.; Jensen, H.H.; Sinning, S.; Schiøtt, B. Binding of mazindol and analogs to the human serotonin and dopamine transporters. Molecular pharmacology 2014, 85, 208–217. [Google Scholar] [CrossRef]
- Henry, L.K.; Field, J.R.; Adkins, E.M.; Parnas, M.L.; Vaughan, R.A.; Zou, M.-F.; Newman, A.H.; Blakely, R.D. Tyr-95 and Ile-172 in transmembrane segments 1 and 3 of human serotonin transporters interact to establish high affinity recognition of antidepressants. Journal of Biological Chemistry 2006, 281, 2012–2023. [Google Scholar] [CrossRef]
- Andersen, J.; Taboureau, O.; Hansen, K.B.; Olsen, L.; Egebjerg, J.; Strømgaard, K.; Kristensen, A.S. Location of the antidepressant binding site in the serotonin transporter. Journal of Biological Chemistry 2009, 284, 10276–10284. [Google Scholar] [CrossRef] [PubMed]
- Coleman, J.A.; Gouaux, E. Structural basis for recognition of diverse antidepressants by the human serotonin transporter. Nature structural & molecular biology 2018, 25, 170–175. [Google Scholar] [CrossRef] [PubMed]
- Gradisch, R.; Szöllősi, D.; Niello, M.; Lazzarin, E.; Sitte, H.H.; Stockner, T. Occlusion of the human serotonin transporter is mediated by serotonin-induced conformational changes in the bundle domain. Journal of Biological Chemistry 2022, 298. [Google Scholar] [CrossRef]
- Yamashita, A.; Singh, S.K.; Kawate, T.; Jin, Y.; Gouaux, E. Crystal structure of a bacterial homologue of Na+/Cl--dependent neurotransmitter transporters. Nature 2005, 437, 215–223. [Google Scholar] [CrossRef] [PubMed]
- Coleman, J.A.; Green, E.M.; Gouaux, E. X-ray structures and mechanism of the human serotonin transporter. Nature 2016, 532, 334–339. [Google Scholar] [CrossRef] [PubMed]
- Zhu, R.; Sinwel, D.; Hasenhuetl, P.S.; Saha, K.; Kumar, V.; Zhang, P.; Rankl, C.; Holy, M.; Sucic, S.; Kudlacek, O. Nanopharmacological force sensing to reveal allosteric coupling in transporter binding sites. Angewandte Chemie 2016, 128, 1751–1754. [Google Scholar] [CrossRef]
- Jørgensen, A.M.; Topiol, S. Driving forces for ligand migration in the leucine transporter. Chemical biology & drug design 2008, 72, 265–272. [Google Scholar] [CrossRef] [PubMed]
- Topiol, S.; Bang-Andersen, B.; Sanchez, C.; Bøgesø, K.P. Exploration of insights, opportunities and caveats provided by the X-ray structures of hSERT. Bioorganic & Medicinal Chemistry Letters 2016, 26, 5058–5064. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Zheng, G.; Fu, T.; Hong, J.; Li, F.; Yao, X.; Xue, W.; Zhu, F. The binding mode of vilazodone in the human serotonin transporter elucidated by ligand docking and molecular dynamics simulations. Physical Chemistry Chemical Physics 2020, 22, 5132–5144. [Google Scholar] [CrossRef]
- Millan, M.J. Dual-and triple-acting agents for treating core and co-morbid symptoms of major depression: novel concepts, new drugs. Neurotherapeutics 2009, 6, 53–77. [Google Scholar] [CrossRef]
- Stuivenga, M.; Giltay, E.J.; Cools, O.; Roosens, L.; Neels, H.; Sabbe, B. Evaluation of vilazodone for the treatment of depressive and anxiety disorders. Expert Opinion on Pharmacotherapy 2019, 20, 251–260. [Google Scholar] [CrossRef] [PubMed]
- Crits-Christoph, P.; Siqueland, L.; Blaine, J.; Frank, A.; Luborsky, L.; Onken, L.S.; Muenz, L.R.; Thase, M.E.; Weiss, R.D.; Gastfriend, D.R. Psychosocial treatments for cocaine dependence: National Institute on Drug Abuse collaborative cocaine treatment study. Archives of general psychiatry 1999, 56, 493–502. [Google Scholar] [CrossRef] [PubMed]
- Lambert, N.M.; McLeod, M.; Schenk, S. Subjective responses to initial experience with cocaine: an exploration of the incentive–sensitization theory of drug abuse. Addiction 2006, 101, 713–725. [Google Scholar] [CrossRef] [PubMed]
- Larsen, M.A.B.; Plenge, P.; Andersen, J.; Eildal, J.N.; Kristensen, A.S.; Bøgesø, K.P.; Gether, U.; Strømgaard, K.; Bang-Andersen, B.; Loland, C.J. Structure–activity relationship studies of citalopram derivatives: examining substituents conferring selectivity for the allosteric site in the 5-HT transporter. British Journal of Pharmacology 2016, 173, 925–936. [Google Scholar] [CrossRef] [PubMed]
- Frauenfelder, H.; Wolynes, P.G. Biomolecules: where the physics of complexity and simplicity meet. Physics Today;(United States) 1994, 47. [Google Scholar] [CrossRef]
- García, A.E.; Sanbonmatsu, K.Y. α-Helical stabilization by side chain shielding of backbone hydrogen bonds. Proceedings of the National Academy of Sciences 2002, 99, 2782–2787. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Song, L.; Zhou, T.; Zeng, C.; Jia, Y.; Zhao, Y. A computational study of Tat–CDK9–Cyclin binding dynamics and its implication in transcription-dependent HIV latency. Physical Chemistry Chemical Physics 2020, 22, 25474–25482. [Google Scholar] [CrossRef] [PubMed]
- Ning, S.; Zeng, C.; Zeng, C.; Zhao, Y. The TAR binding dynamics and its implication in Tat degradation mechanism. Biophysical Journal 2021, 120, 5158–5168. [Google Scholar] [CrossRef]
- Huang, X.; Zhao, X.; Zheng, F.; Zhan, C.-G. Cocaine esterase–cocaine binding process and the free energy profiles by molecular dynamics and potential of mean force simulations. The Journal of Physical Chemistry B 2012, 116, 3361–3368. [Google Scholar] [CrossRef]
- Wang, K.H.; Penmatsa, A.; Gouaux, E. Neurotransmitter and psychostimulant recognition by the dopamine transporter. Nature 2015, 521, 322–327. [Google Scholar] [CrossRef]
- Coleman, J.A.; Yang, D.; Zhao, Z.; Wen, P.-C.; Yoshioka, C.; Tajkhorshid, E.; Gouaux, E. Serotonin transporter–ibogaine complexes illuminate mechanisms of inhibition and transport. Nature 2019, 569, 141–145. [Google Scholar] [CrossRef]
- Singh, S.K.; Piscitelli, C.L.; Yamashita, A.; Gouaux, E. A competitive inhibitor traps LeuT in an open-to-out conformation. Science 2008, 322, 1655–1661. [Google Scholar] [CrossRef]
- Krishnamurthy, H.; Gouaux, E. X-ray structures of LeuT in substrate-free outward-open and apo inward-open states. Nature 2012, 481, 469–474. [Google Scholar] [CrossRef]
- Polovinkin, L.; Hassaine, G.; Perot, J.; Neumann, E.; Jensen, A.A.; Lefebvre, S.N.; Corringer, P.-J.; Neyton, J.; Chipot, C.; Dehez, F. Conformational transitions of the serotonin 5-HT3 receptor. Nature 2018, 563, 275–279. [Google Scholar] [CrossRef] [PubMed]
- Pozharski, E.; Moulin, A.; Hewagama, A.; Shanafelt, A.B.; Petsko, G.A.; Ringe, D. Diversity in hapten recognition: structural study of an anti-cocaine antibody M82G2. Journal of molecular biology 2005, 349, 570–582. [Google Scholar] [CrossRef] [PubMed]
- Vilar, S.; Cozza, G.; Moro, S. Medicinal chemistry and the molecular operating environment (MOE): application of QSAR and molecular docking to drug discovery. Current topics in medicinal chemistry 2008, 8, 1555–1572. [Google Scholar] [CrossRef] [PubMed]
- Pearlman, D.A.; Case, D.A.; Caldwell, J.W.; Ross, W.S.; Cheatham III, T.E.; DeBolt, S.; Ferguson, D.; Seibel, G.; Kollman, P. AMBER, a package of computer programs for applying molecular mechanics, normal mode analysis, molecular dynamics and free energy calculations to simulate the structural and energetic properties of molecules. Computer Physics Communications 1995, 91, 1–41. [Google Scholar] [CrossRef]
- Case, D.A.; Cheatham III, T.E.; Darden, T.; Gohlke, H.; Luo, R.; Merz Jr, K.M.; Onufriev, A.; Simmerling, C.; Wang, B.; Woods, R.J. The Amber biomolecular simulation programs. Journal of computational chemistry 2005, 26, 1668–1688. [Google Scholar] [CrossRef]
- Frank, M.; Gutbrod, P.; Hassayoun, C.; von Der Lieth, C.-W. Dynamic molecules: molecular dynamics for everyone. An internet-based access to molecular dynamic simulations: basic concepts. Journal of Molecular Modeling 2003, 9, 308–315. [Google Scholar] [CrossRef]
- Karplus, M. Molecular dynamics of biological macromolecules: a brief history and perspective. Biopolymers: Original Research on Biomolecules 2003, 68, 350–358. [Google Scholar] [CrossRef]
- Adcock, S.A.; McCammon, J.A. Molecular dynamics: survey of methods for simulating the activity of proteins. Chemical reviews 2006, 106, 1589–1615. [Google Scholar] [CrossRef]
- Maier, J.A.; Martinez, C.; Kasavajhala, K.; Wickstrom, L.; Hauser, K.E.; Simmerling, C. ff14SB: improving the accuracy of protein side chain and backbone parameters from ff99SB. Journal of chemical theory and computation 2015, 11, 3696–3713. [Google Scholar] [CrossRef] [PubMed]
- Dickson, C.J.; Madej, B.D.; Skjevik, Å.A.; Betz, R.M.; Teigen, K.; Gould, I.R.; Walker, R.C. Lipid14: the amber lipid force field. Journal of chemical theory and computation 2014, 10, 865–879. [Google Scholar] [CrossRef] [PubMed]
- He, X.; Man, V.H.; Yang, W.; Lee, T.-S.; Wang, J. A fast and high-quality charge model for the next generation general AMBER force field. The Journal of Chemical Physics 2020, 153, 114502. [Google Scholar] [CrossRef]
- Lu, T.; Chen, F. Multiwfn: a multifunctional wavefunction analyzer. Journal of computational chemistry 2012, 33, 580–592. [Google Scholar] [CrossRef]
- Frisch, M.; Trucks, G.; Schlegel, H.; Scuseria, G.; Robb, M.; Cheeseman, J.; Scalmani, G.; Barone, V.; Mennucci, B.; Petersson, G. Gaussian 09, rev. Gaussian Inc, Wallingford 2009.
- Zhang, J.; Lu, T. Efficient evaluation of electrostatic potential with computerized optimized code. Physical Chemistry Chemical Physics 2021, 23, 20323–20328. [Google Scholar] [CrossRef]
- Morishita, T. Fluctuation formulas in molecular-dynamics simulations with the weak coupling heat bath. The Journal of Chemical Physics 2000, 113, 2976–2982. [Google Scholar] [CrossRef]
- Toukmaji, A.; Sagui, C.; Board, J.; Darden, T. Efficient particle-mesh Ewald based approach to fixed and induced dipolar interactions. The Journal of chemical physics 2000, 113, 10913–10927. [Google Scholar] [CrossRef]
- Ryckaert, J.-P.; Ciccotti, G.; Berendsen, H.J. Numerical integration of the cartesian equations of motion of a system with constraints: molecular dynamics of n-alkanes. Journal of computational physics 1977, 23, 327–341. [Google Scholar] [CrossRef]
- Berendsen, H.J.; Postma, J.v.; Van Gunsteren, W.F.; DiNola, A.; Haak, J.R. Molecular dynamics with coupling to an external bath. The Journal of chemical physics 1984, 81, 3684–3690. [Google Scholar] [CrossRef]
- Kollman, P.A.; Massova, I.; Reyes, C.; Kuhn, B.; Huo, S.; Chong, L.; Lee, M.; Lee, T.; Duan, Y.; Wang, W. Calculating structures and free energies of complex molecules: combining molecular mechanics and continuum models. Accounts of chemical research 2000, 33, 889–897. [Google Scholar] [CrossRef] [PubMed]
- Fu, T.; Zheng, G.; Tu, G.; Yang, F.; Chen, Y.; Yao, X.; Li, X.; Xue, W.; Zhu, F. Exploring the binding mechanism of metabotropic glutamate receptor 5 negative allosteric modulators in clinical trials by molecular dynamics simulations. ACS chemical neuroscience 2018, 9, 1492–1502. [Google Scholar] [CrossRef] [PubMed]
- Xue, W.; Wang, P.; Li, B.; Li, Y.; Xu, X.; Yang, F.; Yao, X.; Chen, Y.Z.; Xu, F.; Zhu, F. Identification of the inhibitory mechanism of FDA approved selective serotonin reuptake inhibitors: an insight from molecular dynamics simulation study. Physical Chemistry Chemical Physics 2016, 18, 3260–3271. [Google Scholar] [CrossRef] [PubMed]
- Xue, W.; Wang, P.; Tu, G.; Yang, F.; Zheng, G.; Li, X.; Li, X.; Chen, Y.; Yao, X.; Zhu, F. Computational identification of the binding mechanism of a triple reuptake inhibitor amitifadine for the treatment of major depressive disorder. Physical Chemistry Chemical Physics 2018, 20, 6606–6616. [Google Scholar] [CrossRef] [PubMed]
- Zheng, G.; Xue, W.; Wang, P.; Yang, F.; Li, B.; Li, X.; Li, Y.; Yao, X.; Zhu, F. Exploring the inhibitory mechanism of approved selective norepinephrine reuptake inhibitors and reboxetine enantiomers by molecular dynamics study. Scientific reports 2016, 6, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Fu, T.; Zhang, X.; Yang, F.; Zheng, G.; Xue, W.; Chen, Y.; Yao, X.; Zhu, F. Differentiating physicochemical properties between NDRIs and sNRIs clinically important for the treatment of ADHD. Biochimica et Biophysica Acta (BBA)-General Subjects 2017, 1861, 2766–2777. [Google Scholar] [CrossRef] [PubMed]
- Torrie, G.M.; Valleau, J.P. Nonphysical sampling distributions in Monte Carlo free-energy estimation: Umbrella sampling. Journal of Computational Physics 1977, 23, 187–199. [Google Scholar] [CrossRef]
- Kirkwood, J.G. Statistical mechanics of fluid mixtures. The Journal of chemical physics 1935, 3, 300–313. [Google Scholar] [CrossRef]
- Berneche, S.; Roux, B. Energetics of ion conduction through the K+ channel. Nature 2001, 414, 73–77. [Google Scholar] [CrossRef]
- Roux, B. The calculation of the potential of mean force using computer simulations. Computer physics communications 1995, 91, 275–282. [Google Scholar] [CrossRef]
| substrate/drug | target | ΔGcale (Kcal/mol)a | ΔGexp(Kcal/mol)b | Ki(nM)b |
|---|---|---|---|---|
| serotonin | SERT (S1)c | -28.38 | -6.62 | 14.2 μM[16] |
| SERT (S2)d | -18.88 | -5.90 | 48 μM | |
| cocaine | SERT (S1)c | -46.42 | f | |
| SERT (S2)d | -39.60 | |||
| escitalopram | SERT (S1)c | -51.47 | -10.23 | 32e[15] |
| SERT (S2)d | -38.76 | -7.16 | 5800 (IC50) | |
| paroxetine | SERT (S1)c | -51.34 | -10.40 | 24 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




