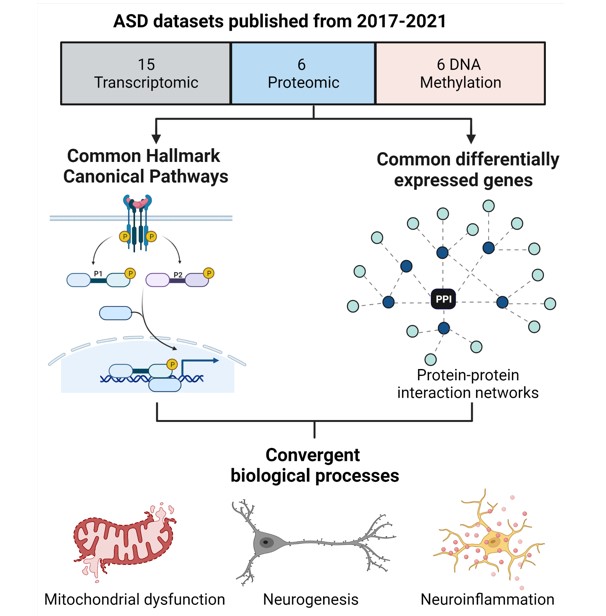
Abstract: Autism Spectrum Disorder (ASD) is a complex neurodevelopmental disorder with extensive genetic and aetiological heterogeneity. While the underlying molecular mechanisms involved remain unclear, significant progress has been facilitated by recent advances in high-throughput transcriptomic, epigenomic and proteomic technologies. Here, we review recently published ASD proteomic data and compare proteomic func-tional enrichment signatures to those of transcriptomic and epigenomic data. We iden-tify canonical pathways that are consistently implicated in ASD molecular data and find an enrichment of pathways involved in mitochondrial metabolism and neurogenesis. We identify a subset of differentially expressed proteins that are supported by ASD tran-scriptomic and DNA methylation data. Furthermore, these differentially expressed proteins are enriched for disease phenotype pathways associated with ASD aetiology. These proteins converge on protein-protein interaction networks that regulate cell pro-liferation and differentiation, metabolism and inflammation which demonstrates a link between canonical pathways, biological processes and the ASD phenotype. This review highlights how proteomics can uncover potential molecular mechanisms to explain a link between mitochondrial dysfunction and neurodevelopmental pathology.