Submitted:
05 February 2025
Posted:
05 February 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
1.1. MRI, Cartesian and Non-Cartesian
1.2. Cartesian k-Space MRI Reconstruction Methods
1.3. Non-Cartesian k-Space MRI Reconstruction Methods
1.4. The Rosette Trajectory
2. Materials and Methods
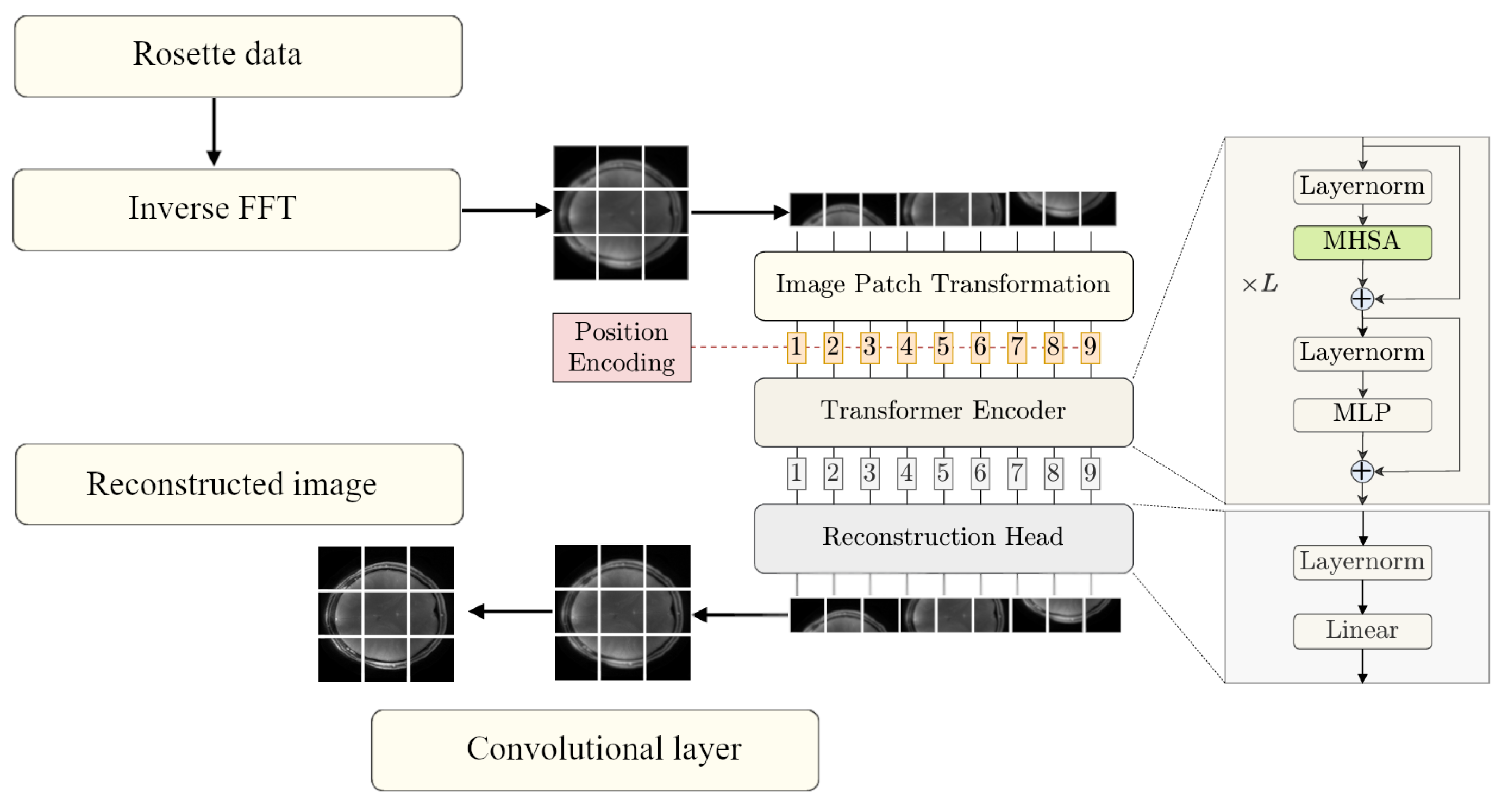
2.1. Method Overview
2.2. Vision Transformer
2.3. Dataset and Pre-Processing
- Repetition Time (TR): 2.4 seconds
- Echo Time (TE) (dual): 1 and 9 milliseconds:
- Acceleration factor: 4
- Number of petals: 189
- Nominal in-plane resolution: 0.468 millimeters
- Slice thickness: 2 millimeters
- Flip angle: 7 degrees
- Number of slices: 1
- Random Horizontal Flip, probability=0.5
- Random Vertical Flip, probability=0.5
- Random Rotation, 0 to 180 degrees
- Color Jitter, brightness/contrast/saturation, range= 0.8 to 1.2
- Random Resized Crop, scale= 0.3 to 1.1
2.4. Evaluation Methods
- The Structural Similarity Index Measure (SSIM) measures image similarity between a reference image and a processed image. Higher scores are preferred.
- Normalized Root Mean Square Error (NRMSE) is the root mean squared error between the images normalized by the sum of the observed values. Lower error is preferred.
- Normalized Mutual Information (NMI) is a normalized Mutual Information (MI) score, where the scale between no mutual information and full correlation is given as 0 to 1.
- Relative contrast is the ratio between the difference in maximum and minimum intensity and the sum of the same values.
- Peak Signal-to-Noise Ratio (PSNR) measures the ratio between the maximum possible pixel value and the noise power. Higher PSNR values indicate better image quality.
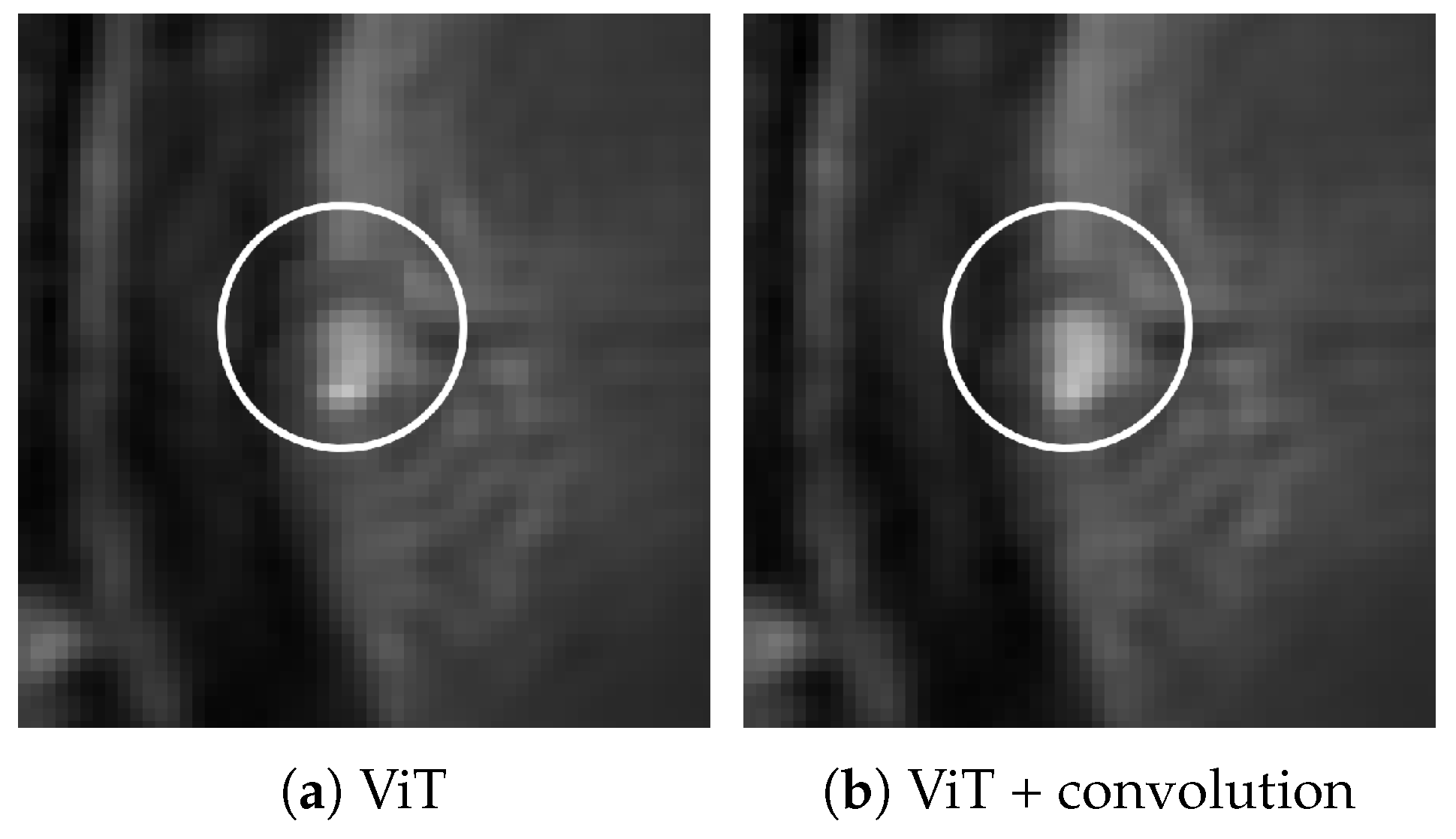
2.5. Visualization
2.6. Training Procedure
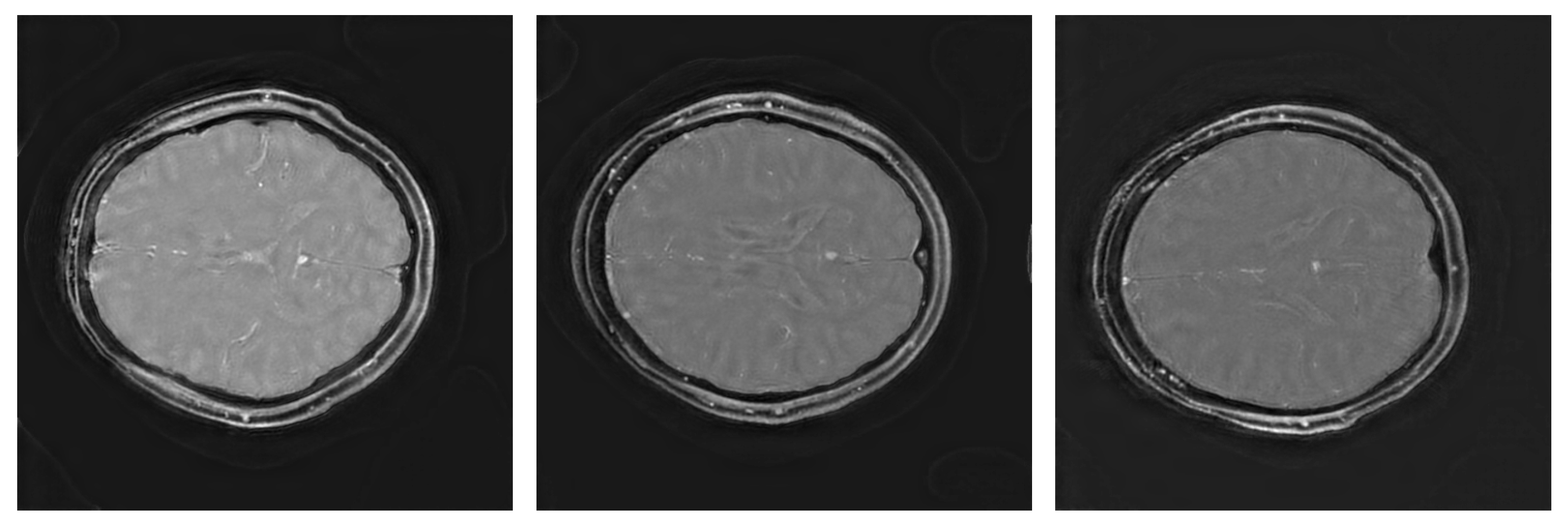
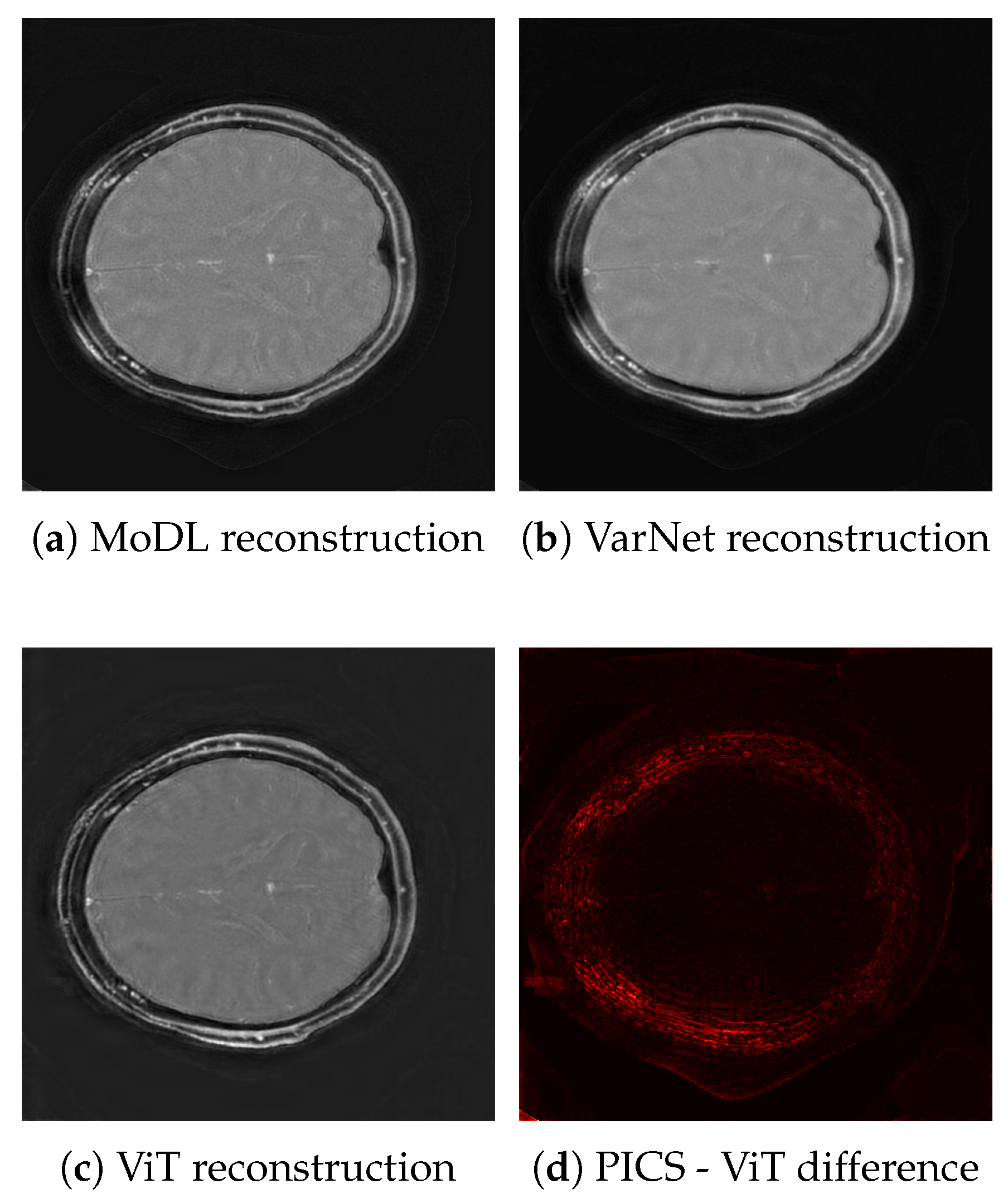
3. Results
3.1. Image Scores
3.2. Network Runtime Performance
4. Discussion
Author Contributions
Data Availability Statement
Acknowledgments
References
- Geethanath, S.; Vaughan Jr, J.T. Accessible magnetic resonance imaging: A review. Journal of Magnetic Resonance Imaging 2019, 49, e65–e77. [Google Scholar] [CrossRef] [PubMed]
- Moratal, D.; Vallés-Luch, A.; Martí-Bonmatí, L.; Brummer, M.E. k-Space tutorial: an MRI educational tool for a better understanding of k-space. Biomedical imaging and intervention journal 2008, 4. [Google Scholar] [CrossRef] [PubMed]
- Gallagher, T.A.; Nemeth, A.J.; Hacein-Bey, L. An introduction to the Fourier transform: relationship to MRI. American journal of roentgenology 2008, 190, 1396–1405. [Google Scholar] [CrossRef] [PubMed]
- Hollingsworth, K.G. Reducing acquisition time in clinical MRI by data undersampling and compressed sensing reconstruction. Physics in Medicine & Biology 2015, 60, R297. [Google Scholar]
- Wright, K.L.; Hamilton, J.I.; Griswold, M.A.; Gulani, V.; Seiberlich, N. Non-Cartesian parallel imaging reconstruction. Journal of Magnetic Resonance Imaging 2014, 40, 1022–1040. [Google Scholar] [CrossRef]
- Lin, D.J.; Johnson, P.M.; Knoll, F.; Lui, Y.W. Artificial Intelligence for MR Image Reconstruction: An Overview for Clinicians. Journal of Magnetic Resonance Imaging 2021, 53, 1015–1028. [Google Scholar] [CrossRef]
- Geethanath, S.; Reddy, R.; Konar, A.S.; Imam, S.; Sundaresan, R.; R. , R.B.D.; Venkatesan, R. Compressed Sensing MRI: A Review. Critical Reviews™ in Biomedical Engineering 2013, 41, 183–204. [Google Scholar]
- Ye, J.C. Compressed sensing MRI: a review from signal processing perspective. BMC Biomedical Engineering 2019, 1, 8. [Google Scholar] [CrossRef]
- Pal, A.; Rathi, Y. A review and experimental evaluation of deep learning methods for MRI reconstruction. The journal of machine learning for biomedical imaging 2022, 1. [Google Scholar] [CrossRef]
- Zhang, H.M.; Dong, B. A review on deep learning in medical image reconstruction. Journal of the Operations Research Society of China 2020, 8, 311–340. [Google Scholar] [CrossRef]
- Chen, Y.; Schönlieb, C.B.; Liò, P.; Leiner, T.; Dragotti, P.L.; Wang, G.; Rueckert, D.; Firmin, D.; Yang, G. AI-based reconstruction for fast MRI—A systematic review and meta-analysis. Proceedings of the IEEE 2022, 110, 224–245. [Google Scholar] [CrossRef]
- Liang, D.; Cheng, J.; Ke, Z.; Ying, L. Deep magnetic resonance image reconstruction: Inverse problems meet neural networks. IEEE Signal Processing Magazine 2020, 37, 141–151. [Google Scholar] [CrossRef] [PubMed]
- Chandra, S.S.; Bran Lorenzana, M.; Liu, X.; Liu, S.; Bollmann, S.; Crozier, S. Deep learning in magnetic resonance image reconstruction. Journal of Medical Imaging and Radiation Oncology 2021, 65, 564–577. [Google Scholar] [CrossRef] [PubMed]
- Montalt-Tordera, J.; Muthurangu, V.; Hauptmann, A.; Steeden, J.A. Machine learning in magnetic resonance imaging: image reconstruction. Physica Medica 2021, 83, 79–87. [Google Scholar] [CrossRef]
- Hammernik, K.; Klatzer, T.; Kobler, E.; Recht, M.P.; Sodickson, D.K.; Pock, T.; Knoll, F. Learning a variational network for reconstruction of accelerated MRI data. Magnetic resonance in medicine 2018, 79, 3055–3071. [Google Scholar] [CrossRef]
- Lv, J.; Zhu, J.; Yang, G. Which GAN? A comparative study of generative adversarial network-based fast MRI reconstruction. Philosophical Transactions of the Royal Society A 2021, 379, 20200203. [Google Scholar] [CrossRef]
- Zhou, B.; Schlemper, J.; Dey, N.; Salehi, S.S.M.; Sheth, K.; Liu, C.; Duncan, J.S.; Sofka, M. Dual-domain self-supervised learning for accelerated non-Cartesian MRI reconstruction. Medical Image Analysis 2022, 81, 102538. [Google Scholar] [CrossRef]
- Fessler, J.A. On NUFFT-based gridding for non-Cartesian MRI. Journal of magnetic resonance 2007, 188, 191–195. [Google Scholar] [CrossRef]
- Wang, S.; Xiao, T.; Liu, Q.; Zheng, H. Deep learning for fast MR imaging: A review for learning reconstruction from incomplete k-space data. Biomedical Signal Processing and Control 2021, 68, 102579. [Google Scholar] [CrossRef]
- Sriram, A.; Zbontar, J.; Murrell, T.; Defazio, A.; Zitnick, C.L.; Yakubova, N.; Knoll, F.; Johnson, P. End-to-end variational networks for accelerated MRI reconstruction. In Proceedings of the Medical Image Computing and Computer Assisted Intervention–MICCAI 2020: 23rd International Conference, Lima, Peru, 2020, Proceedings, Part II 23. Springer, 2020, October 4–8; pp. 64–73.
- Aggarwal, H.K.; Mani, M.P.; Jacob, M. MoDL: Model-based deep learning architecture for inverse problems. IEEE transactions on medical imaging 2018, 38, 394–405. [Google Scholar] [CrossRef]
- Lin, K.; Heckel, R. Vision Transformers Enable Fast and Robust Accelerated MRI. In Proceedings of the Proceedings of The 5th International Conference on Medical Imaging with Deep Learning; Konukoglu, E.; Menze, B.; Venkataraman, A.; Baumgartner, C.; Dou, Q.; Albarqouni, S., Eds. PMLR, 06–08 Jul 2022, Vol. 172, Proceedings of Machine Learning Research, pp.
- Parvaiz, A.; Khalid, M.A.; Zafar, R.; Ameer, H.; Ali, M.; Fraz, M.M. Vision Transformers in medical computer vision—A contemplative retrospection. Engineering Applications of Artificial Intelligence 2023, 122, 106126. [Google Scholar] [CrossRef]
- Chen, Z.; Chen, Y.; Xie, Y.; Li, D.; Christodoulou, A.G. Data-consistent non-Cartesian deep subspace learning for efficient dynamic MR image reconstruction. In Proceedings of the 2022 IEEE 19th International Symposium on Biomedical Imaging (ISBI). IEEE; 2022; pp. 1–5. [Google Scholar]
- Singh, D.; Monga, A.; de Moura, H.L.; Zhang, X.; Zibetti, M.V.; Regatte, R.R. Emerging trends in fast MRI using deep-learning reconstruction on undersampled k-space data: a systematic review. Bioengineering 2023, 10, 1012. [Google Scholar] [CrossRef]
- Shen, X.; Özen, A.C.; Sunjar, A.; Ilbey, S.; Sawiak, S.; Shi, R.; Chiew, M.; Emir, U. Ultra-short T2 components imaging of the whole brain using 3D dual-echo UTE MRI with rosette k-space pattern. Magnetic resonance in medicine 2023, 89, 508–521. [Google Scholar] [CrossRef]
- Li, Y.; Yang, R.; Zhang, C.; Zhang, J.; Jia, S.; Zhou, Z. Analysis of generalized rosette trajectory for compressed sensing MRI. Medical physics 2015, 42, 5530–5544. [Google Scholar] [CrossRef]
- Bucholz, E.K.; Song, J.; Johnson, G.A.; Hancu, I. Multispectral imaging with three-dimensional rosette trajectories. Magnetic Resonance in Medicine: An Official Journal of the International Society for Magnetic Resonance in Medicine 2008, 59, 581–589. [Google Scholar] [CrossRef]
- Shen, X.; Özen, A.C.; Sunjar, A.; Ilbey, S.; Shi, R.; Chiew, M.; Emir, U. Myelin Imaging Using Dual-echo 3D Ultra-short Echo Time MRI with Rosette k-Space Pattern. bioRxiv, 2021. [Google Scholar]
- Mahmud, S.Z.; Denney, T.S.; Bashir, A. Feasibility of spinal cord imaging at 7 T using rosette trajectory with magnetization transfer preparation and compressed sensing. Scientific Reports 2023, 13, 8777. [Google Scholar] [CrossRef]
- Alcicek, S.; Craig-Craven, A.R.; Shen, X.; Chiew, M.; Ozen, A.; Sawiak, S.; Pilatus, U.; emir, u. Multi-site ultrashort echo time 3D phosphorous MRSI repeatability using novel rosette trajectory (PETALUTE). bioRxiv, 2024. [Google Scholar]
- Dosovitskiy, A.; Beyer, L.; Kolesnikov, A.; Weissenborn, D.; Zhai, X.; Unterthiner, T.; Dehghani, M.; Minderer, M.; Heigold, G.; Gelly, S.; et al. An image is worth 16x16 words: Transformers for image recognition at scale. arXiv preprint arXiv:2010.11929, arXiv:2010.11929 2020.
- Zhang, C.; Jiang, W.; Zhang, Y.; Wang, W.; Zhao, Q.; Wang, C. Transformer and CNN hybrid deep neural network for semantic segmentation of very-high-resolution remote sensing imagery. IEEE Transactions on Geoscience and Remote Sensing 2022, 60, 1–20. [Google Scholar] [CrossRef]
- Blumenthal, M.; Luo, G.; Schilling, M.; Holme, H.C.M.; Uecker, M. Deep, deep learning with BART. Magnetic resonance in medicine 2023, 89, 678–693. [Google Scholar] [CrossRef]
- Kingma, D.P.; Ba, J. Adam: A method for stochastic optimization. arXiv preprint arXiv:1412.6980, arXiv:1412.6980 2014.
- Smith, L.N.; Topin, N. Super-convergence: Very fast training of neural networks using large learning rates. In Proceedings of the Artificial intelligence and machine learning for multi-domain operations applications. SPIE, Vol. 11006; 2019; pp. 369–386. [Google Scholar]
| Technique | Advantages | Disadvantages |
|---|---|---|
| IFFT | - Robust initial approximation from k-space to image domain. - Computationally efficient for uniformly sampled Cartesian data. | - Struggles with non-Cartesian trajectories. - Susceptible to artifacts due to irregular and undersampled data. |
| CS | - Balances fidelity to acquired data with image sparsity. - Effective for undersampled data, reducing scan times. - Works well with Cartesian and non-Cartesian data. | - Requires complex optimization, leading to higher computational costs. - Sensitive to parameter tuning and model assumptions. |
| VarNet, MoDL | - Learns complex mappings from data, improving reconstruction at high acceleration rates.
- Handles both Cartesian and non-Cartesian data without regridding. |
- Requires large datasets and computational resources for training.
- Long inference times |
| ViT | - Models long-range dependencies, capturing complex spatial patterns. - No need for extensive pre-processing of non-Cartesian data. | - Requires augmented training data, increasing training time and resource consumption. |
| Metric | Formula |
|---|---|
| SSIM |
|
| NRMSE |
|
| NMI |
|
| Relative Contrast |
|
| PSNR |
|
| Reconstruction Method | SSIM ↑ | NRMSE ↓ | PSNR ↑ | NMI ↑ | Relative Contrast ↑ |
|---|---|---|---|---|---|
| VarNet | 0.946 | 0.250 | 23.63 | 0.537 | 0.277 |
| MoDL | 0.987 | 0.059 | 38.21 | 0.575 | 0.327 |
| Non-augmented ViT | 0.982 | 0.040 | 42.07 | 0.438 | 0.296 |
| Augmented ViT (X1) | 0.983 | 0.029 | 44.98 | 0.458 | 0.306 |
| Augmented ViT (X3) | 0.987 | 0.026 | 46.11 | 0.493 | 0.302 |
| Network | Total CPU Time (minutes) | Max GPU Memory Used (MB) |
| VarNet (10 images) | 06:58 | 2785 |
| VarNet (20 images) | 13:51 | |
| MoDL (10 images) | 07:05 | 6369 |
| MoDL (20 images) | 14:08 | |
| ViT (10 images) | 00:45 | 4895 |
| ViT (20 images) | 01:25 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




