1. Introduction
Chest X-ray radiography (CXR) is usually the first imaging technique employed to perform an early diagnosis of cardiopulmonary diseases. Typical pathologies detected in CXRs include pneumonia, atelectasis, consolidation, pneumothorax or pleural and pericardial effusion [
1]. Since the COVID-19 pandemic started in 2020, it is also used as a tool to detect and assess the evolution of the pneumonia caused by this condition [
2]. CXR is used worldwide thanks to the simplicity with which it can be completed, the low cost, low radiation dose, and its sensitivity [
3,
4].
Regarding the position and orientation of the patient in relation to the X-ray source, the desired and most frequently used setup is the posteroanterior (PA) projection [
5], since it provides better visualization of lungs [
6]. However, this configuration requires the patient to be standing erect, which is not possible for critically ill patients, intubated patients or some elderly people [
6,
7]. In these situations, it is more convenient to carry out the anteroposterior (AP) projection, in which the patient can be sitting up in bed or lying in supine position. Thus, AP image can be acquired with a portable X-ray unit outside the radiology department too, when the patient’s condition advises not to shift him/her [
6].
X-ray imaging is based on the attenuation that photons suffer when they traverse the human body, so that photons that cross the body and reach the detector without interacting with the media, i.e. primary photons, form the image [
8]. However, there are also photons that interact with the media and are not absorbed, but they are scattered. This secondary radiation can also get to the detector, representing background noise and causing blurring in the primary image and a considerable loss in contrast-to-noise ratio (CNR) [
9]. This effect is especially relevant in CXR due to the requirement of large detectors (at least 35 cm side) to be able to cover the whole region of interest (ROI) [
10]. To overcome this problem, several techniques have been proposed, which can be split into two types: scatter suppression and scatter estimation [
11]. Scatter suppression methods aim to remove, or at least reduce, the scattered photons that arrive at the detector, while scatter estimation methods try to obtain the signal just of the secondary photons, and then subtract it from the total projection.
The most widely used scatter correction method is the anti-scatter grid, where a grid is interposed between the patient and the detector in a way that scattered photons are absorbed while primary photons are allowed to pass [
7,
12]. This technique is successfully implemented in clinical practice [
10] since many clinical systems incorporate this grid [
13], although it has some disadvantages. Firstly, an anti-scatter grid can also attenuate primary radiation leading to noisy images [
9,
10]. Besides, it can generate grid line artifacts in the image [
14] and it can increase the radiation exposure between two and five times [
5,
15]. Finally, in AP acquisitions it is difficult to accurately align the grid with respect to the beam, so the use of anti-scatter grid in combination with portable X-ray units used at intensive care units or with patients that cannot stand up is more time-consuming and does not guarantee a good result [
7,
13]. Another method studied to physically remove the scattered X-rays consists in increasing the air gap between the patient and the detector, which enlarges the probability that secondary photons miss the detector due to large scatter angles [
16,
17]. This approach causes a smaller increase in the radiation dose than the anti-scatter grid, but it can magnify the acquired image [
5]. Other alternatives to the use of the anti-scatter grid are the slit scanning, with the drawback of an increase of acquisition times, or a strict collimation, which compromises the adaptability of the imaging equipment [
10,
11,
13].
Regarding scatter estimation, there are some experimental methods such as the beam stop array (BSA), which acquires a projection of just the scatter radiation (primary photons are removed) and then it is subtracted from the standard projection [
18]. However, most techniques to estimate the scatter radiation are software-based [
19,
20,
21]. In this field, Monte Carlo (MC) simulations are gold-standard. MC codes reproduce in a realistic and very accurate way the interactions of photons (photoelectric absorption and Rayleigh and Compton scattering) in their path through the human body. Therefore, these models are able to provide precise estimations of scatter [
9,
13,
19]. The major drawback of MC simulations is the high computational time they require, which makes it difficult to implement in real-time clinical practice [
9,
19]. Model-based methods make use of simpler physical models, so they are faster than MC at the cost of much less accuracy [
13]. Similarly, kernel-based models approximate the scatter signal by an integral transform of a scatter source term multiplied by a scatter propagation kernel [
11,
21]. Nevertheless, this method depends on each acquisition setup (geometry, image object or X-ray beam spectrum), so it is not easy to generalize [
19].
Recently, deep learning algorithms have been widely used for several medical imaging analysis tasks, like object localization [
22,
23], object classification [
24,
25] or image segmentation [
26,
27]. Particularly, convolutional neural networks (CNN) have succeeded in image processing, outperforming traditional and state of the art techniques [
3,
26,
28]. Among them, the U-Net, first proposed by Ronneberger et al in 2015 [
26] for biomedical image purposes, and its subsequent variants such as the MultiResUNet are the most popular networks [
28]. In particular, there are some works that make use of CNN to estimate scatter either in CXR or computed tomography (CT). In Maier et al [
19], they used a U-Net like deep CNN to estimate the scatter in CT images, training the network with MC simulations. In a similar way, Lee & Lee [
9] used MC simulations to train a CNN for image restoration, that is, the scatter image is estimated, and then it is subtracted from the CNN input image (the scattered image), obtaining as output the scatter-corrected image. In that case, they applied the CNN to CXRs, which are then used to reconstruct the corresponding cone-beam CT image. Roser et al [
13] also used a U-Net in combination with prior knowledge of the X-ray scattering, which is approximated by bivariate B-splines, so that spline coefficients help to calculate the scatter image for head and thorax datasets.
Dual-energy X-ray imaging was first proposed by Alvarez and Macovski in 1976 [
29]. This technique takes advantage of the dependence of the attenuation coefficient with the energy of the photon beam and the material properties, i.e. its mass density and atomic number. In dual-energy X-ray, two projections using two different energy spectra in the range of 20-200 kV are acquired. They can provide extra information with respect to a common single-energy radiography, allowing to better distinguish different materials [
30,
31,
32]. In this way, dual-energy X-ray imaging may improve the diagnosis of oncological, vascular or osseous pathologies [
30,
33]. Specifically, in CXR dual acquisitions are utilized to perform dual energy subtraction (DES), generating two separate images: one of the soft tissue and one of the bone. Soft tissue selective images, in which ribs and clavicle shadows are removed, have been proven to enhance the detection of nodules and pulmonary diseases [
34,
35].
Several studies have made use of dual-energy X-ray absorption images along with deep-learning methods to enhance medical image analysis. Some examples include image segmentation of bones to diagnose and quantify osteoporosis [
36], or estimation of phase contrast signal from X-ray images acquired with two energy spectra [
37]. Regarding chest X-rays, the combination of dual energy and deep learning has been used to obtain two separated images with bone and soft tissue components [
38]. However, to the best of our knowledge, the application of dual-energy images to obtain the scatter signal has not been yet examined.
In this work, we present a study of robustness of three U-Net-like CNN that estimate and correct the scatter contribution using either single-energy CXRs or dual-energy CXRs to perform the network training. All images were simulated with a Monte Carlo code from actual CTs of patients affected with COVID-19. The scatter-corrected CXRs were obtained by subtracting the estimation of the scatter contribution from the image affected by the scattered rays, which we call uncorrected scatter image. Two different analysis were performed to evaluate the robustness of the models. First, several metrics were calculated to assess the accuracy of the scatter correction, taking the MC simulations as the ground truth, after the algorithms were applied to images simulated with various source-to-detector distances (SDD), including the original SDD with which the training images were simulated. The accuracy of the models with CXRs acquired with the original SDD was taken as reference, and it was compared with the values obtained for the other distances. Secondly, the contrast between the area of the lesion (COVID-19) and the healthy area of the lung was evaluated on soft tissue Dual-Energy Subtraction (DES) images to quantify how scatter removal helps to better identify the affected region. Then, values of contrast in the ground truth were compared with the ones obtained in the estimated scatter-corrected images, just with the original SDD. Finally, the single-energy neural network was tested with a cohort of varied real CXRs, and the ratio between the values of two regions in the lung (with and without ribs) was calculated to determine how the contrast of the image improves after applying the scatter correction method.
4. Discussion
In this work, we have implemented and evaluated the performance of three deep learning models that estimate and correct the scatter in CXRs. One model is based on standard single energy acquisitions, while the other two models assume that two CXRs with different energies (dual energy) were acquired per patient. The impact of varying the distance of the X-ray source on the accuracy of the scatter correction with these methods was studied.
Results in
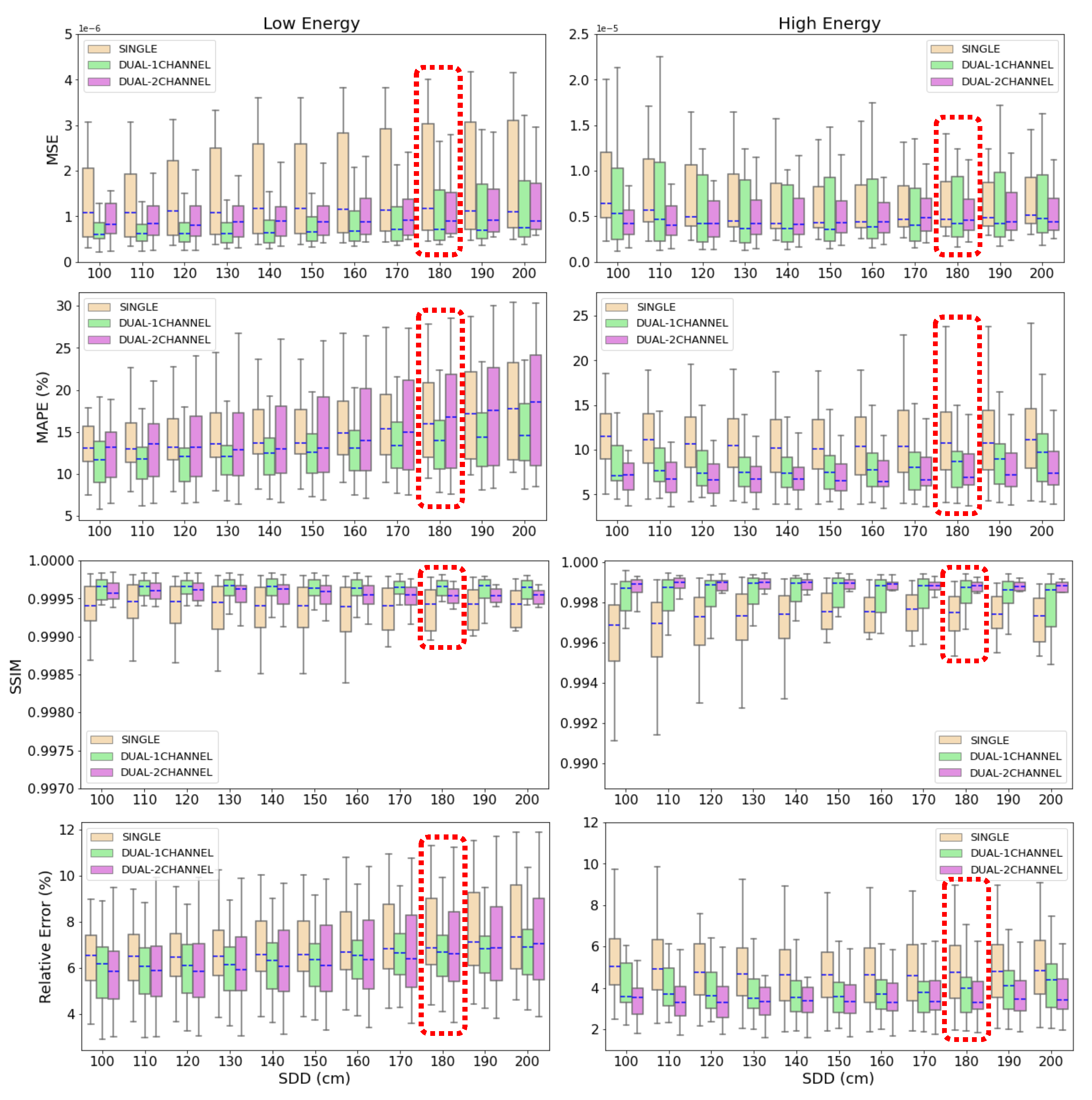
Figure 8 demonstrate that the three deep learning-based models are able to accurately estimate and correct scatter contribution to simulated CXRs. Moreover, the three algorithms keep their accuracy on scatter correction when the source-to-detector distance (SDD) is varied in a range of 100 cm from the SDD used to train the neural networks. Thus, the results prove that the models are robust to variations on the setup parameters like air gap or SDD and their application is not limited to a specific configuration.
The highest decrease in accuracy is found in SDD=100 cm and SDD=200 cm (the neural networks were trained with SDD=180 cm). For SDD=100 cm, this result could be expected since it is the case with the largest distance with respect to the reference SDD. The fact that less accurate metrics are obtained for SDD=200 cm, which is the second closest distance to the one of reference, indicates that the models are more accurate if the images are well focused on the lungs and the air-background areas are reduced or cropped. In this case, lungs get smaller in the image as the SDD increases, which explains the lost in accuracy.
According to the mean values of MAPE and relative error shown in
Section 3.1, the results obtained with the dual-energy NN models are more accurate than the ones provided by the single-energy model. In the estimations of the scatter-corrected high energy CXRs, the p-value between the metrics yielded by the 2-output dual-energy and the single-energy models is
for the MAPE and
for the relative error. That is, in both metrics the p-value is below the classic threshold of 0.05 [
54,
55], indicating that the difference is statistically significant. In the low energy estimations, the p-value between the 1-output dual-energy and the single-energy algorithms is
for the MAPE and
for the relative error, so the difference is significant only in the first metric. As it was explained in
Section 3.1, the difference in the SSIM is minor. Regarding the MSE, there is not a substantial difference between the three models due to the fact that this is the metric employed as loss function in the training of the algorithms, so it is minimized in both cases. In the comparison between the two proposed models of scatter correction, it is also observed in
Figure 8 that there is less deviation in the values of MAPE and relative error applying the dual-energy model. In the high energy case, the standard deviation in MAPE is 4.9%, 3.4% and 2.6% for single-energy, 1-output and 2-output dual-energy models, respectively, and the deviation in the relative error is 1.8% for the single-energy network, 1.4% for the 1-output dual-energy algorithm and 1.1% for the 2-output-dual-energy algorithm. In the low energy case, the standard deviation in MAPE is 6.0%, 4.1% and 6.3% for the single-energy, 1-output and 2-output dual-energy models, and the corresponding values in the relative error are 2.0%, 1.5% and 2.2%, respectively. All these results point out the superior performance of the dual-energy models for scatter correction of CXRs, being more accurate the 1-output model for low energy acquisitions, and the 2-output model for high energy images.
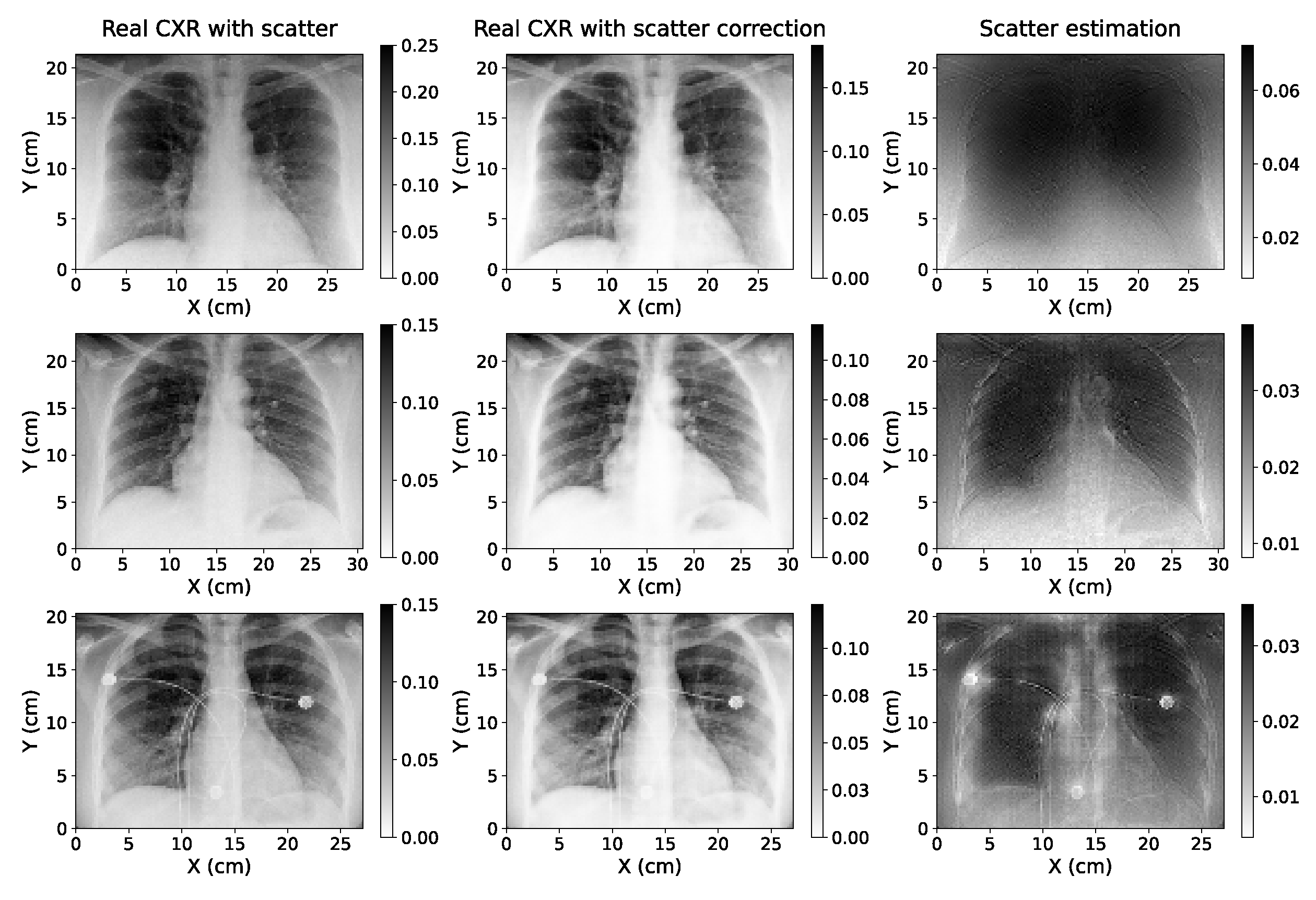
The application of the single-energy model to real CXRs acquired without an anti-scatter grid suggests that the algorithm can be easily adapted to be used with true data acquired with different setups, since in this case the information of the parameters with which the projections were taken was not available in the anonymized Dicom headers used. Furthermore, the accuracy of the model applied to these images could not be quantitatively determined as there was no access to ground truth scatter-free CXRs. However,
Figure 10 shows that the estimations are robust, even for a radiography with artifacts such as wires, as it happens in the third case of this figure. This is a key aspect since the application of these models can be specially useful for critically ill patients where it is harder to utilize the anti-scatter grid (as it was explained in
Section 1). In addition, the values of the ratio between the regions of the lung with and without ribs (
Table 2) point out that the scatter correction improves the contrast between different tissues.
The two dual-energy models could not be verified against real data since we currently do not have access to CXRs acquired with two different X-ray kilovoltages. Nevertheless, it is expected that good estimations of scatter-corrected images can be obtained in actual images, in light of the results shown in
Section 3.1 and
Section 3.3. Moreover, the superior accuracy shown by the 1-output and 2-output dual-energy models when evaluating the MAPE and the relative error, which has been previously discussed in this section and in
Section 3.1, would be worthy to be studied in a future research with real CXRs acquired without the anti-scatter grid with two different energy spectra. This way, it could be determined if dual-energy approaches truly provide better scatter correction, and it also could be tested the gain in DES images after scatter correction.
In the works of Lee & Lee [
9], they perform a similar study on CXR scatter correction using CNN and Monte Carlo simulations to generate training cases. They evaluated the SSIM, among other metrics, obtaining an average value of 0.992. This is in the same order of the 0.997 value yielded by the models herein proposed, being our SSIM a 0.5% better. As stated by these authors [
9], the application of deep learning to scatter correction of CXR has recently started, so there are not many studies with which results can be compared. Some literature can be found on scatter correction (or estimation) on cone beam CT images. Roser et al. [
13] presented a scatter estimation method based on a deep-learning approach helped by bivariate B-spline approximation. For thorax CT, the MAPE ranges from 3% to 20% approximately for the five fold cross-validations, while our models get average values of 11.2% and 8.6%. Additionally, the SSIM for the thorax CT study is between 0.96 and 0.99. Although CT and CXR have some obvious differences and they are not exactly comparable, it can be noticed that our models present a similar precision.
The models displayed in this paper still have some limitations that would need further study. First, we uniquely employed simulated images for the training and validation, as well as for the test cases in
Section 3.1. Thus, when these NN are applied to real chest X-ray acquisitions, the accuracy of the scatter correction depends on the precision of the Monte Carlo codes. Despite the fact that MC simulations are gold-standard in the field and provide very realistic images, some works for scatter estimation in CT showed that the accuracy of a deep-learning model could decrease when applying a NN trained only with simulated images to a real image [
19]. For CXRs, more studies are needed to determine the accuracy of neural networks in this situation. Since having a sufficient amount of cases to train only with real data is complex in this field, it would be convenient to get images as realistic as possible. For this purpose, if at least some real cases are available, the use of generative adversarial networks (GANs) [
56] in combination with MC simulations could achieve this goal and thus improve the accuracy of the models applied to real data.
It is important to note that the input-output pairs of images of the neural network are exclusively uncorrected scatter–scatter ratio images. That is to say, the models are not trained to distinguish if the input image is affected by the scatter contribution, so if giving a scatter-free projection to the network, it will still yield some scatter estimation. The subtraction of this estimated image from a scatter-free input image would entail some loss of information in the final result. Therefore, these models cannot be applied to acquisitions taken with an anti-scatter grid or any other scatter suppression technique, neither can be used to check the effectiveness in scatter removal of those hardware-based scatter suppression methods. A neural network that identifies if the input image is affected by scattered rays is currently being implemented, but it is beyond the scope of this work.
As explained in
Section 2.3, a ROI focus primarily on the lungs region was selected in the training, validation and test images (see
Figure 1). This way, the edges of the images that could cause the appearance of artifacts in the scatter estimation and the scatter-corrected images are removed. This fact allows us to get very accurate results, but it must be taken into account before putting the models into practice. An input image with a large amount of empty regions could compromise the robustness of the models, just as it is suggested by the results obtained for images simulated SDD=200 cm.
Although the number of training and validation cases might seem small, the variety in the patient sizes within the dataset (explained in
Section 2.1) along with the data augmentation described in
Section 2.3 guarantees the accuracy of the models for images with different amounts of scatter ratio.
Regarding the study of the contrast improvement after scatter correction, in this work we focused on COVID-19 since a large number of databases related to this disease has been gathered in the past two years as a consequence of the worldwide pandemic. However, the same analysis could be performed for any other pulmonary affection, as pneumonia, tumors, atelectasis or pneumothorax. It is highly expected that the scatter correction will also entail a contrast improvement in areas affected by any of these lesions, and therefore facilitate its identification.
It should be taken into account that an overestimation in the contrast value could result in a loss of important information for medical diagnosis in the final scatter-corrected image. Results in
Table A1 show that the contrast is overestimated in 7 out of 31 cases in the single-energy model, in 14 cases in the 1-output dual-energy model, and in 11 cases in the 2-output dual-energy model. Nevertheless, the difference with the ground truth value is on average just 1.11% for the single-energy method, 1.48% for the 1-output dual-energy network, and 1.01% for the 2-output dual-energy model. Thus, the overestimation is not significant and will not jeopardize the image information.
5. Conclusions
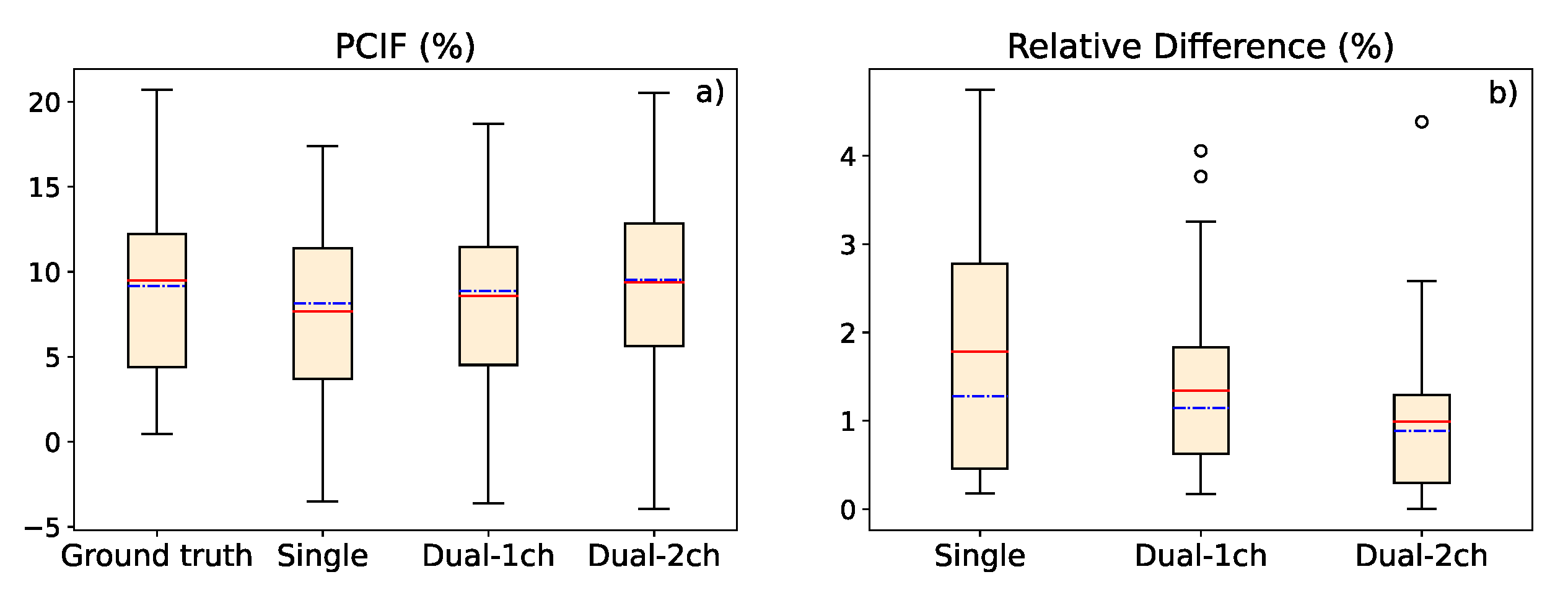
In this work, we presented three deep learning-based methods to estimate the scatter contribution in CXRs and obtain the scatter-corrected projections: a single-energy model, with one input and one output image; a 1-output dual-energy model, in which a projection acquired with a different energy spectrum is set as an additional input channel, but the output has only one channel; and a 2-output dual-energy model, in which the scatter estimation is provided for the two energies introduced in the two input channels. The three models were robust to variations in the SDD, obtaining a high precision for distances in a range between 100 and 200 cm, and proving that a similar accuracy with respect to images with the original SDD of the training data (SDD=180 cm) can be maintained. Besides, the contrast values between the lung region affected by COVID-19 and the healthy region in soft tissue images (obtained by means of dual energy subtraction) demonstrated that scatter correction in CXRs provides better contrast to the area of the lesion, yielding a PCIF of up to 20.5%.
In both studies (accuracy in scatter correction for several SDD and contrast value in COVID-19 region), the dual-energy algorithms provide results with a better accuracy. The analysis of the p-value points out that the difference in the accuracy between the single-energy and dual-energy models can be statistically significant, especially in the scatter-corrected estimations of high energy CXRs with the 2-output method. The single-energy algorithm is accurate enough for scatter correction of CXRs, so it might not be worthy to acquire an extra CXR just for this purpose. However, dual-energy models are proven to be a useful tool for scatter correction on soft tissue DES CXRs.
The single-energy model was tested with a cohort of real CXRs acquired without an anti-scatter grid, yielding robust, qualitative estimations of the scatter correction, even for images with artifacts. Further studies with real phantoms and patients should be performed to quantitatively determine the precision of the three models with real data, and to analyze whether the difference between the performance of the single and dual-energy algorithms is relevant.
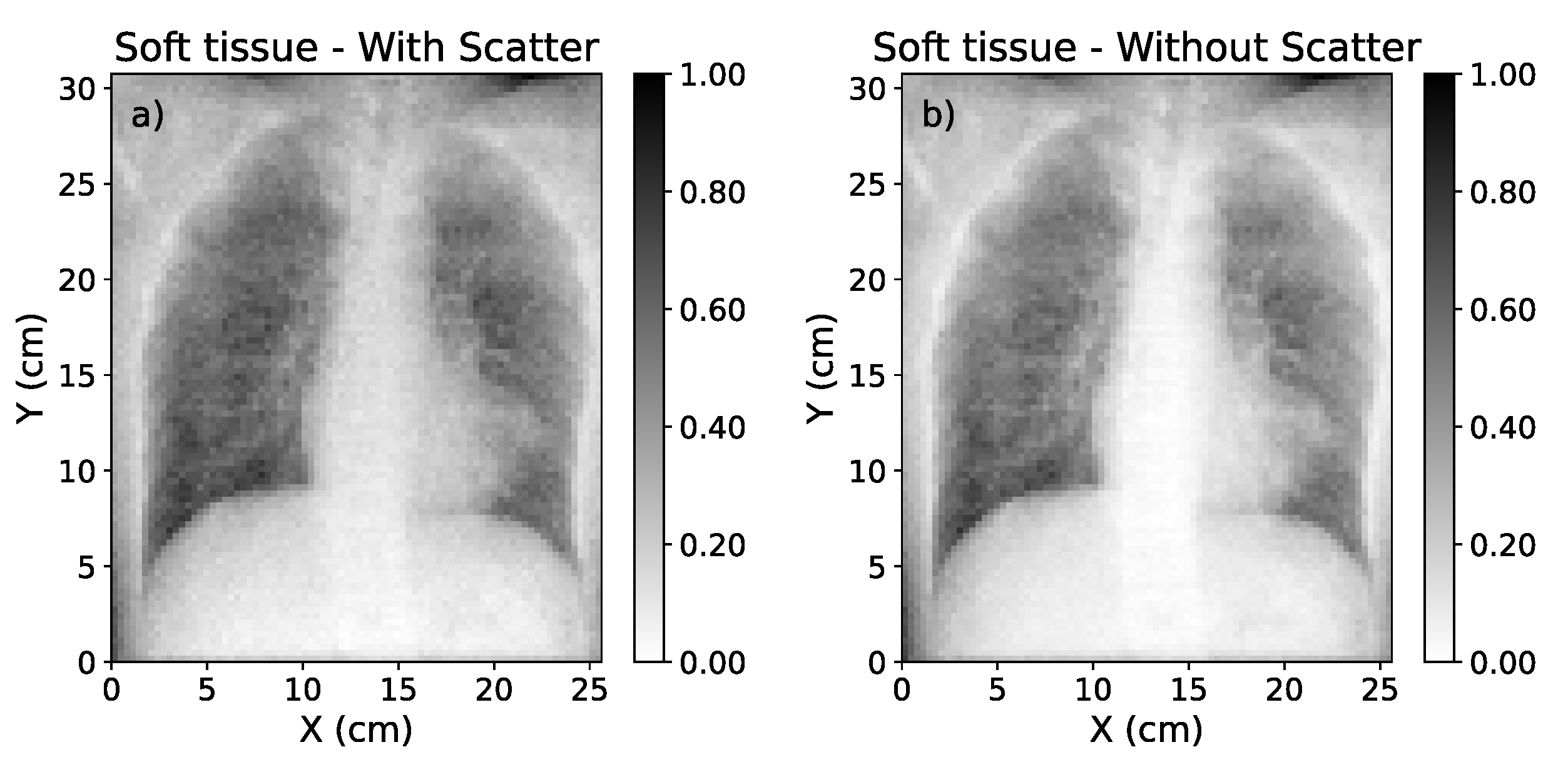
Figure 1.
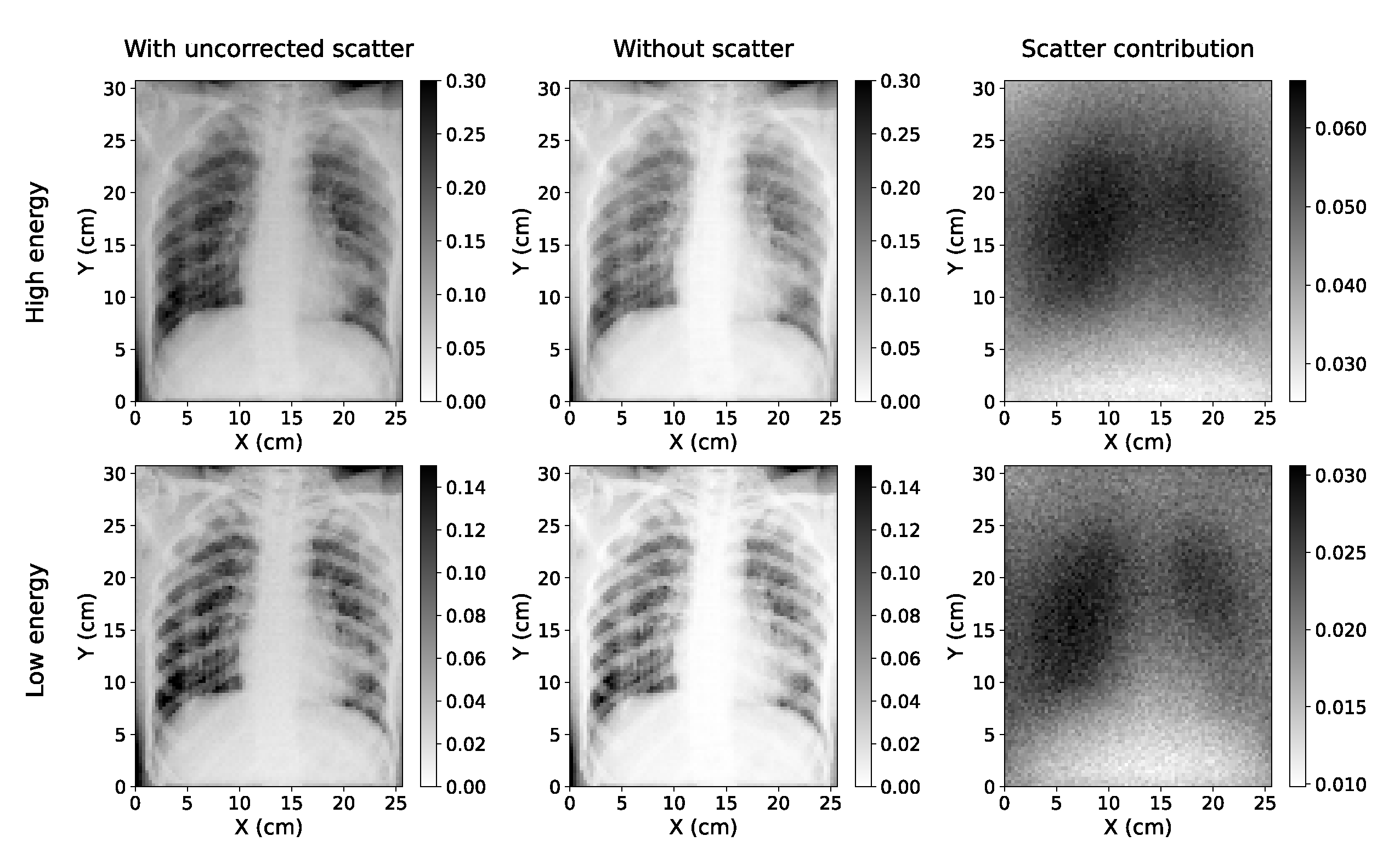
Simulated Chest X-rays for two cases considered (low energy=60 kVp; high energy=130 kVp). The simulation with scatter (left) can be decomposed into a direct component ("without scatter", center) and the scatter contribution (right).
Figure 1.
Simulated Chest X-rays for two cases considered (low energy=60 kVp; high energy=130 kVp). The simulation with scatter (left) can be decomposed into a direct component ("without scatter", center) and the scatter contribution (right).
Figure 2.
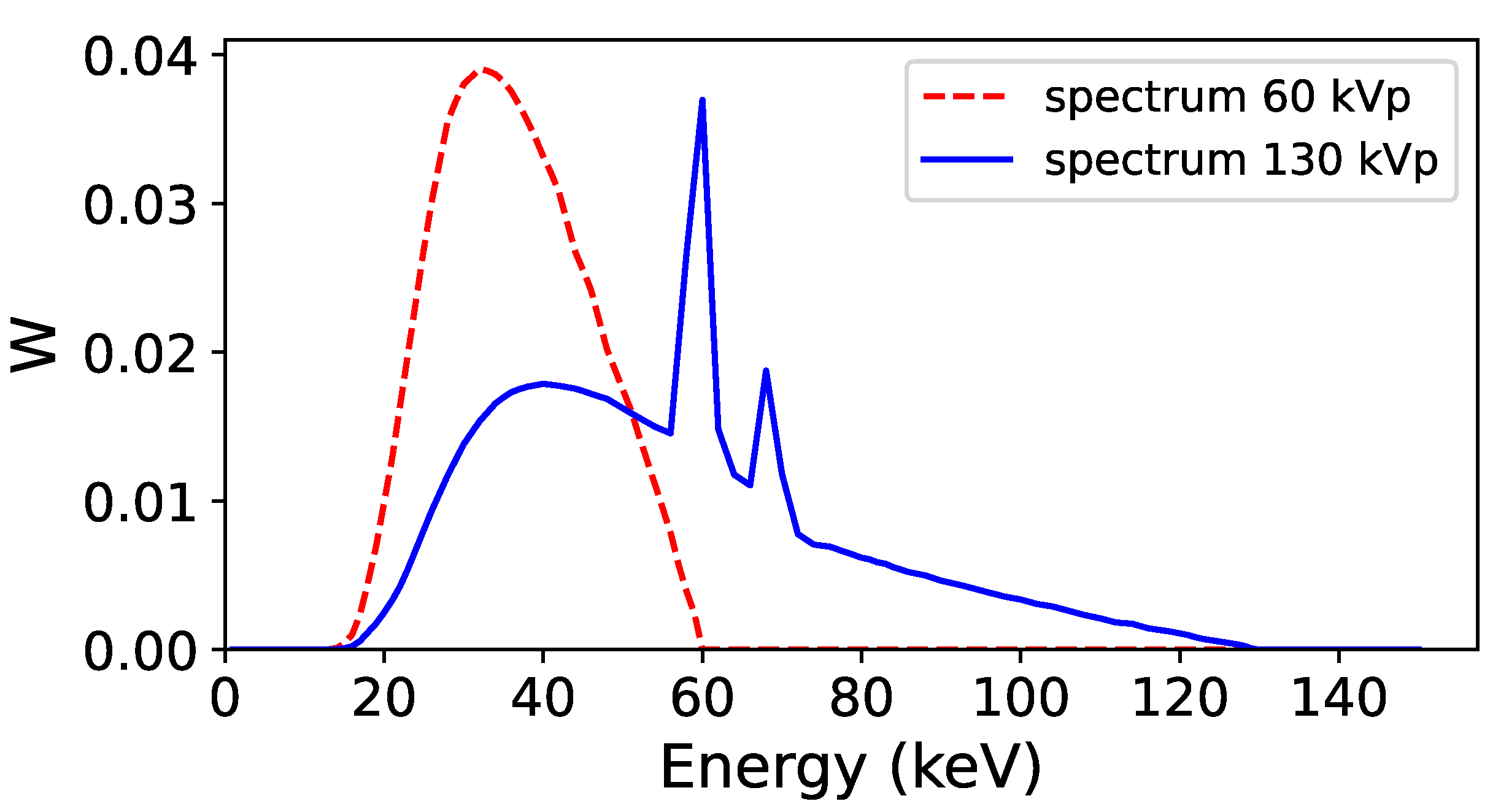
Energy spectra used to acquire the low energy (60 kVp, red line) and high energy (130 kVp, blue line) projections in the Monte Carlo simulation. The two spectra were obtained with Specktr toolkit [
42,
43], with 1.6 mm Al inherent filtration of the tube.
Figure 2.
Energy spectra used to acquire the low energy (60 kVp, red line) and high energy (130 kVp, blue line) projections in the Monte Carlo simulation. The two spectra were obtained with Specktr toolkit [
42,
43], with 1.6 mm Al inherent filtration of the tube.
Figure 3.
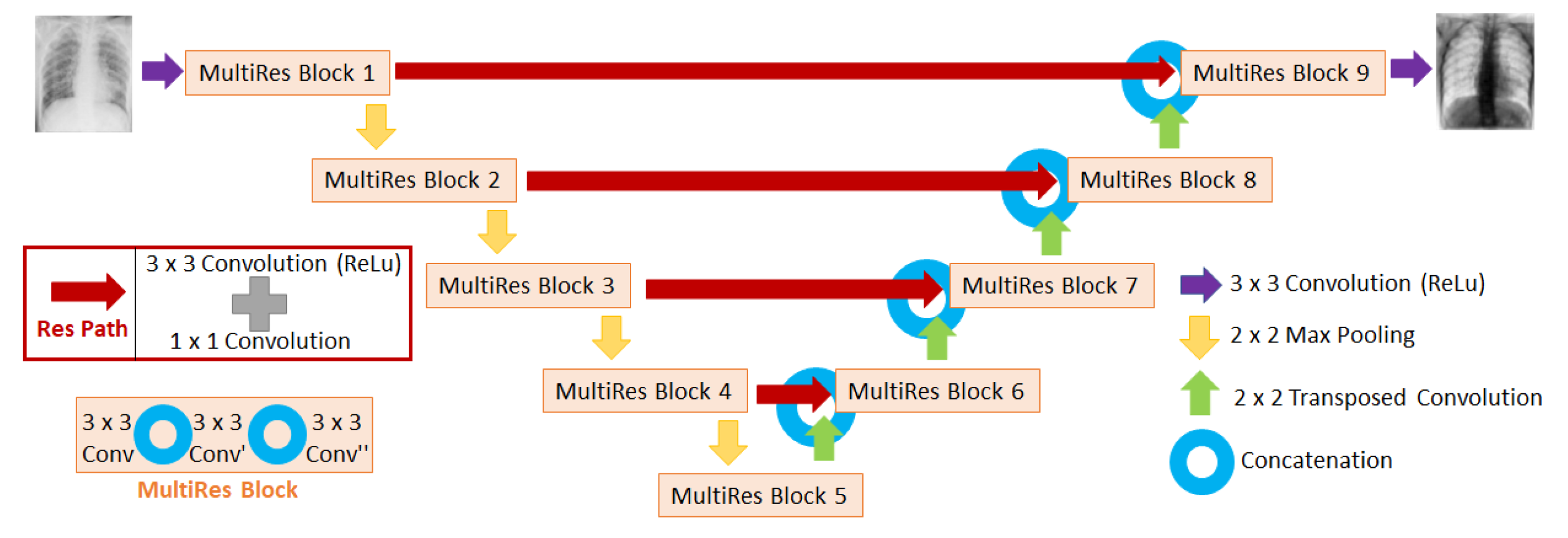
Diagram of the MultiResUNet architecture used in this work to train the neural networks. The input of the network is the image affected by scatter (i.e. uncorrected scatter image), and the output is the fraction of the image of scatter with respect to the uncorrected scatter image.
Figure 3.
Diagram of the MultiResUNet architecture used in this work to train the neural networks. The input of the network is the image affected by scatter (i.e. uncorrected scatter image), and the output is the fraction of the image of scatter with respect to the uncorrected scatter image.
Figure 4.
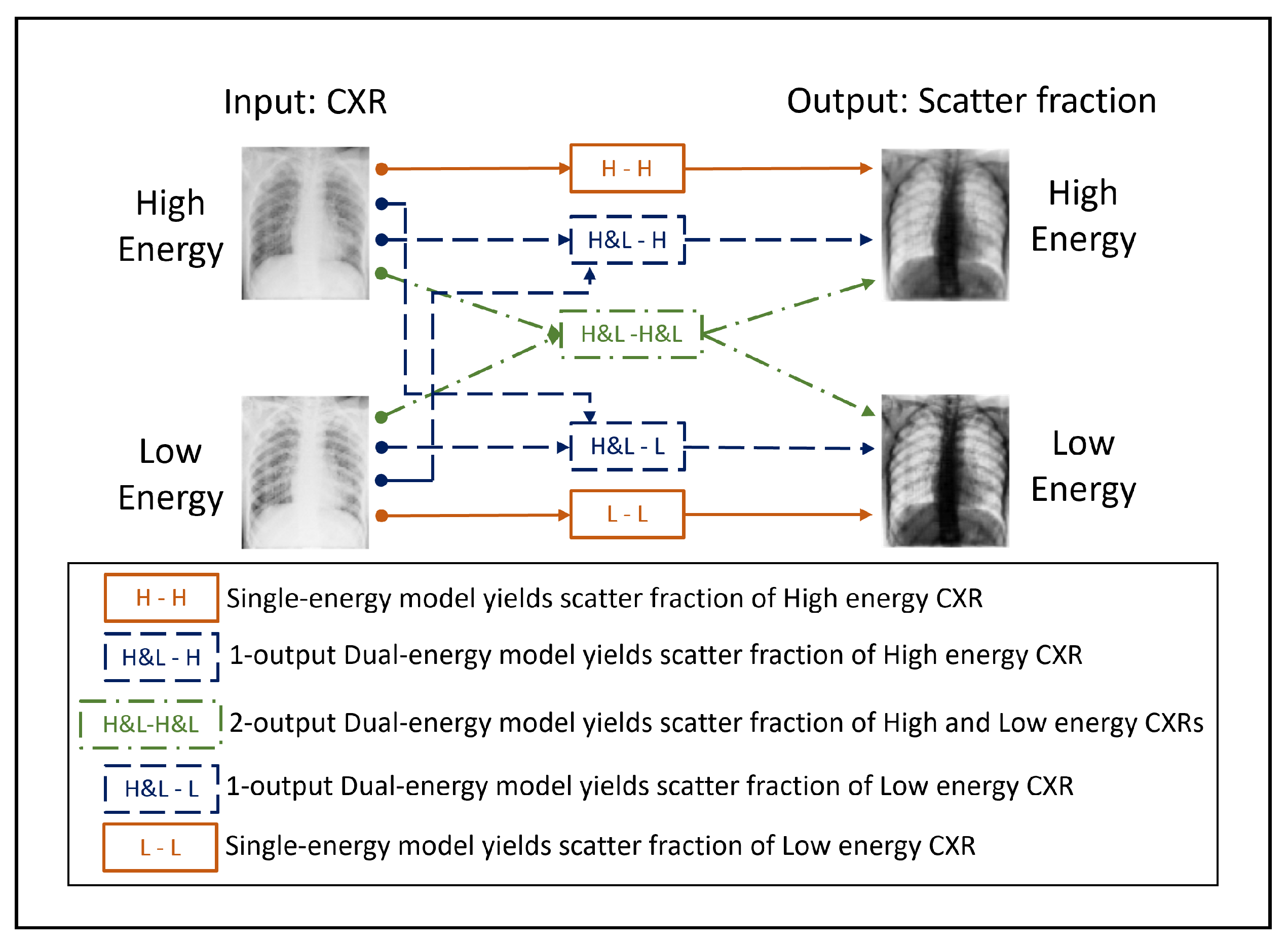
Scheme of the input and output images corresponding to the 3 neural network models presented in this work. The NNs differ on the amount of input and output channels used.
Figure 4.
Scheme of the input and output images corresponding to the 3 neural network models presented in this work. The NNs differ on the amount of input and output channels used.
Figure 5.
Soft tissue dual energy subtraction images: (a) Calculated from uncorrected-scatter CXRs of 60 kVp and 130 kVp. (b) Calculated from ground-truth scatter corrected images of 60 kVp and 130 kVp.
Figure 5.
Soft tissue dual energy subtraction images: (a) Calculated from uncorrected-scatter CXRs of 60 kVp and 130 kVp. (b) Calculated from ground-truth scatter corrected images of 60 kVp and 130 kVp.
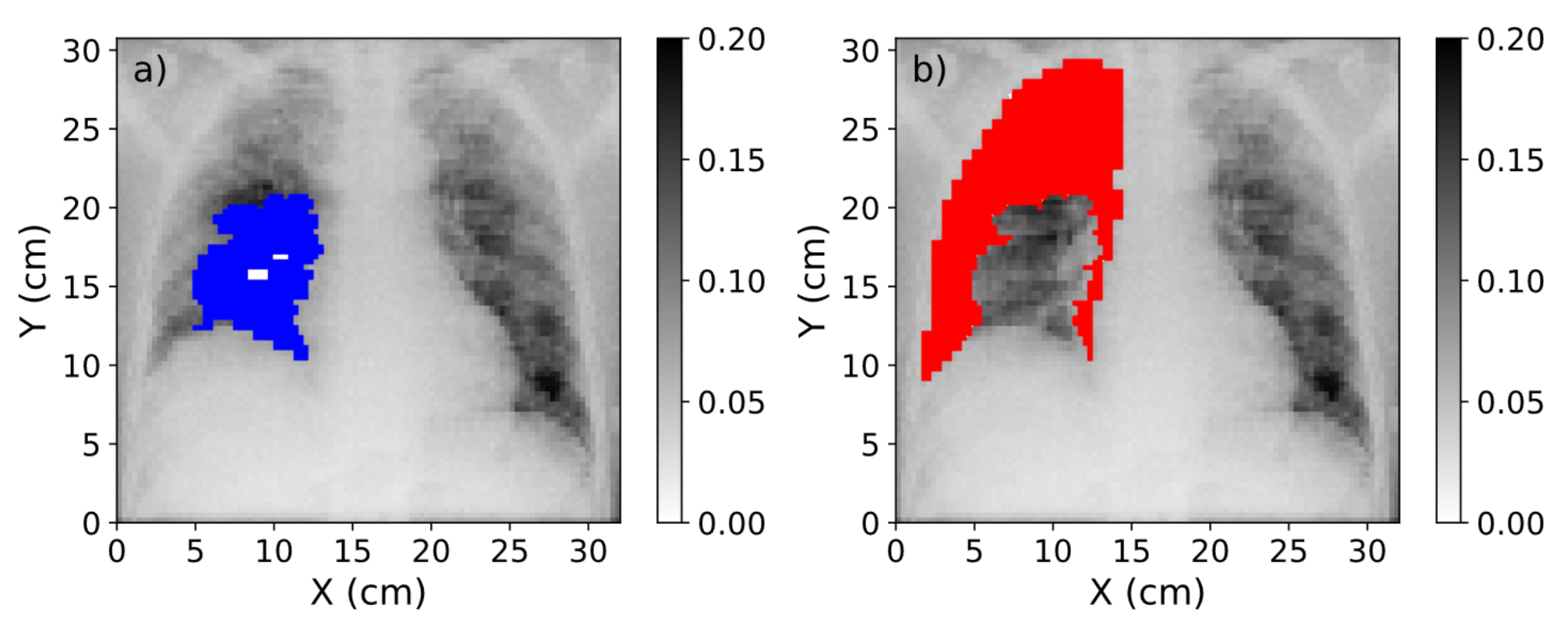
Figure 6.
(
a) Mask of the region affected by COVID-19 (blue) over the CXR. (
b) Mask of the healthy region in the lung (red). The range of values of these images was obtained after the normalization procedure explained in
Section 2.2.
Figure 6.
(
a) Mask of the region affected by COVID-19 (blue) over the CXR. (
b) Mask of the healthy region in the lung (red). The range of values of these images was obtained after the normalization procedure explained in
Section 2.2.
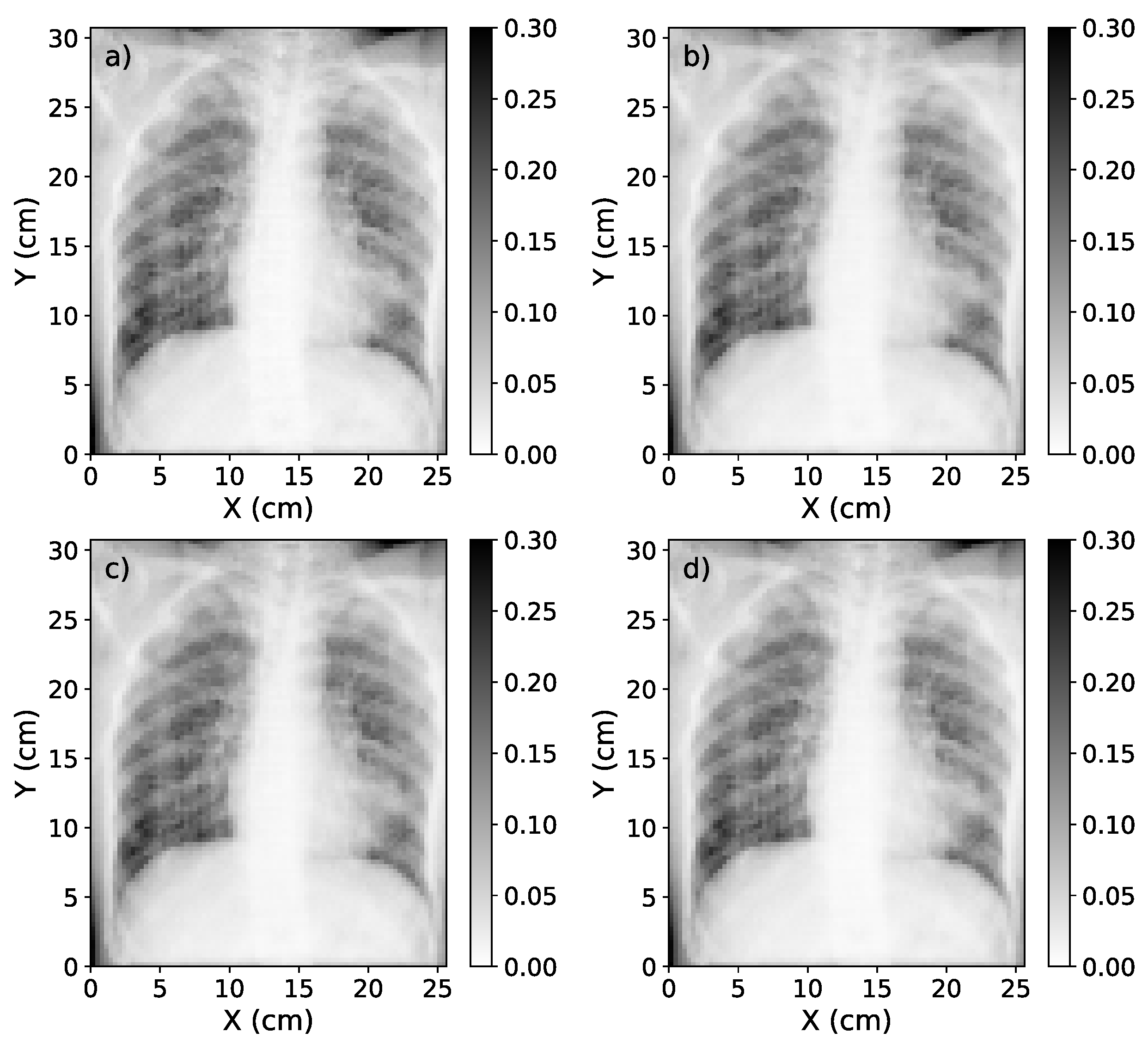
Figure 7.
(a) Representation of the scatter-corrected ground truth image. (b) The scatter-corrected image estimated by the single-energy model. (c) The scatter-corrected image estimated by the 1-output dual-energy model. (d) The scatter-corrected image estimated by the 2-output dual-energy model.
Figure 7.
(a) Representation of the scatter-corrected ground truth image. (b) The scatter-corrected image estimated by the single-energy model. (c) The scatter-corrected image estimated by the 1-output dual-energy model. (d) The scatter-corrected image estimated by the 2-output dual-energy model.
Figure 8.
Box plot of the MSE, MAPE, SSIM and relative error for the 22 test cases at different source-to-detector distances. Blue, dashed line represents the median value of the metric for each SDD. The box extends from the lower to the upper quartile values of the data, while the range (also referred to as whiskers) shows the rest of the distribution.
Figure 8.
Box plot of the MSE, MAPE, SSIM and relative error for the 22 test cases at different source-to-detector distances. Blue, dashed line represents the median value of the metric for each SDD. The box extends from the lower to the upper quartile values of the data, while the range (also referred to as whiskers) shows the rest of the distribution.
Figure 9.
a) Graphic representation of the percentage contrast improvement factor for ground truth image, single-energy model estimation, 1-output dual-energy model and 2-output dual-energy model estimation. (b) Relative difference in the contrast value between ground truth and deep learning-based estimations. In both graphics, the red, solid line represents the average value, while the blue, dash-dotted line represents the median value.
Figure 9.
a) Graphic representation of the percentage contrast improvement factor for ground truth image, single-energy model estimation, 1-output dual-energy model and 2-output dual-energy model estimation. (b) Relative difference in the contrast value between ground truth and deep learning-based estimations. In both graphics, the red, solid line represents the average value, while the blue, dash-dotted line represents the median value.
Figure 10.
Original CXR (with scatter) with pixel value conversion (left); estimation of the scatter-corrected CXR (center); and estimation of the scatter contribution on real CXR (right) of three of the real chest X-ray images used to test the single-energy model of scatter correction.
Figure 10.
Original CXR (with scatter) with pixel value conversion (left); estimation of the scatter-corrected CXR (center); and estimation of the scatter contribution on real CXR (right) of three of the real chest X-ray images used to test the single-energy model of scatter correction.
Table 1.
Parameters of the MC simulation.
Table 1.
Parameters of the MC simulation.
| Parameter |
Specification |
| Source-Detector Distance (cm) |
180 |
| X-ray Detector Size (cm) |
|
| X-ray Detector Resolution (pixel) |
|
Table 2.
Ratio between a region of the lung with and without rib in the original, real CXRs and in the scatter-corrected CXRs yielded by the single-energy algorithm for the 10 CXRs taken as test set (listed as C1-C10), and the resulting average value.
Table 2.
Ratio between a region of the lung with and without rib in the original, real CXRs and in the scatter-corrected CXRs yielded by the single-energy algorithm for the 10 CXRs taken as test set (listed as C1-C10), and the resulting average value.
| |
C1 |
C2 |
C3 |
C4 |
C5 |
C6 |
C7 |
C8 |
C9 |
C10 |
Avg |
| Original |
2.71 |
1.66 |
1.36 |
1.11 |
1.35 |
1.16 |
1.71 |
1.47 |
1.28 |
1.59 |
1.54 |
| Scatter-corrected |
3.80 |
2.02 |
1.54 |
1.17 |
1.49 |
1.23 |
2.14 |
1.68 |
1.41 |
1.64 |
1.81 |