1. Summary
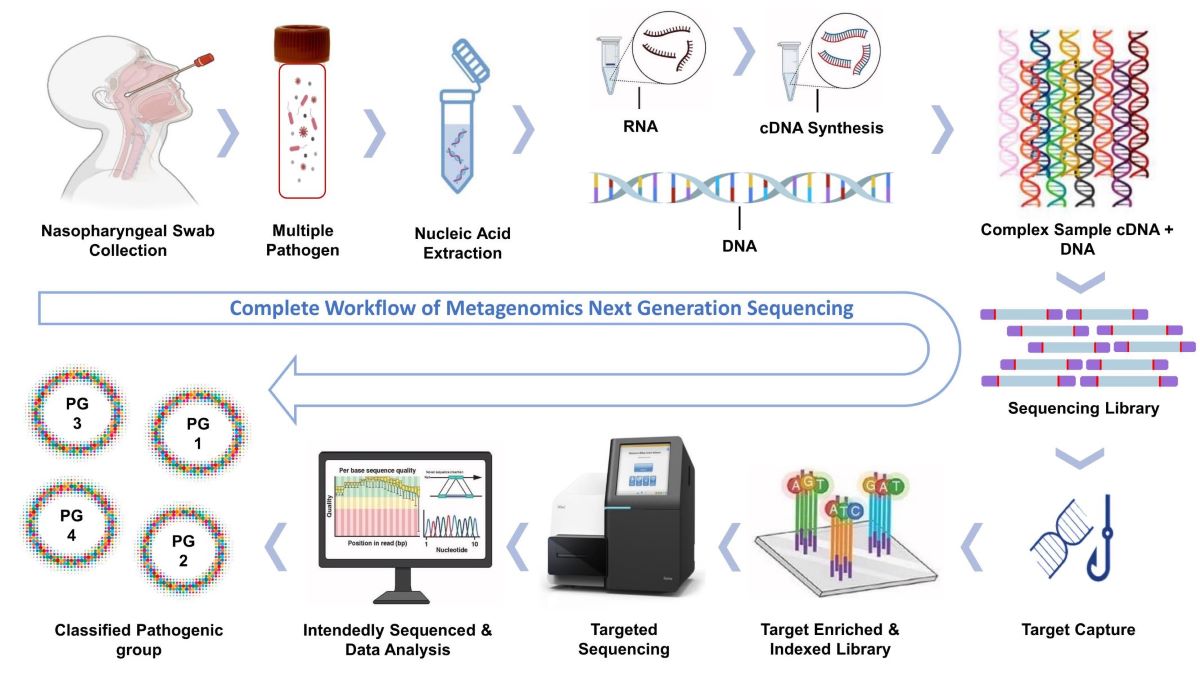
This study collected 29 nasopharyngeal swab samples from patients undergoing PCR testing for respiratory pathogens at Advanta Genetics (Tyler, TX, USA) between January 2022 and June 2022. Nucleic acids, DNA and RNA were extracted separately using Zymo research reagents and following the protocol in the Explify Respiratory Pathogen ID /AMR Panel User Guide. T7 bacteriophage DNA was added to each sample to determine the absolute concentration of the target copies. Next-generation metagenomic sequencing (mNGS) libraries were prepared using the Illumina®/IDbyDNA Respiratory Pathogen ID /AMR Panel (RPIP) protocol[
1].
The data in this study represents a comprehensive metagenomic analysis of acute upper respiratory infections (URIs), offering comparative insights for researchers and clinicians in respiratory medicine, infectious diseases, epidemiology, and public health. The dataset provides unique precision in identifying URI-related pathogens. It stratifies identified organisms based on their potential to induce lower respiratory infections, presenting new avenues for microbiological diagnostics and treatment[
1]. The study illustrates the multifaceted benefits of this dataset, encompassing comprehensive microbiota characterization, identification of uncharacterized microorganisms, implications for respiratory medicine, infectious diseases, epidemiology, and public health, and its integration potential with other omics datasets. Accordingly, the dataset promises advancements in microbiological diagnostics [
2].
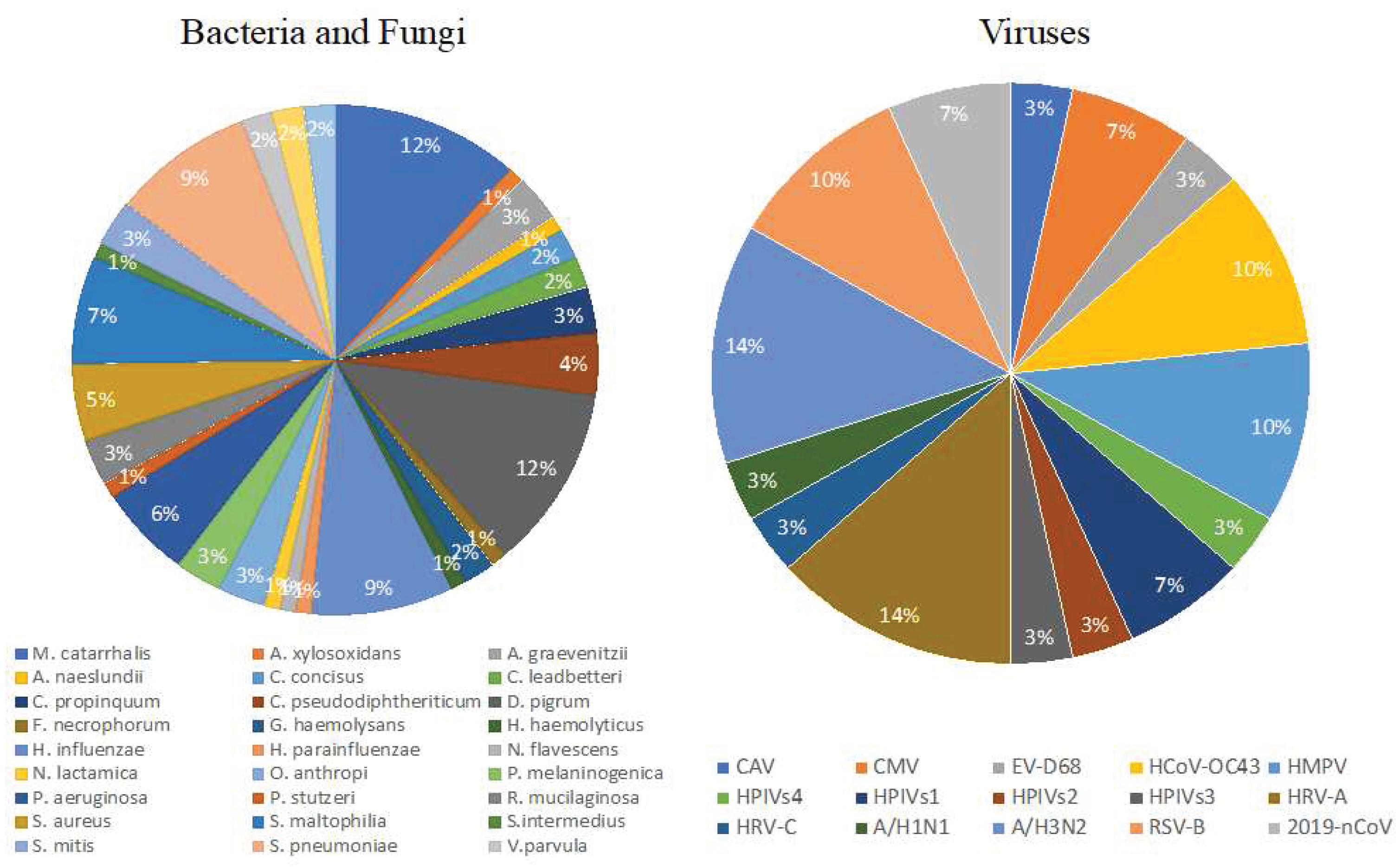
This dataset of URI microbiota offers several clinical benefits that aid in understanding the microbial communities present in the upper respiratory tract. The data provides detailed information on the prevalence, abundance, and diversity of known microbial species, accurately identifying microbes, and shedding light on the etiology and pathogenesis of URIs and their trafficking potential[
3]. Furthermore, the dataset's ability to discern mixed infections involving multiple species is instrumental in addressing polymicrobial pathogenesis. Enabling the detection and differentiation of these mixed infections helps clinicians tailor treatment strategies to manage complex respiratory disease effectively[
4]. The data also offers a snapshot into a better understanding of microbial dysbiosis in microbial communities, aiding in the understanding of how shifts in the microbiota might contribute to infections and related conditions and help scientists and epidemiologists study the prevalence, distribution, and evolution of pathogens causing URIs. Consequently, this information can also aid in monitoring outbreaks and tracking the emergence of new strains[
5].
The metagenomics component of the dataset enables the detection and comprehensive characterization of previously uncharacterized or unculturable microorganisms and provides a potential avenue for biomarker discovery. Microbiota profiling through mNGS can potentially reveal specific microbial signatures or biomarkers associated with different URIs. The discovery of novel microbial species or strains can expand our knowledge of respiratory infections and potentially lead to the development of new diagnostic tools or therapeutic interventions[
6].
This study augments the microbial context of URIs, enriching respiratory disease research. The dataset unveils complex host-microbe interplays, exposing potential pathogenesis, epidemiology, and therapeutic possibilities not readily available by conventional laboratory techniques. Supplementary advantages of the metagenomics data are that it allows clinicians and researchers to identify previously uncultivable or fastidious pathogens, enabling the characterization of dysbiosis, which has been implicated in respiratory diseases like chronic obstructive pulmonary disease (COPD) and asthma[
7]. Furthermore, the dataset illuminates host-microbe interactions, enhancing understanding of immune responses and disease mechanisms. It unravels microbial factors and pathways pivotal for disease, potentially guiding innovative therapeutic strategies. Manipulating these factors might lead to developing novel therapeutic approaches, such as targeting specific virulence factors or modulating dysbiotic microbial populations in many pulmonary diseases[
8].
The data affords epidemiologists and public health experts to gain better insights into URIs epidemiology and transmission patterns. Microbiota composition and lower respiratory trafficking potential data inform surveillance and outbreak investigations. Furthermore, metagenomics-based microbiota characterization offers valuable insights into the distribution and prevalence of specific pathogens across cultures, populations, and geographic domains. This contributes to epidemiological studies that assess patterns of infection spread, potential outbreaks, and transmission dynamics. Such URI data represents a paradigm shift in respiratory disease research and epidemiological investigation and can help guide public health interventions and the development of targeted preventive strategies[
9].
Overall, this dataset's insights into acute URI microbiota contribute to respiratory research, diagnosis, and treatment domains. Integration with omics datasets augments its potential, propelling optimization and advancements in microbiological laboratory workflows and clinical diagnostics[
9]. This work paves the way for a deeper comprehension of URI-related complexities, underscoring the collaborative efforts of respiratory medicine, infectious diseases, epidemiology, and public health.